* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
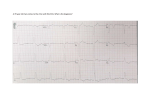
Download Therapeutic Area Data Standards User Guide for QT
Survey
Document related concepts
Transcript
Therapeutic Area Data Standards User Guide for QT Version 1.0 (Draft) Prepared by the CFAST QT Team CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Notes to Readers • • • This is the draft version 1.0 of the Therapeutic Area Data Standards User Guide for QT. This document corresponds to the SDTM v1.4 and SDTMIG v3.2 and to the ADaM v2.1 and ADaMIG v1.0. The TAUG-QT v1.0 package contains a user guide, a draft domain model, and a draft specification for a new relationships dataset. Revision History Date TBD 2014-07-31 Version 1.0 1.0 Draft Summary of Changes Provisional Draft for Public Review See Appendix F for Representations and Warranties, Limitations of Liability, and Disclaimers. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 2 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) CONTENTS 1 INTRODUCTION .............................................................................................................. 5 1.1 1.2 1.3 1.4 1.5 1.6 1.7 PURPOSE.............................................................................................................................................................5 CLINICAL GUIDELINES FOR QT STUDIES ............................................................................................................6 ORGANIZATION OF THIS DOCUMENT...................................................................................................................7 CONCEPT MAPS ..................................................................................................................................................7 CONTROLLED TERMINOLOGY .............................................................................................................................8 RELATIONSHIPS TO OTHER STANDARDS .............................................................................................................8 KNOWN ISSUES...................................................................................................................................................9 2 ECG OVERVIEW .............................................................................................................. 9 2.1 QT/QTC ASSESSMENTS IN CLINICAL STUDIES ...................................................................................................9 2.2 EXPLANATION OF COMMON TERMS ..................................................................................................................10 2.3 THE FUNDAMENTALS OF ECGS ........................................................................................................................12 2.3.1 What is an ECG and What Information Does it Provide? ........................................................................12 2.3.2 ECG Machinery .......................................................................................................................................16 2.3.3 ECG Device Types ...................................................................................................................................20 2.3.4 How is the QT Interval Adjusted (Corrected) for Heart Rate? .................................................................21 3 THE THOROUGH QT (TQT) STUDY .......................................................................... 23 4 TRIAL DESIGN ................................................................................................................ 26 4.1 TQT STUDY DESIGN.........................................................................................................................................26 4.1.1 Parallel Studies ........................................................................................................................................26 4.1.2 Crossover Studies.....................................................................................................................................27 4.1.3 Trial Elements ..........................................................................................................................................29 4.1.4 Trial Summary Parameters .......................................................................................................................30 4.2 TIME POINT PLANNING .....................................................................................................................................32 5 SUBJECT CHARACTERISTICS AND ELIGIBILITY............................................... 35 5.1 5.2 INCLUSION/EXCLUSION CRITERIA ....................................................................................................................35 PHARMACOGENETICS .......................................................................................................................................37 6 STUDY ASSESSMENTS.................................................................................................. 38 6.1 ECG ASSESSMENTS..........................................................................................................................................38 6.1.1 Specification for ECG Test Results ..........................................................................................................40 6.1.2 Examples for ECG Test Results ...............................................................................................................44 6.2 QT CORRECTION ........................................................................................ERROR! BOOKMARK NOT DEFINED. 6.2.1 Specification for ECG QT Correction ......................................................................................................51 6.2.2 Examples for ECG QT Corrections .........................................................................................................53 6.3 PK ASSESSMENTS ......................................................................................ERROR! BOOKMARK NOT DEFINED. 6.4 HEMODYNAMICS/VITAL SIGNS .........................................................................................................................55 6.4.1 Example for Orthostatic Challenge ..........................................................................................................58 7 DATA ANALYSIS ............................................................................................................. 59 7.1 ADEG EXAMPLE ..............................................................................................................................................60 7.1.1 Analysis Data Metadata ...........................................................................................................................60 7.1.2 Analysis Dataset.......................................................................................................................................64 7.2 ADQT EXAMPLE ..............................................................................................................................................67 7.2.1 Analysis Data Metadata ...........................................................................................................................67 7.2.2 Analysis Dataset.......................................................................................................................................68 7.3 BASELINE ALTERNATIVES AND DEFINITION OF TREATMENT DIFFERENCES ......................................................69 Page 3 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 7.3.1 7.3.2 7.3.3 7.3.4 Time-Matched Baseline; Double-Delta Treatment Difference.................................................................69 Time-Averaged Baseline; Double-Delta Treatment Difference ...............................................................71 Predose Averaged Baseline; Double-Delta Treatment Difference ...........................................................72 No Baseline; Single-Delta Treatment Difference .....................................................................................74 APPENDICES ............................................................................................................................. 75 APPENDIX A: PROJECT PROPOSAL ................................................................................................................................75 APPENDIX B: CFAST ORGANIZATIONS ........................................................................................................................75 APPENDIX C: WORKGROUP ..........................................................................................................................................76 APPENDIX D: GLOSSARY AND ABBREVIATIONS ............................................................................................................77 APPENDIX E: REFERENCES ...........................................................................................................................................78 Appendix E1: Figures ..............................................................................................................................................79 Appendix E2: Further Reading ................................................................................................................................79 APPENDIX F: REPRESENTATIONS AND WARRANTIES, LIMITATIONS OF LIABILITY, AND DISCLAIMERS..........................80 LIST OF FIGURES Figure 1: CDISC Industry Wide Data Standards ...........................................................................................................5 Figure 2: Concept Classification Coding Key for Concept Maps..................................................................................8 Figure 3: Diagram of the Heart’s Conduction System................................................................................................. 13 Figure 4: Sequence of Heart Excitation and the Associated ECG Waveforms............................................................ 14 Figure 5: Electrocardiogram Waveform Illustration.................................................................................................... 15 Figure 6: Standard Electrode/Lead Configuration for a 12-Lead/Waveform ECG ..................................................... 17 Figure 7: 12-Lead Electrocardiogram Waveform ........................................................................................................ 18 Figure 8: High-Level View of ECG Hardware-Software: From Subject’s Chest to Data for Analysis....................... 19 Figure 9: Relationship between the Bazett- and Fridericia-Corrected QT Interval and RR Interval ........................... 22 Figure 10: Parallel Study Design Schema for Example TQT Study 1 ......................................................................... 26 Figure 11: Crossover Study Design for Example TQT Study 2 with Washout Period Combined with each Treatment Element ...................................................................................................................................................... 28 Figure 12: Crossover Study Design for Example TQT Study 2 with Separate Elements for each Washout Period.... 28 Figure 13: Individual Beats and Relevant Identifying Information Within a 24-hour Continuous Recording ............ 33 Figure 14: ECG Replicates and Nominal Time Points ................................................................................................ 34 LIST OF CONCEPT MAPS Concept Map 1: Recording of ECG Digital Waveforms ............................................................................................. 19 Concept Map 2: Measurement of RR and Intervals for Use in Analysis ..................................................................... 20 Concept Map 3: Potential Parameters and Their Order of Collection ......................................................................... 35 Concept Map 4: ECG Quantitative Results and Morphological (Qualitative) Findings Determination ...................... 39 Concept Map 5: Population QT Correction for a Pre-specified Model ....................................................................... 49 Concept Map 6: Individual QT Correction for a Pre-specified Model ........................................................................ 49 Concept Map 7: Individual QT Correction (QTc) ‘Best Fit’ Model ............................................................................ 50 Concept Map 8: PK/PD Analysis Steps and Timing ................................................................................................... 55 Concept Map 9: Assessment of Orthostasis ................................................................................................................ 56 © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 4 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 1 Introduction This Therapeutic Area Data Standards User Guide for QT (TAUG-QT) was developed under the Coalition for Accelerating Standards and Therapies (CFAST) initiative (Appendix A). CFAST, a joint initiative of Clinical Data Interchange Standards Consortium (CDISC) and the Critical Path Institute (C-Path), was launched to accelerate clinical research and medical product development by facilitating the establishment and maintenance of data standards, tools, and methods for conducting research in therapeutic areas important to public health. CFAST partners include TransCelerate BioPharma Inc. (TCB), the U.S. Food and Drug Administration (FDA), and the National Cancer Institute Enterprise Vocabulary Services (NCI EVS), with participation and input from many other organizations. See Appendix B for a description of CFAST participating organizations. CDISC has developed industry-wide data standards enabling the harmonization of clinical data and streamlining research processes from protocol (study plan) through analysis and reporting, including the use of electronic health records to facilitate study recruitment, study conduct and the collection of high quality research data. CDISC standards, implementations and innovations can improve the time/cost/quality ratio of medical research, to speed the development of safer and more effective medical products and enable a learning healthcare system. 19 20 21 22 23 24 The goal of the CFAST initiative is to identify a core set of clinical therapeutic area concepts and endpoints for targeted therapeutic areas and translate them into CDISC standards to improve semantic understanding, support data sharing and facilitate global regulatory submission. 25 1.1 Purpose 26 27 28 29 30 31 32 33 34 35 36 37 Figure 1: CDISC Industry Wide Data Standards This draft version 1.0 (v1.0) of the TAUG-QT highlights the data endpoints for clinical studies characterizing the QT effects of drugs in healthy volunteers or in patients. The TAUG-QT endeavours to cover all studies specifically directed at the QT invertal, with extra attention to a specific type of study, the “thorough QT (TQT) study,” which is operationally defined by an ICH E14 guidance document 1 and the associated Question & Answer document 2 (see Section 3 for additional details). See Appendix A for the project proposal that was approved by the CFAST Steering Committee. Because this TAUG is directed at the evaluation of the QT in clinical studies and even more specifically at the conduct and analysis of the TQT study, certain aspects of the processing of QT data (e.g., correction of the QT interval for heart rate, potential complexities of evaluating and interpreting a change in QT) will differ from the processing of QT data used in routine clinical practice. While routine clinical practices focus on the individual patient and substantial changes in QT and/or absolute QT values, TQT studies can find even relatively small Page 5 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 differences between groups of patients treated with an experimental drug compared to a control treatment to be of interest. This difference in interests leads to the need for different processing. Also, TQT studies are generally limited to adult healthy volunteers of a non-geriatric age and having normal ECGs at screening. Data collection is also performed under highly controlled, inpatient conditions. Therefore, a number of considerations in other clinical trials are not emphasized in this TAUG. For example, while potential differences in incidence of adverse events based on various demographic characteristics might be included for some large Phase 3 studies, these would be unusual analyses to include in a TQT study. It would also be highly unexpected to observe adverse events related to cardiac ischemia, congestive heart failure, or other non-electrophysiological aspects of cardiac function, and therefore such events do not receive special attention in this TAUG. An analysis of all adverse events during the course of any study would be expected as a routine component of any study report. This TAUG-QT v1.0 describes the most common data needed for QT interval or heart rate corrected QT interval (QTc) studies, especially TQT studies, so that those handling the data (e.g., data managers, statisticians, programmers) understand the data and can apply standards appropriately. Descriptions addressed in this TAUG-QT v1.0 include the clinical situations from which the data arise, and the reasons these data are relevant for QT/QTc studies. The TAUG-QT v1.0 also strives to define research concepts unambiguously, so that consistent terminology can be used in QT/QTc studies to enable aggregation and comparison of data across studies and drug programs. And finally, the TAUG-QT v1.0 describes how to use CDISC standards to represent the data: • For the Study Data Tabulation Model (SDTM) and the SDTM Implementation Guide for Human Clinical Trials (SDTMIG), these instructions include guidance on which domains data should be stored in, how variables should be used, and example datasets and controlled terminology for ECG replicate (generally conventional 10-second, 12-lead ECGs recorded as such or extracted from continuous recording of longer duration) and single beat measurements. • For the Analysis Data Model (ADaM) and ADaM Implementation Guide (ADaMIG), an example of metadata and dataset for ADEG are provided, and baseline alternatives for treatment differences are described and modeled with ADaM variables. These CDISC standards are freely available at http://www.cdisc.org. It is recommended that implementers consult the SDTM prior to implementing these QT clinical data standards. It is recommended that the reader be familiar with common electrocardiogram (ECG) terminology, which can be found in Section 2.2, as these terms have very specific connotations within the context of this document. A comprehensive ECG overview is also provided in Section 2.3 to ensure that the reader has a strong foundation to understand the intricacies of these QT data standards. It is important to note that the inclusion of concepts in this user guide should not be construed as a requirement to collect data on these concepts in any particular study of QT/QTc. The examples included are intended to show how data of particular kinds can be represented using CDISC standards, if such data are collected. They are not intended to imply that such data must be collected, or even that such data should be collected. This document does not contain guidance from a regulatory authority and should not be construed as substituting for or replacing any documents published by such an authority. 83 1.2 Clinical Guidelines for QT Studies 84 85 86 87 88 89 90 91 A research concept is a unit of knowledge created by a unique combination of characteristics that define observations of real world clinical research phenomena, including the question and assessment that result in a measurable outcome. Research concepts specify what should be observed for a specific subject assessment in a clinical study, but not how to capture the data or how to group observations together. Research concepts include data that represent a set of roles such as a topic or qualifiers. Research concepts represent clinical research knowledge that borrows from medical knowledge, statistical knowledge, BRIDG, and the CDISC standards. The authors of this user guide considered the International Conference on Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use (ICH) Efficacy (E) 14 Guidance for Industry: The Clinical Evaluation of QT/QTc © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 6 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 Interval Prolongation and Proarrhythmic Potential for Non-Antiarrhythmic Drugs (May 2005) 1 and associated Question and Answers (R1 and R2) documents (May 2012 and March 2014, respectively) 2, 3 when drafting this user guide. The guidance provides recommendations to sponsors concerning the design, conduct, analysis, and interpretation of clinical studies to assess the potential of a drug to delay cardiac repolarization. Tests for the effects of new agents on the QT interval and QTc, as well as the collection of cardiovascular adverse events, were noted as recommended assessments. The guidance also recommends that the investigational approach used for a particular drug should be individualized, depending on the pharmacodynamic (PD), pharmacokinetic (PK), and safety characteristics of the product, as well as on its proposed clinical use. The assessment of the effects of drugs on cardiac repolarization remains the subject of active investigation. Further details regarding ICH E14 guidance can be found in Section 3. This user guide focuses primarily on how to represent study design elements, time point planning, recommended exclusion criteria, pharmacogenetic considerations, study assessments and key data analysis components for QT studies. This document also covers elements of routinely collected data (e.g., medical history, adverse events) that might be of particular interest in QT studies. 1.3 Organization of this Document The TAUG-QT v1.0 differs from that of other CDISC therapeutic area user guides, which categorize the concepts covered into subject/disease characteristics, disease assessments, and routinely collected data, because QT studies are associated with drug safety rather than with a particular disease or condition to be treated. This is reflected in the organization of this document, as follows: • • • • • • • • Section 1, Introduction, provides an overall introduction to the purpose and goals of the QT project. Section 2, ECG Overview, provides a general overview of ECG terminology and concepts. Section 3, The Thorough QT (TQT) Study, provides background information regarding TQT studies. Section 4, Trial Design, covers trial design for studies evaluating QT, particularly TQT studies. Section 5, Subject Characteristics and Eligibility, highlights considerations for inclusion/exclusion criteria and pharmacogenetics in QT studies. Section 6, Study Assessments, provides insight into ECG assessments and machinery, PK assessments and hemodynamics/vital signs. Section 7, Data Analysis, includes key data analysis elements for a QT study. Appendices provide additional background material and describe other supplemental material relevant to the QT/QTc studies. A list of domains used in the examples in this document, and the sections in which these examples appear, is given below: 129 Domains from SDTMIG Findings EG – ECG Test Results QT – ECG QT Correction Model Data* VS – Vital Signs Trial Design TA – Trial Arms TE – Trial Elements TS – Trial Summary * Domain is not final. 130 1.4 Concept Maps 131 132 This document uses concept maps to explain clinical processes and research concepts. Concept maps, also sometimes called mind maps, are diagrams which include “bubbles” representing concepts/ideas/things and labeled Page 7 Draft Section 6.1.2 6.1.2 6.4.1 Domains from SDTMIG-MD Section DI – Device Identifier 2.3.3.1 DO – Device Properties 2.3.3.1 4.1.1, 4.1.2 4.1.3 4.1.4 © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 arrows that represent the relationships between the concepts/ideas/things. They are generally easier to draw and more accessible than more formal modeling diagrams, such as Unified Modeling Language (UML) diagrams. The diagrams in this document use the following coding for classification of concepts. This classification is based on classes in the Biomedical Research Integrated Domain Group (BRIDG) model. These color-symbol pairs have been used to highlight kinds of things that occur commonly in clinical data and therefore give rise to common patterns of data. Some concepts are not coded; they have a thinner, black outline, and no accompanying symbol. These may include the subject of an observation, as well as characteristics, or attributes, of the coded concepts. Figure 2: Concept Classification Coding Key for Concept Maps 1.5 Controlled Terminology CDISC Controlled Terminology is a set of standard value lists that are used throughout the clinical research process, from data collection through analysis and submission. Terminology applicable to SDTM and/or ADaM is either in production or under development by the CDISC Terminology Team at the time of publication of this document. Production terminology is published by the National Cancer Institute’s Enterprise Vocabulary Services (NCI EVS) and is available at: http://www.cancer.gov/cancertopics/cancerlibrary/terminologyresources/cdisc. CDISC Controlled Terminology is updated quarterly. Because this document is a static publication, it refers readers to the NCI EVS page for CDISC terminology (at the link given above). For the same reason, this document cannot claim to use controlled terminology in either the lists of laboratory tests, assessment procedures, or in the examples provided; users should not refer to these as the ultimate authority on what terms to use or to not use. 1.6 Relationships to Other Standards This section describes the relationship of this document to other standards, whether CDISC or external. This document does not replace the foundational CDISC standards or their implementation guides. The user should read those standards and implementation guides before applying the advice in this user guide. There are multiple types of data that have existing CDISC-based standards that can be used in QT studies without additional development or customization (e.g. demographics, clinical laboratory tests, and pharmacokinetics), and so are not covered in special detail in this document. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 8 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 164 165 166 167 168 169 170 171 172 173 This document uses a domain model, a dataset specification, and two variables which are not final at the time of publication and therefore are subject to change or deletion without formal notice. Please check the most recent version of the SDTM, SDTMIG, and SDTMIG-MD to ascertain their current status. • The ECG QT Correction Data Model (QT) domain is still a work in progress (i.e. a “draft” domain), which has not appeared prior to this TAUG-QT. • The Device-Device Relationships (RELDEV) dataset is a draft relationships dataset, designed for eventual inclusion in the SDTMIG-MD, which has also not appeared prior to this TAUG-QT. • The --REPNUM (Repetition Number) variable is a variable which has been approved by SDTM Governance but has not yet been included in a formally published version of the SDTM. • The EGBEATNO (ECG Beat Number) variable is also an approved but unpublished variable. 174 1.7 Known Issues 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 Location of QT correction coefficients in submission data For studies in which QTc is a major or the primary endpoint, advanced statistical methods may be used for the correction the QT interval. Unlike the straightforward population correction formula such as Fridericia's or Bazette's, these other methods are based on regression modeling techniques and may utilize external data or within-subject data to derive the corrected values. The correction formula produced by these methods, along with the model coefficients, is considered important data to be included in a submission. Such statistical analyses are not always performed by the sponsor and may be done by the same vendor that creates the database of uncorrected, collected measurements. There are different perspectives on whether the output from these statistical methods should be submitted in the SDTM or ADaM folder. There is precedence to consider any data originating from a central lab or other vendor as raw data, regardless of the number or complexity of calculations performed to obtain them, and therefore should be submitted in SDTM. However, clearly QT correction coefficients that are determined via statistical modeling represent derived, analysis-related data, and would be considered data submitted in the ADaM folder. This TAUG-QT v1.0 was developed using the rationale that data that is vendor-supplied should be submitted in SDTM (Section 6.2) and sponsor-derived data should be submitted in ADaM (Section 7.2). It is recognized that this rationale results in variability with respect to the location of these data in submissions. Therefore, this approach may change. SDTM baseline flag It is recognized that the SDTM baseline flag is inadequate for analyses in certain situations, among them crossover studies where each treatment period could have its own baseline. There are ongoing discussions about whether, in these situations, the SDTM baseline flag should be populated since it could be misleading. In the vital signs example in this document (Section 6.4.1), the baseline flag VSBLFL was not populated, and it was noted that this omission would be explained in the Study Data Reviewers Guide. Other approaches are possible, and future implementation advice on a preferred approach may be different from that shown in the example 202 2 ECG Overview 203 2.1 QT/QTc Assessments in Clinical Studies 204 205 206 207 208 209 210 When ventricular repolarization (i.e. the relaxing of the bottom chambers of the heart, which perform the majority of its pumping action) is delayed, it can lead to cardiac arrhythmias which may be fatal. Historically, a number of drugs have been found to cause such a delay, thus, determining whether a drug delays ventricular repolarization is important in assessing its safety. One measure of the time required for ventricular repolarization is the QT interval on an ECG, corrected for heart rate; or QTc. QTc is clearly recognized as an imperfect biomarker for increased risk of fatal arrhythmia because it can be increased by a number of drugs that are not associated with a significant incidence of such arrhythmias. On an individual basis, the increase in QTc generally needs to be substantial to place Page 9 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 the patient at risk. However, for a potential new drug, even a slight mean increase * in QTc can be clinically meaningful, in that some degree of risk cannot be excluded in a small number of individuals in a large population that will receive the drug during its use in clinical medicine. QTc can be assessed in a variety of study types with a variety of analyses: • QTc is one of many safety parameters assessed in early Phase 1 Single Ascending Dose (SAD) and Multiple Ascending Dose (MAD) safety studies. These studies are typically small, with participants that are adult, non-geriatric, healthy, and with normal ECGs at screening. These studies collect and analyze ECG, PK, and PD data from multiple time points both before and after study treatment. • QTc is also one of the multiple safety parameters that might be assessed in Phase 2–4 clinical trials. Such trials collect baseline ECGs and then collect additional ECGs at one or more time points subsequent to initiation of study treatments. Analyses generally consist of comparing change from baseline for active drug with the comparable change for placebo and/or active control and also comparing the incidences of values exceeding absolute upper reference limits (e.g., values above 500 ms regardless of magnitude of change) and positive change from baseline reference limits (e.g., increases from baseline ≥60 ms regardless of absolute value). • The TQT study has QTc as its primary outcome of interest. Data collected in these types of studies are discussed in this document. Sometimes QTc is an incorrect or insufficient measure: • When the drug under study causes substantial changes in autonomic tone, this can cause significant changes in heart rate, complicate the correction of QT for heart rate, and even alter the relationship between QT and heart rate. This circumstance is the primary one of those under which a conventional TQT might not be appropriate and alternative/complementary methods and/or analyses are required. A White Paper has recognized five such alternative/complementary methods and/or analyses 4. • Alternatives to the TQT study of QTc as assessment of the potential for a drug to cause fatal arrhythmias have been published 5. These alternatives may predict the potential for a drug to induce a fatal cardiac arrhythmia or, conversely, be able to demonstrate absence of such a liability. Furthermore, some of these proposals have suggested using morphological aspects of the T wave that can be quantitated (e.g., ascending and descending slopes height (voltage)) as primary outcomes of interest 6. • Several groups of authors have described a set of properties of a drug (i.e., triangulation, reverse use dependence, instability, and dispersion; or TRIaD) that relate to the depolarization and repolarization, or action potential (AP), of the ventricle, which they believe are more relevant to prediction of potential for induction of a fatal arrhythmia than is prolongation of the QTc obtained from surface ECGs in human subjects 7. Detailed discussion of these topics is out of scope for this document. 2.2 Explanation of Common Terms The terms discussed below have very specific connotations within the context of this document. Familiarity with this terminology should improve the understanding of the content. Common Term Cardiologist over-read ECG Central ECG laboratory * Explanation An ECG for which a cardiologist, using ECG processing hardware and software, makes a final determination of ECG quantitative data (interval measurements) and an assessment of ECG qualitative findings (abnormalities of axis, rhythm, conduction, evidence of ischemia, etc.). See also Over-read (ECG). A commercial facility (some pharmaceutical companies maintain internal central ECG laboratories) that provides a broad range of services, globally, supporting the acquisition of ECGs during clinical studies, including project management, ECG management, data management, ECG over-reading and analyses, and reporting of >5 milliseconds (msec) would be considered to exceed random variability 19 and a mean increase of ≥10 msec could be clinically significant. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 10 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Common Term Clinical study ECG, EKG ECG machine Continuous, high-fidelity, 12-lead ECG recording HR Machine-read ECG Over-read (ECG) QT QTc RR Interval Page 11 Draft Explanation ECG results. This term is synonymous with ECG core laboratory, vendor, central vendor, and core lab. A study (also known as a trial) conducted in human subjects according to procedures and methods described in a written protocol. Electrocardiogram. A graphical recording (also termed a tracing) of the electrical activity of the heart, conventionally recorded from electrodes placed on the body surface (surface electrodes). The standard ECG records and displays 12 wave forms referred to as leads. Equipment designed to record and graphically display signals from the electrical activity of the heart as detected from electrodes placed on the body surface (ECGs). This equipment is also known as an ECG cart or an ECG acquisition device. ECG machines used for most clinical studies must record high-fidelity (1000 Hz) digital data for 10-second recording intervals and provide automated machine-read measurements of quantitative ECG data. Equipment that continuously records the electrocardiogram for up to 24 hours (or longer, depending on memory capacity of the equipment being used) for all 12 conventional ECG leads with the same digital quality as electrocardiograms recorded by a standard, 10-sec, 12-lead ECG machine described immediately above. Conventional 10-sec ECGs can be extracted from continuous recordings. Sometimes referred to as a Holter recording; however, the original Holter device recorded at a lower fidelity and with fewer leads. Heart rate. The measurement of the number of electrical depolarizations of cardiac muscle, usually leading to contraction of the heart muscle, and under normal circumstances, resulting in pumping of blood into the arterial system. The rates of these electrical depolarizations are often referred to as cycles per minute (cpm) but can be referred to as heart beats per minute (bpm). HR is assessed on an ECG tracing. HR is not equivalent to pulse rate because pulse rate is obtained by palpation of blood being pumped through the arteries by effective contraction of the heart muscle. Under normal circumstances, pulse rate will be the same numerically as HR. However, with certain abnormalities, these measurements will be numerically different. Pulse rate is always referred to as bpm. Automated measurements of ECGs generated by ECG machines (ECG cart, ECG acquisition device). The corrected QT (QTc) is generally reported with Bazett’s correction (QTcB) for HR. This information is useful for patient management but may not be recommended for data analyses that will be used for definitive internal decision making or for regulatory submission. Some machines can report Fridericia’s correction (QTcF) or a proprietary correction. ECG machines also provide automated measurements of heart rate, PR interval, QRS width and axis (QRS vector measured in degrees), and other quantitative parameters as well as reporting abnormal qualitative findings The process of measuring and interpreting the waveforms recorded on each ECG as related to the electrical conductivity and functioning of the heart. Methods for overreading may be: 1) fully manual by cardiologist or medical professional (ECG technician or non-cardiologist physician); 2) computer assisted by cardiologist or medical professional; or 3) fully automated (computerized interpretation of ECGs). QT interval. The portion of an ECG between the onset of the QRS complex and the end of the T-wave, representing the total time for ventricular depolarization and repolarization. HR-corrected QT interval. The QT interval is inversely related to HR. When the QT interval is corrected for HR by use of various formulas, it is expressed as QTc and allows an assessment of the QT interval that is intended to be independent of HR. The time between 2 consecutive heart beats/cycles (P-QRS-T complexes), measured as the time between the peaks of 2 consecutive R waves. The RR interval is the time © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Common Term Subject TQT study Explanation between individual heart beats and is related to HR in that HR is essentially the number of R waves in 1 minute. In most QT correction formulas the RR interval is used for correction. An individual under study. Subjects may be normal healthy individuals or patients with a particular disease or condition. A clinical study focused on the investigation of possible study drug effects on ventricular repolarization as evidenced by changes in the QTc of the ECG following the ICH E14 guidelines 1. The primary statistical analysis is a noninferiority comparison of study drug to placebo. That is to say, the goal of the study is to prove the absence of a drug-prolonging effect on QTc within a predefined upper bound on the confidence interval about the difference between drug and placebo that will be accepted as demonstrating no prolongation. 253 2.3 The Fundamentals of ECGs 254 2.3.1 What is an ECG and What Information Does it Provide? 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 An electrocardiogram (ECG, or EKG) is a graphical recording, or tracing, of the electrical activity of the heart recorded at the body surface and depicted as a series of waveforms that illustrate voltage over time. A 12-lead ECG is useful as an independent marker of cardiac disease and reflects abnormalities of the heart that can arise from anatomic, hemodynamic, molecular, ionic, and/or drug-induced causes. The heart is a specialized muscle consisting of four chambers: the right and left atria (or receiving chambers) and the right and left ventricles (or discharging chambers). The right atrium receives blood from the body, the right ventricle discharges blood to the lungs, the left atrium receives blood from the lungs, and the left ventricle discharges blood to the body. The cardiac cycle consists of a period of relaxation called diastole, during which the heart fills with blood, followed by a period of contraction called systole. Alternating contraction and relaxation of the muscle produces the mechanical pumping action of the heart that carries blood around the circulatory system. Atrial contraction precedes ventricular contraction by a short time, which is important for the pumping’s effectiveness. The contraction of the heart is initiated by small amounts of electrical activity in the heart muscle cells. Certain muscle cells are organized into a specialized conduction system, much as a set of wires would carry electrical impulses in a computer (Figure 3 and Figure 4). For the heart to function properly as a pump, this electrical activity must be accurately timed and coordinated in the various parts of the heart muscle in order for contraction and relaxation to occur properly. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 12 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 274 275 276 277 278 279 280 281 Sinoatrial node Atrioventricular node Bundle of His Left bundle branch Left posterior fascicle Left-anterior fascicle Left ventricle Ventricular septum Right ventricle Right bundle branch Figure 3: Diagram of the Heart’s Conduction System † As illustrated in more detail in Figure 4 below, electrical activity (depolarization and repolarization) in differing components of the specialized conduction system are responsible for different components of the surface ECG waveforms. In Figure 4, the action potentials are the electrical activity recorded from specific local areas in the conduction system. These multiple action potentials give rise to the surface ECG waveform at the bottom of Figure. The specific contribution of the action potentials to the surface wave form are color coded. † Heuser J, based on Lynch PJ, Jaffe CC. CC-BY 2.5. Available at: http://commons.wikimedia.org/wiki/File:RLS_12blauLeg.png. Page 13 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Abbreviations: SA - sinoatrial AV - atrioventricular 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 Figure 4: Sequence of Heart Excitation and the Associated ECG Waveforms ‡ ECG waveforms include single electrophysiological events and combinations of electrophysiological events (Figure 4 and Figure 5; see also Figure 7). Figure 5 illustrates an ECG waveform complex and the measurement points for its various components and intervals. The first part of an ECG waveform is called the P-wave and represents atrial depolarization (also called activation; electrical activity in heart muscle cells that causes muscular contraction). There is then a slight time delay before the next ECG waveform component, the QRS complex, which represents ventricle depolarization. Another slight time delay occurs before the T-wave, which represents ventricular repolarization (or recovery). The U-wave may not always be observed. There is no ECG component representing atrial repolarization because this occurs at the same time as ventricular depolarization. An ECG waveform is captured by an ECG machine and recorded on 1-millimeter (mm) grid paper, with the Y-axis representing changes in electrical voltage and the X-axis representing change over time. When the ECG machine is set to a paper speed of 25 mm/second, each mm on the X-axis represents 40 milliseconds (msec). Each mm on the Y-axis customarily represents 0.1 microvolt (mV). ‡ Public domain. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 14 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) A b b r 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 Figure 5: Electrocardiogram Waveform Illustration Electrocardiogram recorded on grid paper with lines 1mm apart. X-axis is the time axis. Y-axis is the voltage axis. Figure assumes a paper speed of 25mm/sec and a calibration of 10mm/mV. Waveform components: • PR interval − time from onset of atrial activation to onset of ventricular activation; includes P-wave and PR segment. The PR interval actually ends at the beginning of the QRS complex and not the peak of the Rwave. o P-wave − atrial depolarization o PR segment − atrial ventricular (AV) nodal depolarization • QT interval − duration of ventricular activation and recovery; includes the QRS, ST segment, and T-wave o QRS complex − ventricular depolarization; obscures atrial repolarization Q-wave − the first negative wave in QRS R-wave − the first positive wave in QRS S-wave − the first negative wave following the R-wave o ST segment − delay between ventricular depolarization and repolarization; all ventricular cells are depolarized o T-wave − ventricular repolarization • Corrected QT interval (QTc) − QT interval corrected for heart rate (HR) • U-wave − Purkinje fiber repolarization or possibly represents “after depolarizations” in the ventricles caused by delayed repolarization of the ventricle • RR interval − ventricular rate; time between cycles of depolarization/repolarization of the entire heart HR represents the number of times the heart beats per minute. On an ECG, HR usually implies ventricular rate, which means the number of QRS complexes in a minute. Normally, each QRS is preceded by a P-wave; hence, HR also refers to atrial rate (the number of P-waves in a minute). There are different methods to calculate HR on an ECG. The most widely used ECG machine algorithm derives the median elapsed time from the peaks of the R-waves on the dominant beat type within the 10-second recording and, as HR = 60,000/RR when RR is expressed in msec, derives the HR from the RR. Page 15 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 The PR interval and QRS complex are of interest because they reflect muscle depolarization. Some drugs cause a delay in muscle depolarization that can lead to a lack of coordination of cardiac electrical activity and impairment of cardiac pump function. The QT interval is primarily driven by the ventricular repolarization time, including the delay after depolarization (the ST segment). Ventricular repolarization time is particularly important because if it is too long, ventricular depolarization can begin to occur before repolarization is complete, causing the electrical activity of the heart to become highly rapid and chaotic, resulting in loss of effective cardiac pump function and death. This abnormal electrical activity is referred to as ventricular tachydysrhythmia (also termed ventricular tachyarrhythmia, ventricular tachycardia). Specific types of ventricular tachydysrhythmia can occur, such as Torsades de Pointes, and ventricular fibrillation. The regulation of HR changes is based on the body’s need for energy sources. With respect to the ECG waveform, an increase in HR is equivalent to an increase in the frequency of depolarization and to a decreased time between individual depolarizations as denoted by a shorter RR interval. Changes in the frequency of cardiac depolarization are detected by heart muscle cells and by the brain centers that influence cardiac electrical activity, which then alter the rate of repolarization appropriately to maintain effective cardiac function. As noted above, the time required for ventricular repolarization to occur (using the QT interval as a biomarker) shortens as heart rate increases and lengthens as heart rate decreases to maintain normal physiological function. Therefore, it is necessary to correct QT interval for the HR, by means of the RR interval measurement, in order to meaningfully compare QT intervals that were measured at different HRs. The QT interval corrected for HR is referred to as the HR-corrected QT (QTc) interval. QTc intervals are useful when comparing the QT intervals within an individual from ECGs collected at different times when HRs were different or for comparisons between individuals or groups. Approaches to QT interval correction are covered in Section 2.3.4. Unfortunately, a number of factors can delay ventricular repolarization and thereby put individuals at risk of ventricular tachyarrhythmia and sudden cardiac death. Use of drugs is a common cause of delayed ventricular repolarization, so the effect of a drug on cardiac function must be clearly determined. Towards this end, the ECG QTc is used as the standard method of assessing the potential for a drug to influence cardiac repolarization. ECG assessments are covered in Section 6.1. 2.3.2 ECG Machinery An ECG acquisition device (i.e. ECG machine, also called an ECG cart or cardiograph) records and graphically displays the electrical impulses that stimulate the heart muscle to contract (electrical activity → muscle contraction → heart pumps blood). ECG acquisition devices used in clinical studies capture the heart muscle’s electrical activity from electrodes on the torso and left leg (right and left arm leads are placed on the respective shoulders) that are configured to form various types of leads (Figure 6). Data collected from these electrodes are stored electronically in the ECG machine (Concept Map 1) and are printed graphically as waveforms, which represent the electrical potential difference over time recorded from the ECG leads. Bipolar limb leads (I, II, and III) record the electrical potential differences between two limb electrodes. The unipolar precordial leads record the electrical potential difference between each of six designated sites on the torso and a theoretical zero reference potential site, known as the Wilson central terminal that is composed of the mean of 3 limb leads. Three augmented limb leads (aVR, aVL, and aVF) are modified or augmented unipolar leads. These augmented leads record the electrical potential difference between the limb leads and a different central terminal site composed of the mean of the two other limb leads. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 16 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 Figure 6: Standard Electrode/Lead Configuration for a 12-Lead/Waveform ECG § Precordial Electrodes locations: • • • • • • V1: 4th intercostal space to right of sternum V2: 4th intercostal space to left of sternum V3: Half way between V2 and V4 V4: Left mid-clavicular line in the 5th intercostal space V5: Left anterior axillary line in the 5th intercostal space V6: Left mid-axillary line in the 5th intercostal space Leads – potential differences between Abbreviations: 3 bipolar limb leads: aVR/aVL/aVF = augmented vector right/left/foot; I: R and L arms (L is+) R = right; L = left; V = vector II: R arm and L leg (leg is +) III: L arm and L leg (leg is +) 6 precordial leads (central terminal is average of the 3 limb leads not recorded) V1: V1 (+) and central terminal (R and L arms and L leg) V2: V2 (+) and central terminal (R and L arms and L leg) V3: V3 (+) and central terminal (R and L arms and L leg) V4: V4 (+) and central terminal (R and L arms and L leg) V5: V5 (+) and central terminal (R and L arms and L leg) V6: V6 (+) and central terminal (R and L arms and L leg) 3 augmented unipolar limb leads (central terminal is average of the 2 limb leads not recorded) aVR: R arm (+) and central terminal (L arm and L leg) aVL: L arm (+) and central terminal (R arm and L leg) aVF: L leg (+) and central terminal (R and L arms) On an ECG printout, the name for each lead is associated with its waveform. Each ECG tracing has 12 individual waveforms (resulting from leads) on the standard ECG printout (Figure 7). ECG machines also construct a median beat for each lead from all the beats for that lead, and then superimpose them by aligning the peak of the R-wave for each median beat. This creates a superimposed median beat, which is used for measurement purposes of PR, QRS, and QT. One lead is used to measure multiple RR intervals to obtain HR. § Left image: Public domain. Right image: CC0 1.0. Page 17 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 Figure 7: 12-Lead Electrocardiogram Waveform In addition to waveforms, ECG machines provide automated machine-read interval measurements and qualitative assessments (Concept Map 1). The ECG machine stores the ECG waveforms, interval measurements (annotation marks are placed on the appropriate lead for RR and on the superimposed median beat for other intervals for automated measurement purposes, but these are not displayed on the ECG print out) and qualitative assessments for electronic transmission to a server that is housed at a centralized ECG vendor. The server at the vendor (Concept Map 2) can be used for human over-read of the ECG that consists of human adjustment of the annotation lines for interval measurements and correction of any qualitative assessment. The human over-reader can adjust the measurements made on the superimposed median beat and make measurements in any individual lead. The server and its software allow high precision visualization of the waveforms by the human over-reader. If human over-read is not performed, the automated measurements are transmitted to the sponsor for analysis. In collecting and processing ECG data, moving from the subject to a dataset for analysis, there are generally two distinct hardware-software devices that are employed. These devices are generally linked as a pair. Figure 8 is a high-level representation of these devices, with more detailed representation in Concept Map 1 and Concept Map 2. These concept maps also describe some of the functions of the two respective devices in going from subject to data output for analysis. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 18 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 432 433 434 435 436 437 438 439 440 441 442 443 444 445 Figure 8: High-Level View of ECG Hardware-Software: From Subject’s Chest to Data for Analysis Some vendors have their own proprietary hardware-software (a third device) for further processing of the digital waveform information that is ultimately reported to the sponsor for analysis. In this case, the server for the ECG machine is simply used as a transmission device for the data received from the ECG machine to the vendor’s proprietary device. Concept Map 1: Recording of ECG Digital Waveforms The above concept map shows the process of capturing analog electrical potential differences, converting to digital waveforms, and the automated measurement of intervals as well as morphological interpretation. Page 19 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 446 447 448 449 450 451 452 453 454 This concept map displays the process for automated measurement of intervals, based on a single lead or a superimposed median beat and RR, with potential human over-reading/adjustments. Relationships involved in optional human over-read are shown using blue arrows. Furthermore, this concept map represents the server at the central vendor (second machine). If the central vendor uses a proprietary hardware-software device to generate final measurements to supply to the sponsor, this machine would simply receive the digital information and transmit it to the proprietary third device. 455 2.3.3 ECG Device Types Concept Map 2: Measurement of RR and Intervals for Use in Analysis 456 457 458 459 460 461 462 463 In a multisite study, there would be ECG equipment at each site. Even at a single site, there may be multiple ECG machines used. However, in a QT study, the specifications of the ECG equipment would be stated in the protocol, and all individual devices used would have the same characteristics. The SDTM Implementation Guide for Medical Devices (SDTMIG-MD) allows a single sponsor identifier to be assigned to a group of devices that share the same characteristics, and this is how ECG devices would be handled in QT studies. 464 2.3.3.1 Examples for Device Types 465 466 467 468 469 470 471 472 473 474 475 476 477 478 Example 1: This example shows the types of devices used to obtain ECG data for a particular study. Although multiple ECG machines were used, they were all of the same type, so identifiers were assigned to the type of machine, rather than to individual devices. There were two separate devices used to obtain the ECG data: an ECG machine that recorded data from the subject, and a second device that performed further processing of the ECG data from the first machine, and also allowed manual over-reading of the ECG data. Because these two machines were used together to produce the results in the ECG domain, the combination of the two devices is treated as a composite device, and the combination is given its own unique sponsor ID. Data about these devices is represented in the Device Identifier (DI) and Device Properties (DO) domains and in the proposed Device-Device Relationships dataset. For the ECG machine, the device that collects data from the subject, and the device that does further processing and allows manual over-reading, the characteristics of interest are manufacturer, model, and software/firmware version. Row 1: Rows 2-4: Shows the sponsor device identifier for the composite ECG Device. This is the device whose identifier appears in the EG dataset where subject data are represented. Further information about what this device consisted of is provided in the Device-Device Relationships dataset, below. Show the characteristics of the ECG machine, which has been given an SPDEVID value of "2.” © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 20 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 479 480 481 482 Rows 5-7: Show the characteristics of the machine which performs further processing of the ECG data from the first machine. This device has been given an SPDEVID value of “3.” di.xpt Row STUDYID DOMAIN SPDEVID DISEQ DIPARMCD DIPARM DIVAL ABC-123 DI 1 1 TYPE Device Type Composite ECG Device 1 ABC-123 DI 2 1 TYPE Device Type ECG Machine 2 ABC-123 DI 2 2 MANUF Manufacturer Acme 3 ABC-123 DI 2 3 MODEL Model XYZ 2000 4 ABC-123 DI 3 1 TYPE Device Type ECG Analyzer 5 ABC-123 DI 3 2 MANUF Manufacturer Acme 6 ABC-123 DI 3 3 MODEL Model ECG Wizard 7 483 484 485 486 487 488 489 Row 1: Row 2: Shows the software/firmware version of the device with sponsor device identifier “2”. The software/firmware version is not an intrinsic part of the device, so it is represented in the DO domain rather than the DI domain. Shows the software/firmware version of the device with sponsor device identifier “3”. do.xpt Row STUDYID DOMAIN SPDEVID DOSEQ DOTESTCD DOTEST DOORES ABC-123 DO 2 4 SFTWRVER Software/Firmware Version 3.6 1 ABC-123 DO 3 5 SFTWRVER Software/Firmware Version 12B 2 490 491 492 493 494 495 496 497 498 499 500 501 The relationships between the two ECG machines and their combination is a simple one, which requires no additional parameters to describe the relationship, so the permissible variables PARMCD, PARM, and VAL are not included in the Device-Device Relationship dataset. Row 1: Row 2: Row 3: Shows that the device with SPDEVID = 1 has no parent and thus is at Level 1. Shows that the device with SPDEVID = 2 is a part of the composite device with SPDEVID = 1. Since the device with SPDEVID = 1 is at Level 1, this device is at Level 2. Shows that the device with SPDEVID = 3 is also a part of the composite device with SPDEVID = 1 and thus is also at Level 2. reldev.xpt Row STUDYID SPDEVID PARENT LEVEL ABC 1 1 1 ABC 2 1 2 2 ABC 3 1 2 3 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 2.3.4 How is the QT Interval Adjusted (Corrected) for Heart Rate? There are three basic methods to adjust or correct the QT interval for HR. Essentially, all the methods attempt to adjust the individual subject’s QT interval to a value that would be expected if the subject’s HR were constant. The methods are: 1. Population-based formula derived from a historical population 2. Population-based formula derived from the population under study 3. Individual-based formula derived for each individual in the population under study The first two methods are based on finding the mathematical formula (linear or nonlinear) that best describes the relationship between the absolute QT interval and the RR interval from populations of multiple individuals and applying the same formula to all ECGs for which a QTc is being computed. 1. Population-Based Formula from a Historical Population In a historical population, this would be a group of normal, healthy volunteers, generally with 1 ECG from each subject. Due to the normal variance between different populations, multiple researchers using different groups of subjects have derived different formulas. Page 21 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 520 521 The most well-known and clinically used is Bazett’s formula (QTcB), which was derived/developed in 1920 from ECGs recorded in a small group of healthy subjects: QTc = 522 523 √RR Fridericia’s formula (QTcF) was also developed in 1920 from ECGs recorded in a small group of healthy subjects: QTc = 524 525 526 527 528 529 530 531 532 533 QT QT 3 √RR It is reasonably well known that the Bazett formula under-corrects at faster heart rates (over 60 bpm) and conversely over-corrects at slower heart rates. That is, at faster HRs (smaller RR intervals), the computed QTc is ‘larger than it should be’ and at slower HRs (larger RR intervals), the computed QTc is ‘smaller than it should be’. When Bazett corrected QTc is plotted against RR interval and a regression line is plotted, the slope is negative (Figure 9; with a perfect correction, the slope of the regression line would be 0). In spite of this, Bazett’s formula is still the most widely used for clinical correction of QT intervals. However, it is becoming more acceptable in regulatory documents to use the Fridericia formula correction, without use of the Bazett formula, 2, 8 along with additional correction results as described below. Fridericia correction Corrected QT interval (msec) Corrected QT interval (msec) Bazett correction 500 450 400 350 450 400 350 0.6 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 500 0.8 1.0 1.2 0.6 RR interval (sec) 0.8 1.0 1.2 RR interval (sec) Figure 9: Relationship between the Bazett- and Fridericia-Corrected QT Interval and RR Interval 2. Population-Based Formula from the Population under Study A population formula derived from the population under study uses off-treatment, baseline ECGs to construct a population formula in a manner similar to construction of a formula from a historic control population. Because the formula is based on the behavior of the individuals actually under study, such a study populationderived formula presumably accounts for variables (e.g., disease factors, age, and gender distribution) which might influence the QT-RR relationship. Therefore, such a formula should be more accurate for the individuals under study than one based on a historical population. 3. Individual-Based Formula (QTcI) An individual-based QTc requires that a number of ECGs be obtained across a sufficient range of HRs (35 to 50 ECGs covering a range of heart rates of 50 to 80 beats per minute difference 9) for each individual under baseline (nontreatment) conditions. One author has published data to support the position that at least 400 ECGs (QT-RR pairs for each individual subject) are needed to compute an adequate individual correction and that there must definitely be a range of heart rates corresponding to the heart rates that will be observed on the experimental drug 10. These data are then used to compute a specific correction formula for each individual subject in a manner similar to that used to compute a population correction. The rationale for such a method is based on the experimental findings that the QT-RR relationship is different across subjects but relatively stable within subjects. In using QTcI, one subapproach is to use a single, predetermined mathematical model for all subjects. An alternative sub-approach is to fit © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 22 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 the individual subject’s data to several preselected mathematical models and use the best mathematical model for each individual subject (model that results in flattest regression line after correction (QTcI vs. RR)) As such, this QTc method is probably the best available, but is also very labor intensive and costly to use. For example, a study with only 60 subjects would require, at a minimum, approximately 2100 baseline ECGs (60 subjects × 35 ECGs/subject) be collected and measured to compute the 60 individual correction formulae. If 400 ECGs are required per subject then 24,000 ECGs are required and this would require acquiring the ECGs from continuous ECG recording. The 60 individual correction formulae would need to be derived in addition to collecting the actual experimental ECGs. ECGs acquired for baseline values could be used for this purpose. Some researchers have developed methods of assessing changes in ventricular repolarization based on the QT interval that do not rely on an explicit correction of the QT interval for heart rate (the RR interval). One such method is known as the beat-to-beat method 4. These methods are particularly important when the experimental drug results in marked changes in autonomic nervous system tone and heart rate. These changes can be so large that it will be difficult to obtain ECG data at heart rates that will be observed during treatment with the experimental drug, which would raise concerns about the validity of any correction factor. Discussion of these alternatives beyond the introduction of the concept is outside the scope of this document. These methods would generally rely on continuous recording data. Section 6.2 provides further details on QT correction from the dataset perspective. 3 The Thorough QT (TQT) Study This section is based primarily upon the May 2005 ICH-E14 document 1, which describes the basic conduct, purpose, and expected analyses of the TQT study. The purpose of a TQT study is to evaluate the potential for an experimental drug to delay cardiac ventricular repolarization, which it does through evaluation of changes in the heart rate corrected QT interval (QTc) during drug treatment; and also to demonstrate that the study is capable of detecting differences of clinical significance (approximately 5ms), so as to confirm that any lack of detected change is due to actual lack of change rather than lack of assay sensitivity. These TQT studies are generally conducted in healthy volunteers highly screened for normal cardiac electrical activity, for ease of precise measurement of the QT interval and to avoid additional confounding factors. The TQT study can be a crossover design when sufficient experimental drug exposure can be achieved with a single dose or a few doses, or it can be a parallel design when a longer treatment period (several weeks or more) is required to achieve sufficient experimental drug and/or metabolite exposure (i.e. experimental drug and/or metabolites with long half-lives). If the study is a crossover design, the order of the treatments should be balanced across subjects, such as with a Williams design 11. In general, the treatments are: 1. A dose of the experimental drug that is many times higher, if possible, than the intended maximum dose, in order to account for drug-drug interactions and/or genetic metabolic enzyme deficiencies that might lead to greater exposure to the experimental drug than otherwise intended with a given dose during routine clinical use 2. Placebo 3. A positive control for purpose of demonstration of assay sensitivity (most often moxifloxacin, usually oral but sometimes intravenous) 4. Optionally, a dose of the experimental drug that is within the intended therapeutic range (generally the maximum intended therapeutic dose) The administration of the active control has been allowed by regulators to be open-label, but the administrations of the experimental drug dose(s) and placebo are double-blind, and ECG measurements and readings are performed by persons completely blinded to associated treatments, subject details, and date/time of the ECG. ECGs are usually collected at one or more baseline time points: immediately before experimental drug administration, and/or on the day or days preceding day of first treatment administration at time points corresponding to those at which ECGs will be collected following treatment. In crossover studies, these baseline Page 23 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 ECGs are usually collected before each treatment period as opposed to using a single baseline for all treatment periods. Post-treatment time points are determined by the pharmacokinetic properties of the treatment. The specific time points at which ECGs would be acquired correspond to and surround the expected time of maximum concentration of parent experimental drug and relevant metabolites. ECGs are collected as a set of replicates (in close temporal proximity, e.g., 3 ECGs collected at 1-minute intervals) of approximately 10 seconds in duration and utilizing all 12 leads. ECGs can be collected as conventional ECGs or they can be extracted from a continuous high-fidelity ECG recording. Experimental drug and metabolite concentrations are often assayed immediately after the time of ECG collection for PK/PD analysis, which can be a useful secondary analysis pertinent to the potential influence of the experimental drug on ventricular repolarization. An experimental drug is declared to have no influence on QTc if: 1. The comparison of the supratherapeutic dose of the experimental drug versus placebo is a noninferiority comparison and the upper one-sided 95% confidence interval for the maximum difference (across the several time-matched time points of ECG acquisition following treatment administration) between change from baseline (there are multiple potential definitions of baselines discussed in a separate document) for experimental drug compared to placebo is less than 10 ms (with no adjustment for the multiple comparisons across the multiple time points) 1 AND 2. The positive control (e.g., moxifloxacin) versus placebo results in at least one difference from baseline where the lower one-sided 95% confidence interval for the difference is greater than 5 ms (must adjust for the multiple comparisons across time points) 8. Note that this explicit statistical definition requires not only the ability to detect a difference between the positive control and placebo but also a certain minimum difference, based on historical experience with moxifloxacin. Without passing the test on assay sensitivity, the experimental drug is unlikely to be declared to have no effect, regardless of the results of the first statistical co-primary test. Unfortunately, across multiple TQT studies, oral moxifloxacin, while producing a detectable increase in QTc, sometimes fails to result in an increase of sufficient magnitude such that the one-sided 95% confidence interval for the difference between moxifloxacin and placebo is greater than 5 ms after adjusting for multiple comparisons. This lack of sufficient difference might occur for a variety of reasons. As discussed below, PK/PD modeling of moxifloxacin concentration – change in QTc can serve to support assay sensitivity when exposure to moxifloxacin (concentrations) are less than expected due to some identified reason. The ICH-E14 document has specified some secondary, conventional difference (not noninferiority) analyses of the QTc. These are categorical (i.e., outlier, extreme observation) analyses (on the average for the replicate sets) based on both absolute values and changes from baseline: • Incidences of QTc >450 ms, experimental drug vs. placebo • Incidences of QTc >480 ms, experimental drug vs. placebo • Incidences of QTc >500 ms, experimental drug vs. placebo • Incidences of increase from baseline >30 ms, experimental drug vs. placebo • Incidences of increase from baseline >60 ms, experimental drug vs. placebo Although not explicitly specified in detail in ICH-E14, both central tendency (i.e., mean change) as well as categorical (i.e., outlier, extreme observation) analyses for experimental drug vs. placebo of the following are often expected by regulators: • Heart rate / RR interval • PR interval • QRS • Incidence of treatment-emergent abnormal morphological (i.e., qualitative) findings: rhythm, axis, conduction, hypertrophy, ischemia-injury-infarction, ST segment abnormalities, T wave abnormalities, U wave abnormalities, etc. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 24 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 Some treatment-emergent adverse events which suggest the occurrence of delays in ventricular repolarization might not be well accounted for by the standard, objective study analyses. Additional analyses of such adverse events are also important. Such events are likely to be of low frequency in a healthy population. In addition, as noted above, the time course of change in QTc displayed by the positive control is of interest. It is expected to rise to a maximum associated with time of maximum concentration and then decline to baseline consistent with kinetics of elimination. If the observed time course of change in QTc does not parallel change in concentration, assay sensitivity can be questioned. Sponsors will often model changes in QTc (difference between drug and placebo in changes in QTc) versus changes in concentration for the experimental drug, its metabolites, and separately moxifloxacin with linear regression or other appropriate model. If the experimental drug is associated with increases in QTc, this modeling can be helpful in defining the expected magnitude of such an increase at maximum expected exposure. If the positive control results in less increase in QTc than expected, this modeling can be helpful in explaining the results and potentially demonstrating that the positive control behaved similarly to historical expectations when exposure (maximum concentrations) was less than expected based on historical experience. Detailed advice on the specifics of these PK/PD analyses are beyond the scope of this introductory description of TQT study basics. When an experimental drug causes substantial changes in heart rate, the TQT study paradigm described above may not be appropriate because it may be impossible with conventional methods described above to adequately correct the QT interval for heart rate. In such situations, alternative study and/or analytical methodologies may be necessary 4 . Similarly, when the experimental drug is potentially toxic and/or has a narrow therapeutic range, special study adaptations may be necessary. Page 25 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 685 4 Trial Design 686 4.1 TQT Study Design 687 688 689 690 691 692 693 694 695 696 As mentioned in Section 2.1, QT interval and QTc might be an outcome of interest evaluated in a number of other types of studies such as Phase 1 single ascending dose (SAD) and multiple ascending dose (MAD) studies, Phase 3 and Phase 4 studies, and TQT studies with modifications dictated by toxicity considerations relevant to the test drug. In general, these other study types would be of the parallel or crossover design but may have differences in study elements such as study population, inclusion/exclusion criteria, and specific treatments included. 697 4.1.1 Parallel Studies 698 699 700 701 702 703 704 705 706 707 708 The example study designs presented below illustrate specific TQT study designs. In a TQT study, QT interval and QTc are the primary outcomes of interest (see Section 3). A typical TQT study is designed as double-blind (partial double-blind as in some cases the investigator might not be blind to administration of the active control), placebo- and positive-controlled to determine whether the test treatment fails to prolong QTc and QT intervals (primary statistical test is noninferiority) and to demonstrate the assay sensitivity of the positive control treatment in the study population. Traditional TQT studies employ parallel or crossover designs and are often designed with equal study duration and sample size for the different treatment arms or periods 12. Under certain circumstances (related to the PK characteristics of the test drug), a parallel design may be preferred for a TQT study. Such circumstances include 1: • Drugs with long elimination half-lives for which lengthy time intervals would be required to achieve steady-state and complete washout. • If carryover effects are prominent for other reasons, such as irreversible receptor binding or long-lived active metabolites. • If multiple doses of the investigational drug are required to evaluate the effect on QT/QTc. Example TQT Study 1 Below is the study schema diagram for Example TQT Study 1, a parallel trial. This trial has 4 treatment arms (placebo, positive control, therapeutic study drug dose, and supratherapeutic study drug dose), which correspond to the 4 possible left-to-right "paths" through the trial. Moxifloxacin has become the standard positive control with a well characterized (peak effect and time course), expected influence on QTc in healthy subjects with a mean increase in QTc in the range of 10 – 15 msec. Other positive control compounds are possible (e.g., low dose ibutalide). Note: Moxifloxacin is one example of a positive control. Note: This is an optional arm. 709 710 711 Randomization Figure 10: Parallel Study Design Schema for Example TQT Study 1 T = Therapeutic Dose (DRUG A 1 MG), ST = Supratherapeutic Dose (DRUG A 100 MG) © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 26 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 712 713 714 715 716 717 718 719 Below is an example of a trial design matrix for Example TQT Study 1. A trial design matrix is a table with a row for each treatment arm in the trial and a column for each epoch in the trial. The cells in the matrix represent the study cells, which are populated with trial elements. In this trial, each study cell contains exactly one element. Note that randomization is not represented in the trial design matrix. Trial Design Matrix for Example TQT Study 1 Assume the study’s therapeutic and supratherapeutic doses for Drug A are 1 mg and 100 mg, respectively. Placebo Moxifloxacin Drug A 1 MG Drug A 100 MG 720 721 722 723 724 725 726 727 728 729 730 731 Screen Screen Screen Screen Screen Treatment PLACEBO MOXIFLOXACIN DRUG A 1 MG DRUG A 100 MG Below is an example of a trial arms dataset for Example TQT Study 1. For this example, the conversion of the trial design matrix into the trial arms dataset is straightforward. For each cell of the matrix, there is a record in the trial arms dataset. ARM, EPOCH, and ELEMENT can be populated directly from the matrix. TAETORD acts as a sequence number for the elements within a treatment arm, so it can be populated by counting across the cells in the matrix. The randomization information, which is not represented in the trial design matrix, is held in TABRANCH in the trial arms dataset. TABRANCH is populated only if there is a branch at the end of an element for the treatment arm. When TABRANCH is populated, it describes how the decision at the branch point would result in a subject being in this treatment arm. Trial Arms Dataset for Example TQT Study 1 Assume the study’s therapeutic and supratherapeutic doses for Drug A are 1 mg and 100 mg, respectively. ta.xpt Row STUDYID DOMAIN ARMCD ARM TAETORD ETCD ELEMENT TABRANCH TATRANS EPOCH EX1 TA PBO Placebo 1 SCRN Screen Randomized to Placebo Screen 1 EX1 TA PBO Placebo 2 PBO Placebo Treatment 2 EX1 TA MOXI Moxifloxacin 1 SCRN Screen Randomized to Moxifloxacin Screen 3 EX1 TA MOXI Moxifloxacin 2 MOXI Moxifloxacin Treatment 4 EX1 TA D1MG Drug A 1mg 1 SCRN Screen Randomized to Drug A 1 mg Screen 5 EX1 TA D1MG Drug A 1mg 2 DT Drug Therapeutic Dose Treatment 6 EX1 TA D100MG Drug A 100mg 1 SCRN Screen Randomized to Drug A 100 mg Screen 7 EX1 TA D100MG Drug A 100 mg 2 DST Drug Supratherapeutic Dose Treatment 8 732 733 4.1.2 Crossover Studies 734 735 736 In comparison to parallel group studies, crossover studies have at least two potential advantages 1: • A smaller number of subjects are typically required. Subjects serve as their own controls, resulting in reduced variability of differences related to intersubject variability. Page 27 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 • Heart rate correction approaches based on individual subject data may be more feasible. Since baseline ECGs are collected before each treatment period, more ECGs are available for each subject for computation. Example TQT Study 2 This is an example of a Williams Design used for a typical crossover study. The two most common ways this design could be represented are shown in Figure 11 and Figure 12. How the design is described in the protocol and how the sponsor intends to analyze the data will determine which representation a sponsor chooses. A washout period may be “built into” a treatment Element (Figure 11) if observations occurring during the washout portion will be analyzed as part of the treatment. This representation assumes there is a sufficient period at the end of each Element for the drug to be eliminated from the subjects before the next drug is administered. If data from the time periods where dosing does not occur is not intended to be associated with the previous treatment, then separate washout Elements would be created (Figure 12). Randomization Figure 11: Crossover Study Design for Example TQT Study 2 with Washout Period Combined with each Treatment Element T = Therapeutic Dose (DRUG A 1 mg), ST = Supratherapeutic Dose (DRUG A 100 mg) Randomization Figure 12: Crossover Study Design for Example TQT Study 2 with Separate Elements for each Washout Period T = Therapeutic Dose (DRUG A 1 mg), ST = Supratherapeutic Dose (DRUG A 100 mg), WO = Washout The trial design matrix for the crossover example trial is shown below. Trial Design Matrix for Example TQT Study 2 Assume the study’s therapeutic and supratherapeutic doses for Drug A are 1 mg and 100 mg, respectively. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 28 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Treatment Sequence Screen First Treatment Washout Second Treatment Washout Third Treatment Washout Fourth Treatment PBO-D100-MOXI-D1 MOXI-PBO-D1-D100 D1-MOXI-D100-PBO D100-D1-PBO-MOXI 762 763 764 765 766 767 Screen Screen Screen Screen Placebo Moxifloxacin Drug A 1 mg Drug A 100 mg Washout Washout Washout Washout Drug A 100mg Placebo Moxifloxacin Drug A 1mg Washout Washout Washout Washout Moxifloxacin Drug A 1mg Drug A 100 mg Placebo Drug A 1mg Drug A 100mg Placebo Moxifloxacin Below is an example of a trial arms dataset for Example TQT Study 2. For this example, it is straightforward to produce the trial arms dataset for this crossover trial from the diagram showing treatment arms and epochs or from the trial design matrix. Trial Arms Dataset for Example TQT Study 2 ta.xpt Row STUDYID DOMAIN 1 EX2 TA 2 EX2 TA 3 EX2 TA 4 EX2 TA 5 EX2 TA 6 EX2 TA 7 EX2 TA 8 EX2 TA 9 EX2 TA 10 EX2 TA … … … ARMCD PBO-D100MGMOXI-D1MG PBO-D100MGMOXI-D1MG PBO-D100MGMOXI-D1MG PBO-D100MGMOXI-D1MG PBO-D100MGMOXI-D1MG MOXI-PBO-D1MG –D100MG MOXI-PBO-D1MG– D100MG MOXI-PBO-D1MG– D100MG MOXI-PBO-D1MG– D100MG MOXI-PBO-D1MG– D100MG … ARM Placebo-Drug A 100MGMoxifloxacin-Drug A 1MG Placebo-Drug A 100MGMoxifloxacin-Drug A 1MG Placebo-Drug A 100MGMoxifloxacin-Drug A 1MG Placebo-Drug A 100MGMoxifloxacin-Drug A 1MG Placebo-Drug A 100MGMoxifloxacin-Drug A 1MG Moxifloxacin-Placebo-Drug A 1MG-Drug A 100MG Moxifloxacin-Placebo-Drug A 1MG-Drug A 100MG Moxifloxacin-Placebo-Drug A 1MG-Drug A 100MG Moxifloxacin-Placebo-Drug A 1MG-Drug A 100MG Moxifloxacin-Placebo-Drug A 1MG-Drug A 100MG … TAETORD ETCD ELEMENT 1 SCRN Screen 2 PBO Placebo 3 DA100MG Drug A 100mg 4 MOXI Moxifloxacin 5 DA1MG Drug A 1mg 1 SCRN Screen 2 MOXI Moxifloxacin 3 PBO Placebo 4 DA1MG Drug A 1 mg 5 DA100MG … … Drug A 100mg … 768 Note: The above could be modeled using separate elements for each washout period and follow-up 769 4.1.3 Trial Elements 770 771 772 773 774 775 776 Washout Washout Washout Washout TABRANCH TATRANS Randomized to Placebo-Drug A 100mgMoxifloxacin-Drug A 1mg EPOCH Screen First Treatment Second Treatment Third Treatment Fourth Treatment Randomized to Moxifloxacin-PlaceboDrug A 1mg-Drug A 100mg … Screen … First Treatment Second Treatment Third Treatment Fourth Treatment … The Trial Elements (TE) dataset contains the definitions of the elements that appear in the Trial Arms (TA) dataset. An element may appear multiple times in the Trial Arms table because it appears either 1) in multiple arms, 2) multiple times within an arm, or 3) both. However, an element will appear only once in the Trial Elements table. Each row in the TE dataset may be thought of as representing a "unique element" in the sense of "unique" used when a case report form template page for a collecting certain type of data is often referred to as "unique page." An element is a building block for creating study cells and an arm is composed of study cells. Or, from another point of view, an arm is composed of elements (i.e. the trial design assigns subjects to arms, which are comprised of a sequence of steps called elements). Trial elements represent an interval of time that serves Page 29 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 777 778 779 780 781 782 783 784 785 786 787 788 a purpose in the trial and are associated with certain activities affecting the subject. A valid element has a name that describes the purpose of the element and includes a description of the activity or event that marks the subject's transition into the element as well as the conditions for leaving the element. Below are the TE datasets for Example TQT Study 1 and Example TQT Study 2, respectively. These trials are assumed to have fixed-duration Elements. The wording in TESTRL is intended to separate the description of the event that starts the Element into the part that would be visible to a blinded participant in the trial (e.g., "First dose of a treatment Epoch") from the part that is revealed when the study is unblinded (e.g., "where dose is 5 mg"). Care must be taken in choosing these descriptions to be sure that they are "Arm and Epoch neutral." For instance, in a crossover trial such as Example TQT Study 2, where an Element may appear in one of multiple Epochs, the wording must be appropriate for all the possible Epochs. The wording for Example TQT Study 2 uses the wording "a treatment Epoch." Trial Elements Dataset for Example TQT Study 1 te.xpt Row STUDYID DOMAIN ETCD ELEMENT TESTRL EX1 TE SCRN Screen Informed consent 1 EX1 TE PBO Placebo First dose of study drug, where drug is placebo 2 EX1 TE DST Drug A 100mg First dose of study drug, where drug is Drug A 100mg 3 EX1 TE MOXI Moxifloxacin First dose of study drug, where drug is Moxifloxacin 4 EX1 TE DT Drug A 1 mg First dose of study drug, where drug is Drug A 1mg 5 789 790 791 TEENRL TEDUR 3 weeks after start of element P21D 2 weeks after start of element P14D 2 weeks after start of element P14D 2 weeks after start of element P14D 2 weeks after start of element P14D Trial Elements Dataset for Example TQT Study 2 te.xpt Row STUDYID DOMAIN ETCD ELEMENT TESTRL EX2 TE SCRN Screen Informed consent 1 EX2 TE PBO Placebo First dose of treatment epoch, where drug is placebo 2 EX2 TE DA100MG Drug A 100mg First dose treatment epoch, where drug is Drug A 100mg 3 EX2 TE MOXI Moxifloxacin First dose of treatment epoch, where drug is Moxifloxacin 4 EX2 TE DA1MG Drug A 1mg First dose of treatment epoch, where drug is Drug A 1 mg 5 792 793 794 795 796 TEENRL TEDUR 3 weeks after start of element P21D 2 weeks after start of element P14D 2 weeks after start of element P14D 2 weeks after start of element P14D 2 weeks after start of element P14D 4.1.4 Trial Summary Parameters This section shows a partial trial summary dataset for Example TQT Study 1 described in Section 4.1.3, with Drug T (1 mg) and Drug ST (100 mg) that may be applicable to TQT studies. Note that the example includes data for proposed terminology for new trial summary parameters in rows 44-50. ts.xpt Row STUDYID DOMAIN TSSEQ TSGRPID TSPARMCD TSPARM 1 EX1 TS 1 TITLE Trial Title 2 3 4 5 6 7 8 EX1 EX1 EX1 EX1 EX1 EX1 EX1 TS TS TS TS TS TS TS 1 1 1 1 1 2 1 SPONSOR TPHASE RANDOM TBLIND TCNTRL TCNTRL DESIGN Sponsoring Organization Trial Phase Classification Trial is Randomized Trial Blinding Schema Control Type Control Type Description of Trial Design © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 TSVAL A Randomized, Double-Blind, Placebo- and Positive-Controlled, 4-arm Parallel-Group Study to Evaluate the Effect of Drug A on Cardiac Repolarization in Healthy Adult Male Subjects ACME Pharmaceutical Company Phase I Trial Y DOUBLE BLIND ACTIVE PLACEBO PARALLEL ... ... ... ... ... ... ... ... ... Page 30 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row STUDYID DOMAIN TSSEQ TSGRPID TSPARMCD 9 EX1 TS 1 OBJPRIM 10 EX1 TS 1 OBJSEC 11 EX1 TS 2 OBJSEC 12 EX1 TS 3 OBJSEC 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 EX1 TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS TS 1 2 3 1 1 1 1 1 1 1 2 1 2 1 2 3 4 1 2 3 4 1 2 3 4 1 2 3 4 1 1 1 1 1 1 1 1 1 TTYPE TTYPE TTYPE TDIGRP AGEMIN AGEMAX AGEU SEXPOP ADDON TRT TRT COMPTRT COMPTRT DOSE DOSE DOSE DOSE DOSU DOSU DOSU DOSU DOSFRQ DOSFRQ DOSFRQ DOSFRQ ROUTE ROUTE ROUTE ROUTE LENGTH PLANSUB EGCTMON EGRDMETH EGBLIND EGTWVALG EGLEADPR EGREPLTR EGREPLBL Page 31 Draft A-1 A-100 MOXI PBO A-1 A-100 MOXI PBO A-1 A-100 MOXI PBO A-1 A-100 MOXI PBO A-1 A-100 MOXI PBO TSPARM TSVAL The primary objective of this study is to evaluate the effects of therapeutic and Trial Primary Objective supratherapeutic doses of Drug relative to placebo on cardiac repolarization following multiple oral doses in healthy adult male subjects To characterize the pharmacokinetic profiles of 1 mg and 100 mg doses of Drug A and its Trial Secondary Objective metabolites in healthy adult male subjects To evaluate the safety and tolerability of 1 mg and 100 mg doses of Drug A in healthy adult Trial Secondary Objective male subjects To establish assay sensitivity to the response in QT/QTc following a single dose of 400 mg Trial Secondary Objective moxifloxacin in healthy adult male subjects Trial Type SAFETY Trial Type PHARMACOKINETIC Trial Type THOROUGH QT Diagnosis Group HEALTHY SUBJECTS Planned Minimum Age of Subjects 18 Planned Maximum Age of Subjects 45 Age Unit YEARS Sex of Participants M Added on to Existing Treatments N Reported Name of Test Product Drug A - 1 mg Reported Name of Test Product Drug A - 100 mg Comparative Treatment Name moxifloxacin Comparative Treatment Name placebo Dose per Administration 1 Dose per Administration 100 Dose per Administration 400 Dose per Administration 0 Test Product Dose Units mg Test Product Dose Units mg Test Product Dose Units mg Test Product Dose Units mg Test Product Dosing Frequency QD Test Product Dosing Frequency QD Test Product Dosing Frequency ONCE Test Product Dosing Frequency QD Route of Administration ORAL Route of Administration ORAL Route of Administration ORAL Route of Administration ORAL Trial Length P56D Planned Number of Subjects 40 ECG Continuous Monitoring Y ECG Read Method SEMI-AUTOMATIC ECG Reading Blinded Y ECG Twave Algorithm TANGENT METHOD ECG Planned Primary Lead LEAD II ECG Replicates On-Treatment Y ECG Replicates at Baseline Y ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 797 798 799 Proposed terminology for Trial Summary Parameter Test Codes and Trial Summary Parameter Test Names is given below:. CDISC Submission Value TSPARMCD EGCTMON EGRDMETH EGBLIND EGTWVALG EGLEADPR EGREPLTR EGREPLBL EGLEADSM CDISC Submission Value TSPARM ECG Continuous Monitoring ECG Read Method ECG Reading Blinded ECG Twave Algorithm ECG Planned Primary Lead ECG Replicates On-Treatment ECG Replicates at Baseline ECG Used Same Lead CDISC Definition Indicate whether the ECG measurements are extracted from continuous monitoring. The ECG read method, e.g. manual, semi-automatic, automatic. Indicate whether the ECG readers are blinded to subject details, treatment, day, and time. Algorithm used to determine the end of Twave. The ECG lead intended to be used as the primary lead. Indicate whether replicate ECGs were collected on-treatment. Indicate whether replicate ECGs were collected at baseline. Indicate whether the same lead was used for interval measurements for all ECGs used in the analyses. 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 4.2 Time Point Planning In clinical studies, the timing and collection of ECGs are guided by the known properties of the drug and its metabolites. ECGs are recorded at baseline *, on the day on which steady-state or other intended concentration of the experimental drug is reached (and the same day relative to baseline for placebo), and periodically thereafter. It has been recommended that 13: • In order to study the peak effect on QTc and the QT interval, ECGs should be recorded several times shortly before and around the time of the maximum drug concentration (Tmax). • To evaluate any delayed effects of the drug or its metabolites on cardiac repolarization, ECG recording should continue even after Tmax. • To demonstrate assay sensitivity, ECGs should also be recorded close to the Tmax of the positive control. • To ensure that blinding is maintained, ECGs should be recorded on the same days and at the same time points in all treatment groups. The diagrams below show how ECG data are organized within 10-second ECGs, and how those 10-second ECGs are organized within and across time points (Figure 13 and Figure 14). Although analysis methods that use all the data from continuous monitoring over a long period of time (e.g., 24 hours) are being developed, analysis usually assumes that data are organized by time points. ECGs are recorded in replicates (usually three or more), 30-120 seconds apart, to account for inherent variability. Each replicate lasts 10 seconds. Replicates may be individual recordings or extracted from a continuous recording (typically lasting 24 hours or a multiple thereof). The ECG record is digitally stored for later processing. * Several alternatives for baseline exist, such as: the day before treatment is first administered, or just a 1- or 2-hour period before treatment is first administered. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 32 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 24 hours Beat 1 (7:59:00.00) 817 818 819 820 821 Beat 2 (7:59:01.30) Beat 3 (7:59:02.15) … Beat 12 (7:59:10.05) … Beat 100,800 Extracted 10-second ECG ≈ 12 P-QRS-T complexes Figure 13: Individual Beats and Relevant Identifying Information Within a 24-hour Continuous Recording Each cycle, in a normal ECG obtained from a healthy person, consists of a P-QRS-T complex and the subsequent isoelectric activity before the next P-QRS-T complex. The pertinent information is the sequence number and the time. Page 33 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 822 823 824 825 826 827 828 829 830 831 832 833 Figure 14: ECG Replicates and Nominal Time Points The concept map below displays the process and timing of potential parameters (such as vital signs and PK) and their order of collection in a QT study. In a typical QT study, ECGs are recorded at baseline on the day of treatment (day 1) and at the same pre-defined clock times (time matched) within each of the treatment groups. A stable heart rate is typically obtained via ECGs taken after subjects have rested (but not while sleeping) for at least 5 to 10 minutes in the supine position. Vital signs such as blood pressure and heart rate in the supine and/or standing positions can be collected at the same time points as the ECG recordings, immediately after collecting the ECGs. However, the assessment of orthostatic challenge is optional and dependent on the study objectives. To investigate potential exposure-response relationships and confirm the availability of the drug, blood samples for PK measurements can be collected at the same time points as the ECG recordings. Blood draws are performed after the ECG recordings to avoid confounding stress. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 34 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 834 835 836 837 838 This concept map displays the process and timing of a collection of parameters if used in a QT study. Note that multiple ECG replicates are recorded (the number can vary but is generally 3 or more), 30-120 seconds apart, to account for inherent variability; each recording lasting 10 seconds. 839 5 Subject Characteristics and Eligibility 840 5.1 Inclusion/Exclusion Criteria 841 842 843 844 845 Typically, TQT studies are conducted in a healthy population unless there are safety and tolerability concerns with administration of the drug to healthy subjects. Inclusion of healthy subjects minimizes confounding factors of underlying disease, co-morbidities and/or concomitant medications. Thus, standard inclusion/exclusion criteria for healthy subject studies are often applied in TQT studies. TQT studies are designed to protect subject safety. Furthermore, consideration may be given to excluding individuals who have a risk for Torsades de pointes (e.g., cardiac disease, hypokalemia, personal history of unexplained syncopal episodes, or known personal or family history of Long QT Syndrome) and individuals who have repeatedly high QTcF values at baseline. TQT studies Concept Map 3: Potential Parameters and Their Order of Collection Page 35 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 846 847 848 849 850 851 are also designed to guard against falsely concluding a lack of effect on QT/QTc and to limit all potential sources of QT/QTc variability such as concomitant medications or other substances that may prolong QT intervals. Note that general inclusion and exclusion criteria are also addressed in other CDISC standards. The table below provides example exclusion criteria for studies evaluating QT. Data collection for these data is already well documented in the current SDTM IG using the IE domain. Protect Patient Safety Exclusion Criteria A known or suspected hypersensitivity to 1 study drug, moxifloxacin or any components of the formulations used. Subject has any clinically significant abnormality following the investigator’s review of the physical examination, and protocol-defined clinical laboratory tests at Screening or day -1. 2 a. For the following laboratory analytes, any result outside reference range, even if NOT judged clinically significant is reason for exclusion: sodium, potassium, chloride, calcium, magnesium Any prescription medication, over-thecounter medication, herbal medication or product (except for vitamins supplements 3 without herbal additives and acetaminophen) within 2 weeks of first treatment or during the study. Any use of recreational drugs or a positive 4 drug screen. Excessive methylxanthine (caffeine, etc.) 5 use, exceeding _____ per day Smoking >____ cigarette equivalents per 6 day. Alcohol intake >____ U per day (U = 1.5oz 7 of ≥80 proof spirit equivalent). Methylxanthine, nicotine, alcohol, or 8 grapefruit juice consumption within 2 weeks of any treatment. Resting, basal systolic BP ≥140 mmHg or 9 diastolic BP ≥90 mmHg. 10 Orthostatic hypotension or postural Guard Against Falsely Concluding No Effect and Eliminate Confounding Influence (confounds increase within and between subject variance) Additional Details X X X X X X X © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 X X Examination and lab abnormalities can suggest poor health that predispose to safety concerns when QTc is prolonged. This is especially the case for electrolytes. Alterations in electrolytes can alter the normal configuration of the T wave. Substances can both alter metabolism of test drug and thereby alter any potential influence, and directly alter the QTc itself and alter autonomic tone that alters T wave response to change in heart rate and change heart rate, complicating analyses through correction. Substances may alter the metabolism of test drug, the autonomic tone and/or alter heart rate. X Alcohol intake may alter the autonomic tone and/or alter heart rate. X Substances may alter the metabolism of test drug, the autonomic tone and/or alter heart rate. X These factors suggest a disturbed autonomic tone. X Page 36 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 11 12 Orthostatic Tachycardia Personal or family history of: syncope; ventricular tachydysrhythmia; cardiac arrest / sudden cardiac death; congenital long QT syndrome. Specific ECG exclusionary criteria: a. Heart rate <40 or >99 bpm b. PR <120 or >200 ms c. QRS >110 ms d. QTcF males >430 ms, females >450 ms e. Any qualitative / morphological abnormality except: sinus arrhythmia; isolated premature atrial complexes / premature ventricular complexes f. T-wave / U-wave characteristics making determination of the end of the T-wave difficult such as biphasic T-waves; U-waves of width greater than 1/3 the width of the preceding T-wave Protect Patient Safety Guard Against Falsely Concluding No Effect and Eliminate Confounding Influence (confounds increase within and between subject variance) X X X (d only) X (all) Additional Details Suggests a lack of safety margin if QTc prolonged and potential for prolongation due to factors other than drug influence HR: difficulties in correction - confound; PR & QRS: lack of good normal cardiac electrophysiology - some mild safety concern and confound; QTc: safety concern if further prolonged and confound; Other abnormalities: lack of good normal heart - confound; T wave: abnormality - confound and difficulty in measuring T wave 852 853 854 855 In general, a minimum age of 18 years old is an entry criterion, and in order to promote homogeneity in the experimental population, some maximum age criterion is likely to be in place, which can be variable given enrolment considerations. The list above does not cover all potential exclusion criteria that sponsors might select. 856 5.2 Pharmacogenetics 857 858 859 860 861 862 863 864 865 866 867 868 Several genetic factors that alter the risk of drug-induced QT prolongation have been reported. Some of these genetic factors are associated with Long QT Syndrome. Many forms of Long QT Syndrome have been linked to mutations in genes encoding cardiac ion channel proteins. Polymorphisms can affect ion channels, leading to an increased sensitivity to drugs that affect repolarization. Because of incomplete penetrance, not all carriers of non-wildtype alleles of ion channel genes will manifest QT/QTc interval prolongation in screening ECG evaluations. Other genetic factors may affect drug metabolism genes (mainly CYP2D6 and CYP3A4) 14. Poor metabolizers accumulate excess concentrations of drugs. If the drug blocks relevant cardiac ion channels, this poses a risk of QT interval prolongation. This would be particularly exacerbated when combined with polymorphisms associated with Long QT Syndrome. Thus, the presence of either or both types of genetic variations may predispose a subject to a potentially fatal adverse reaction, even at therapeutic drug doses 15. The ICH E14 2005 guidance on QT prolongation advises considering genotyping patients who experience marked prolongation of QT/QTc or TdP while on drug therapy 1. Page 37 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 869 870 871 872 873 874 875 876 877 878 879 The pharmacogenomics/genetics (PGx) team, a sub-team within the CDISC SDS Team (Clinical Data Interchange Standards Consortium-Study Data Submission), has developed several domains designed to carry PGx data. The PGx findings domain stores key results such as intensity values (both raw and normalized), p-value, fold-change, ratio, nucleotide variation, amino acid variation, etc. The initial development of these CDISC PGx domains was done in parallel with the work being done by the Health Level Seven Clinical Genomics Work Group. Please refer to the SDTM Implementation Guide for Pharmacogenomics/Genetics (STDMIG-PGx)† for these domains. 880 6 Study Assessments 881 6.1 ECG Assessments 882 883 884 885 886 887 Genetic testing may occur prospectively in QT studies to exclude subjects predisposed to genetic Long QT Syndrome or Short QT Syndrome, or it may be executed post hoc in a case of unusual results. Additionally, prospective genetic testing may be performed either to exclude slow or rapid metabolizers or to include only slow metabolizers. Note that the inclusion of slow metabolizers is a potential option to achieve increased drug exposure with lower doses in lieu of using higher doses greater than maximum clinical dose or administering an inhibitor. Quantitative ECG data (e.g., HR, PR interval, QRS, absolute QT interval, and QTc) represent one interest of any ECG data analysis. Qualitative ECG data are descriptive findings based on visual interpretation of the waveforms, sometimes referred to as morphological findings. These data include electrical axis ‡, heart rhythm, conduction, evidence of ischemia, ST segment and T-wave and U-wave shapes. These data also provide extremely important data for analysis (Concept Map 4). It is important to note that nearly all abnormal, outlier quantitative data are reflected in the qualitative findings. For example, a PR interval greater than the upper reference limit of 220 msec would lead to the conduction qualitative finding of 1st-degree AV block. † Currently in development as of the publication of this document. The Virology Therapeutic Area Data Standards User Guide (VR-UG, available at: http://www.cdisc.org/stuff/contentmgr/files/0/2356ae38ac190ab8ca4ae0b222392b37/misc/vr_ug_v1_0_prov.pdf) has prototype versions of the domains. ‡ Also measured as a quantitative finding. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 38 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 888 889 890 891 892 Concept Map 4: ECG Quantitative Results and Morphological (Qualitative) Findings Determination This concept map displays the process for determining the measurements of the PR, QT, RR intervals and QRS complex as well as determining any abnormal qualitative findings on the ECG. Page 39 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 6.1.1 Specification for ECG Test Results The ECG Test Results (EG) domain model is a fully approved and formally published domain model. The table below was modified from the current standard as defined in the SDTMIG 3.2 to extend the record structure to cover beat-to-beat records as well as conventional 10-second ECGs, and to add two pre-existing variables (SPDEVID and EPOCH) and two new variables (EGREPNUM and EGBEATNO §) to the specification. SPDEVID and EPOCH are identifier and timing variables, respectively, which may be added to any domain model. EGBEATNO is used to differentiate between beats in beat-to-beat records, and EGREPNUM is used to differentiate between multiple repetitions of a test within a given time frame. Because they are new, they have been highlighted in yellow. An additional column, BRIDG Mapping, has also been added to the table below. This column contains the BRIDG class and attribute associated with each variable. Sometimes no mapping exists for a particular variable, in which case a brief comment as to why is given in italics. For non-individual ECG beat data, and for aggregate ECG parameter results (e.g., QT interval, RR, PR, QRS), EGREFID is populated for all unique ECGs, so that submitted SDTM data can be matched to the actual ECGs stored in the ECG warehouse. Therefore, this variable is expected for these types of records. For individual-beat parameter results, waveform data will not be stored in the warehouse, so there will be no associated identifier for these beats. Note that EGREFID holds the identifier or accession number for the actual ECG stored in the warehouse, rather than its file name and path. Controlled terminology is used where it exists; however, new terminology such as that for beat-to-beat data has been proposed. Please refer to the current CDISC Controlled Terminology (http://www.cancer.gov/cancertopics/cancerlibrary/terminologyresources/cdisc). eg.xpt, ECG Test Results — Findings. One record per ECG observation per replicate per time point or one record per ECG observation per beat per visit per subject, Tabulation. Variable Name Variable Label STUDYID DOMAIN USUBJID SPDEVID EGSEQ EGGRPID § Study Identifier Domain Abbreviation Unique Subject Identifier Sponsor Device Identifier Sequence Number Group ID DocumentIdentifier.identifier PerformedObservation Controlled Type Terms, Codelist Role or Format Char Identifier Char EG Identifier SubjectIdentifier.identifier Char Identifier MaterialIdentifier.identifier Char Identifier implementation specific record identifier implementation specific record grouping variable Num Identifier Char Identifier BRIDG Mapping CDISC Notes Unique identifier for a study. Two-character abbreviation for the domain. Core Req Req Identifier used to uniquely identify a subject across all studies Req for all applications or submissions involving the product. Sponsor-defined identifier for a device Perm Sequence Number given to ensure uniqueness of subject records within a domain. May be any valid number. Used to tie together a block of related records in a single domain for a subject. Req Perm Details for the new variables are in accordance with SDTM Governance via the SDTM Variable Requests tracking tool as of July 15, 2014, but may be subject to change prior to formal publication. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 40 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Variable Name Variable Label EGREFID EGSPID BRIDG Mapping ECG Reference implementation specific ID Sponsor-Defined implementation specific Identifier Controlled Type Terms, Codelist Role or Format Char Identifier Char Identifier EGTESTCD ECG Test or Examination Short Name DefinedObservation.nameCode Char (EGTESTCD) ! Topic EGTEST ECG Test or Examination Name DefinedObservation.nameCode Char (EGTEST) ! Synonym Qualifier EGCAT Category for ECG Subcategory for ECG ECG Position of Subject ECG Beat Number Result or Finding in Original Units DefinedActivity.categoryCode Char * Grouping Qualifier Grouping Qualifier Record Qualifier Result Qualifier Result Qualifier Original Units PerformedObservationResult.value Char (UNIT) EGSCAT EGPOS EGBEATNO EGORRES EGORRESU Page 41 Draft DefinedActivity.subcategoryCode Char * PerformedObservation. bodyPositionCode new variable Char (POSITION) PerformedObservationResult. value Char Num Variable Qualifier CDISC Notes Core Internal or external ECG identifier. Perm Example: UUID. Sponsor-defined reference number. Perhaps pre-printed on the Perm CRF as an explicit line identifier or defined in the sponsor's operational database. Example: Line number from the ECG page. Short name of the measurement, test, or examination described Req in EGTEST. It can be used as a column name when converting a dataset from a vertical to a horizontal format. The value in EGTESTCD cannot be longer than 8 characters, nor can it start with a number (e.g., “1TEST”). EGTESTCD cannot contain characters other than letters, numbers, or underscores. Examples: PRAG, QTAG Verbatim name of the test or examination used to obtain the Req measurement or finding. The value in EGTEST cannot be longer than 40 characters. Examples: PR Interval, Aggregate, QT Interval, Aggregate Used to categorize ECG observations across subjects. Perm Examples: MEASUREMENT, FINDING, INTERVAL. A further categorization of the ECG. Perm Position of the subject during a measurement or examination. Examples: SUPINE, STANDING, SITTING. Variable describing ECG measurements of individual beat data. Examples: 1,2,3 Result of the ECG measurement or finding as originally received or collected. Examples of expected values are 62 or 0.151 when the result is an interval or measurement, or “ATRIAL FIBRILLATION” or “QT PROLONGATION” when the result is a finding. Original units in which the data were collected. The unit for EGORRES. Examples: sec or msec. Perm Perm Exp Perm © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Variable Name Variable Label EGSTRESC Character Result/Finding in Std Format EGSTRESN Numeric Result/Finding in Standard Units Standard Units EGSTRESU Controlled Type Terms, Codelist Role or Format PerformedObservationResult.value Char Result Qualifier BRIDG Mapping PerformedObservationResult. value Num Result Qualifier PerformedObservationResult. value PerformedObservation. negationIndicator Char (UNIT) Variable Qualifier Record Qualifier EGSTAT Completion Status EGREASND Reason ECG Not PerformedObservation. Performed negationReason Char Record Qualifier EGXFN External File Name Vendor Name PerformedObservationResult. value Organization.name Char Lead Location Used for Measurement Method of ECG Test Baseline Flag complex mapping Char (EGLEAD) Record Qualifier Record Qualifier Record Qualifier PerformedObservation. methodCode PerformedObservationResult. baselineIndicator Char (EGMETHOD) EGNAM EGLEAD EGMETHOD EGBLFL Char (ND) Char Char (NY) © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Record Qualifier Record Qualifier CDISC Notes Core Contains the result value for all findings, copied or derived Exp from EGORRES in a standard format or standard units. EGSTRESC should store all results or findings in character format; if results are numeric, they should also be stored in numeric format in EGSTRESN. For example, if a test has results of “NONE”, “NEG”, and “NEGATIVE” in EGORRES and these results effectively have the same meaning, they could be represented in standard format in EGSTRESC as “NEGATIVE”. For other examples, see general assumptions. Additional examples of result data: SINUS BRADYCARDIA, ATRIAL FLUTTER, ATRIAL FIBRILLATION. . Used for continuous or numeric results or findings in standard Perm format; copied in numeric format from EGSTRESC. EGSTRESN should store all numeric test results or findings. Standardized unit used for EGSTRESC or EGSTRESN. Perm Used to indicate an ECG was not done, or an ECG Perm measurement was not taken. Should be null if a result exists in EGORRES. Describes why a measurement or test was not performed. Perm Examples: BROKEN EQUIPMENT or SUBJECT REFUSED. Used in conjunction with EGSTAT when value is NOT DONE. File name and path for the external ECG Waveform file. Perm Name or identifier of the laboratory or vendor who provided the test results. The lead used for the measurement. Examples: LEAD V1, LEAD V6, LEAD aVR, LEAD I, LEAD II, LEAD III Perm Method of the ECG test. Examples: 12 LEAD STANDARD. Perm Perm Indicator used to identify a baseline value. The value should be Exp “Y” or null. Page 42 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) EGDRVFL Derived Flag Defer to Statistical Analysis Modeling. Controlled Type Terms, Codelist Role or Format Char (NY) Record Qualifier EGEVAL Evaluator Performer.typeCode Char * Record Qualifier VISITNUM Visit Number Num Timing VISIT Visit Name Char Timing VISITDY Planned Study Day of Visit Epoch PlannedSubjectActivityGroup. sequenceNumber PlannedSubjectActivityGroup. name PlannedActivity.studyDayRange Num Timing Epoch.name Char * Timing Variable Name Variable Label EPOCH BRIDG Mapping EGDTC Date/Time of ECG PerformedObservation.dateRange Char ISO 8601 Timing EGDY Study Day of ECG PerformedObservation. studyDayRange Num Timing EGTPT Planned Time Point Name PlannedSubjectActivityGroup. name Char Timing EGTPTNUM Planned Time Point Number Planned Elapsed Time from Time Point Ref PlannedSubjectActivityGroup. Num sequenceNumber PlannedContingentOnRelationship Char ISO 8601 EGELTM Page 43 Draft PlannedCompositionRelationship. pauseQuantity Timing Timing CDISC Notes Used to indicate a derived record. The value should be Y or null. Records which represent the average of other records, or that do not come from the CRF, or are not as originally collected or received are examples of records that would be derived for the submission datasets. If EGDRVFL=Y, then EGORRES could be null, with EGSTRESC, and (if numeric) EGSTRESN having the derived value. Role of the person who provided the evaluation. Used only for results that are subjective (e.g., assigned by a person or a group). Should be null for records that contain collected or derived data. Examples: INVESTIGATOR, ADJUDICATION COMMITTEE, VENDOR. 1. Clinical encounter number. 2. Numeric version of VISIT, used for sorting. 1. Protocol-defined description of clinical encounter. 2. May be used in addition to VISITNUM and/or VISITDY. Planned study day of the visit based upon RFSTDTC in Demographics. Epoch associated with the Element in the planned sequence of Elements for the ARM to which the subject was assigned Date/Time of ECG (note: the next version of the SDTMIG will reflect “date/time of ECG”) 1. Study day of the ECG, measured as integer days. 2. Algorithm for calculations must be relative to the sponsordefined RFSTDTC variable in Demographics. 1. Text Description of time when measurement should be taken. 2. This may be represented as an elapsed time relative to a fixed reference point, such as time of last dose. See EGTPTNUM and EGTPTREF. Examples: Start, 5 min post. Numerical version of EGTPT to aid in sorting. Core Perm Perm Exp Perm Perm Perm Exp Perm Perm Perm Planned elapsed time (in ISO 8601) relative to a fixed time point Perm reference (EGTPTREF). Not a clock time or a date time variable. Represented as an ISO 8601 duration. Examples: “-PT15M” to represent the period of 15 minutes prior to the reference point indicated by EGTPTREF, or “PT8H” to represent the period of 8 hours after the reference point indicated by EGTPTREF. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Variable Name Variable Label BRIDG Mapping EGTPTREF Time Point Reference EGRFTDTC Date/Time of PerformedActivity.dateRange Char ISO 8601 Reference Time Point Repetition PerformedObservation.repetitionN Num Number umber EGREPNUM DefinedActivity.nameCode Controlled Type Terms, Codelist Role or Format Char Timing Timing Name of the fixed reference point referred to by EGELTM, EGTPTNUM, and EGTPT. Examples: PREVIOUS DOSE, PREVIOUS MEAL. Date/time of the reference time point, EGTPTREF. Core Perm Perm Timing The incidence number of a test that is repeated within a given Perm timeframe for the same test. The level of granularity can vary, e.g., within a time point or within a visit. For example, multiple measurements of blood pressure or multiple analyses of a sample. * Indicates variable may be subject to controlled terminology, (Parenthesis indicates CDISC/ NCI codelist code value), ! Indicates terminology to be reviewed by the CDISC Terminology Team. BRIDG Mapping values in italics are comments on why no mapping exists for that variable. 917 918 919 920 For assumptions for the ECG Tests Results domain model, please refer to the current SDTMIG. 921 6.1.2 Examples for ECG Test Results 922 923 924 925 926 927 928 929 CDISC Notes Example 1 This example is from 10-second ECG replicates extracted from a continuous recording. The example shows one subject’s (USUBJID=2324-P0001) extracted 10second ECG replicate results. Three replicates were extracted for planned time points 1 HR and 2 HR. Summary mean measurements are reported for the 10 seconds of extracted data for each replicate. EGDTC is the date/time of the first individual beat in the extracted 10-second ECG. In this example, corrections to QT were performed by the sponsor and are not included; however, this information can be found in the ADaM dataset. In order to save space, some permissible variables (EGREFID, VISITDY, EGTPTNUM, EGTPTREF, EGRFTDTC) have been omitted, as marked by an ellipsis. eg.xpt Row 1 2 3 4 6 6 7 8 9 10 11 12 13 14 STUDYID DOMAIN USUBJID EGSEQ EGTESTCD EGTEST STUDY01 EG 2324-P0001 1 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 2 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 3 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 4 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 5 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 6 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 7 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 8 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 9 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 10 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 11 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 12 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 13 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 14 RRAG RR Interval, Aggregate © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 EGCAT INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL EGPOS EGORRES EGORRESU EGSTRESC EGSTRESN EGSTRESU SUPINE 176 msec 176 176 msec SUPINE 658 msec 658 658 msec SUPINE 97 msec 97 97 msec SUPINE 440 msec 440 440 msec SUPINE 176 msec 176 176 msec SUPINE 679 msec 679 679 msec SUPINE 95 msec 95 95 msec SUPINE 389 msec 389 389 msec SUPINE 169 msec 169 169 msec SUPINE 661 msec 661 661 msec SUPINE 90 msec 90 90 msec SUPINE 377 msec 377 377 msec SUPINE 176 msec 176 176 msec SUPINE 771 msec 771 771 msec Page 44 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row 15 16 17 18 19 20 21 22 23 24 STUDYID DOMAIN USUBJID EGSEQ EGTESTCD EGTEST STUDY01 EG 2324-P0001 15 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 16 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 17 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 18 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 19 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 20 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 21 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 22 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 23 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 24 QTAG QT Interval, Aggregate EGCAT INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL EGPOS EGORRES EGORRESU EGSTRESC EGSTRESN EGSTRESU SUPINE 100 msec 100 100 msec SUPINE 379 msec 379 379 msec SUPINE 179 msec 179 179 msec SUPINE 749 msec 749 749 msec SUPINE 103 msec 103 103 msec SUPINE 402 msec 402 402 msec SUPINE 175 msec 175 175 msec SUPINE 771 msec 771 771 msec SUPINE 98 msec 98 98 msec SUPINE 356 msec 356 356 msec 930 Row EGLEAD EGMETHOD VISITNUM VISIT EGDTC EGTPT … EGREPNUM 2 VISIT 2 2014-03-22T10:00:21 1 HR … 1 1 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:00:21 1 HR … 1 2 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:00:21 1 HR … 1 3 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:00:21 1 HR … 1 4 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:01:35 1 HR … 2 5 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:01:35 1 HR … 2 6 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:01:35 1 HR … 2 7 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:01:35 1 HR … 2 8 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:02:14 1 HR … 3 9 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:02:14 1 HR … 3 10 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:02:14 1 HR … 3 11 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T10:02:14 1 HR … 3 12 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:00:21 2 HR … 1 13 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:00:21 2 HR … 1 14 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:00:21 2 HR … 1 15 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:00:21 2 HR … 1 16 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:01:31 2 HR … 2 17 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:01:31 2 HR … 2 18 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:01:31 2 HR … 2 19 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:01:31 2 HR … 2 20 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:02:40 2 HR … 3 21 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:02:40 2 HR … 3 22 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:02:40 2 HR … 3 23 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 2014-03-22T11:02:40 2 HR … 3 24 (cont) LEAD II 12 LEAD STANDARD 931 932 933 934 935 936 937 938 939 Example 2 This example is from ECG results where beat-to-beat measurements were recorded. The example shows one subject’s (USUBJID=2324-P0001) continuous beatto-beat EG results. Only 3 beats are shown, but there could be measurements for, as an example, 101,000 complexes in 24 hours. The actual number of complexes in 24 hours can be variable and depends on average heart rate. The results are mapped to the EG (ECG Test Results) domain using EGBEATNO. The controlled terminology for EGTESTCD, EGTEST, EGLEAD, and EGMETHOD is represented in the table. If there is no result to be reported, then the row would not be included. Page 45 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 940 941 942 943 944 Rows 1-2: Rows 3-8: There is no RR measurement for the first beat (EGBEATNO=1), because RR is measured as the duration (time) between the peak of the R wave in the reported single beat and peak of the R wave in the preceding single beat. EGDTC is the date/time of the individual beat. There is a 1-hour gap between beats 2 and 3 due to electrical interference or other artifacts that prevented measurements from being recorded. eg.xpt Row 1 2 3 4 5 6 7 8 STUDYID DOMAIN USUBJID EGSEQ EGTESTCD EGTEST STUDY01 EG 2324-P0001 1 PRSB PR Interval, Single Beat STUDY01 EG 2324-P0001 2 QRSSB QRS Duration, Single Beat STUDY01 EG 2324-P0001 3 PRSB PR Interval, Single Beat STUDY01 EG 2324-P0001 4 RRSB RR Interval, Single Measurement STUDY01 EG 2324-P0001 5 QRSSB QRS Duration, Single Beat STUDY01 EG 2324-P0001 6 PRSB PR Interval, Single Beat STUDY01 EG 2324-P0001 7 RRSB RR Duration, Single Measurement STUDY01 EG 2324-P0001 8 QRSSB QRS Duration, Single Beat EGCAT INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL EGPOS EGBEATNO EGORRES EGORRESU SUPINE 1 176 msec SUPINE 1 97 msec SUPINE 2 176 msec SUPINE 2 679 msec SUPINE 2 95 msec SUPINE 3 169 msec SUPINE 3 661 msec SUPINE 3 90 msec 945 Row EGSTRESC EGSTRESN EGSTRESU EGLEAD EGMETHOD VISITNUM VISIT VISITDY EGDTC 176 176 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T14:32:12.3 1 (cont) 97 97 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T14:32:12.3 2 (cont) 176 176 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T14:32:13.3 3 (cont) 679 679 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T14:32:13.3 4 (cont) 95 95 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T14:32:13.3 5 (cont) 169 169 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T15:32:14.2 6 (cont) 661 661 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T15:32:14.2 7 (cont) 90 90 msec LEAD II 12 LEAD STANDARD 1 SCREENING -7 2014-02-11T15:32:14.2 8 (cont) 946 947 948 949 950 951 952 953 954 955 956 Example 3 In order to be able to indicate which ECG records were used in the calculation of the population and individual QT correction coefficients, the SUPPEG dataset was used to flag those records that were used in this evaluation. This example also shows the corresponding EG dataset with pre-dose ECG data for one subject’s (USUBJID=2324-P0001) extracted 10-second ECG replicate results. Summary mean measurements are reported for the 10 seconds of extracted data (for each replicate). EGDTC is the date/time of the start of the 10-second ECG segment. Rows 1-12: Rows 13-16: Extract of pre-dose ECG results showing results for PR, RR, QRS and QT measurements. Extract of post-dose ECG results showing results for PR, RR, QRS and QT measurements. Only the first replicate is included. eg.xpt Row 1 2 3 4 5 6 7 8 STUDYID DOMAIN USUBJID EGSEQ EGTESTCD EGTEST STUDY01 EG 2324-P0001 1 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 2 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 3 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 4 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 5 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 6 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 7 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 8 QTAG QT Interval, Aggregate © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 EGCAT INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL EGPOS EGORRES EGORRESU EGSTRESC EGSTRESN EGSTRESU SUPINE 176 msec 176 176 msec SUPINE 658 msec 658 658 msec SUPINE 97 msec 97 97 msec SUPINE 343 msec 343 343 msec SUPINE 176 msec 176 176 msec SUPINE 679 msec 679 679 msec SUPINE 95 msec 95 95 msec SUPINE 345 msec 345 345 msec Page 46 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row 9 10 11 12 13 14 15 16 STUDYID DOMAIN USUBJID EGSEQ EGTESTCD EGTEST STUDY01 EG 2324-P0001 9 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 10 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 11 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 12 QTAG QT Interval, Aggregate STUDY01 EG 2324-P0001 13 PRAG PR Interval, Aggregate STUDY01 EG 2324-P0001 14 RRAG RR Interval, Aggregate STUDY01 EG 2324-P0001 15 QRSAG QRS Duration, Aggregate STUDY01 EG 2324-P0001 16 QTAG QT Interval, Aggregate EGCAT INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL INTERVAL EGPOS EGORRES EGORRESU EGSTRESC EGSTRESN EGSTRESU SUPINE 169 msec 169 169 msec SUPINE 661 msec 661 661 msec SUPINE 90 msec 90 90 msec SUPINE 352 msec 352 352 Msec SUPINE 176 msec 173 176 msec SUPINE 658 msec 666 658 msec SUPINE 97 msec 94 97 msec SUPINE 343 msec 347 343 msec 957 Row EGLEAD EGMETHOD VISITNUM VISIT VISITDY EGDTC EGTPT … EGREPNUM 2 VISIT 2 1 2014-02-11T14:32:12 Pre-dose … 1 1 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:32:12 Pre-dose … 1 2 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:32:12 Pre-dose … 1 3 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:32:12 Pre-dose … 1 4 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:33:13 Pre-dose … 2 5 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:33:13 Pre-dose … 2 6 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:33:13 Pre-dose … 2 7 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:33:13 Pre-dose … 2 8 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:34:14 Pre-dose … 3 9 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:34:14 Pre-dose … 3 10 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:34:14 Pre-dose … 3 11 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T14:34:14 Pre-dose … 3 12 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T15:02:12 30 min post … 1 13 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T15:02:12 30 min post … 1 14 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T15:02:12 30 min post … 1 15 (cont) LEAD II 12 LEAD STANDARD 2 VISIT 2 1 2014-02-11T15:02:12 30 min post … 1 16 (cont) LEAD II 12 LEAD STANDARD 958 959 960 961 962 Rows 1-12: Links those records from eg.xpt that were used in the calculation of the population and individual QT correction coefficients (i.e. the RR and QT measurements). The data between suppeg.xpt and eg.xpt is linked via the EGSEQ variable in eg.xpt (IDVAR and IDVARVAL in suppeg.xpt). suppeg.xpt Row 1 2 3 4 5 6 7 8 9 10 11 12 STUDYID RDOMAIN USUBJID IDVAR IDVARVAL QNAM QLABEL QVAL QORIG QEVAL STUDY01 EG 2324-P0001 EGSEQ 2 QTCCIFL Individ QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 2 QTCCNFL Populatn QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 4 QTCCIFL Individ QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 4 QTCCNFL Populatn QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 6 QTCCIFL Individ QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 6 QTCCNFL Populatn QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 8 QTCCIFL Individ QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 8 QTCCNFL Populatn QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 10 QTCCIFL Individ QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 10 QTCCNFL Populatn QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 12 QTCCIFL Individ QT Correction Coefficient Flag Y Derived STUDY01 EG 2324-P0001 EGSEQ 12 QTCCNFL Populatn QT Correction Coefficient Flag Y Derived 963 Page 47 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 6.2 QT Correction Building on the information presented in Section 2.3.4, the following is a summary of the correction methods that can be used in TQT studies. Comparing QT intervals to determine whether a drug has affected them works best when the drug is the only variable to be taken into consideration, which means correcting the raw data for the other major influencing factor, i.e., heart rate. HR-corrected QT (QTc) intervals may be derived in a variety of ways: • Historical Population QTc: Derived by fitting a model to data from a historical population of subjects. The formula for correcting QT, including its coefficient(s), is based on a formula which is known before the study starts, such as Bazett’s or Fridericia’s corrections. • Study Population QTc: Derived by fitting a model to data from the population of subjects under study. The formula for correcting QT is known before the study starts, but the coefficient(s) in the correction formula are derived from data collected in the study. • Multi-model Study Population QTc: Derived by fitting several models to data for the study population, then identifying one or more “best” models and using a correction method based on the best model(s). The candidate models, the method for choosing the best model(s) and, for methods that use multiple best models, the way in which the best models are combined, are known before the study starts. The formulas for correcting QT, as well as its coefficient(s), are derived from data collected in the study. • Individual QTc: Derived by fitting a model to data for each study subject individually. The formula for correcting QT is known before the study starts, but the coefficient(s) in the formula are derived from data collected from the individual subject. • Multi-model Individual QTc: Derived by fitting several models to data for each subject, then identifying one or more “best fit” models for the subject and using a correction method based on the best fit model(s). The candidate models, the method for choosing the best model(s) and, for methods that use multiple best models, the way in which the best models are combined, are known before the study starts. The formulas for correcting QT, as well as its coefficient(s), are derived from data collected from the individual subject. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 48 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 986 This concept map displays the process for determining QT correction 987 coefficients for a pre-specified model for a study population. 988 989 990 Concept Map 5: Population QT Correction for a Pre-specified Model This concept map displays the process for determining QT correction coefficients for a pre-specified model for individuals. Concept Map 6: Individual QT Correction for a Pre-specified Model 991 Page 49 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 992 993 994 995 This concept map displays the process for determining QT correction formula and coefficient(s) for Individual QTc ‘Best Fit’ Model. Concept Map 7: Individual QT Correction (QTc) ‘Best Fit’ Model © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 50 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 For each QT correction method used in the study, values of EGTESTCD and EGTEST are assigned at the study level. CDISC Controlled Terminology includes EGTESTCD and EGTEST values for QT corrected by Bazett’s method and QT corrected by Fridericia’s method. CDISC Controlled Terminology does not plan to include any other corrected QT tests in CDISC Controlled Terminology in the near future. This is because of the large and growing number of QT corrections in use, and the fact that there is not scientific consensus on a best fit QT correction method. A protocol may specify that several different QT correction models be assessed and that only the best fit model for the individual be used for calculating the individual’s corrected QT intervals. Another alternative is that the protocol specifies multiple models be computed for each individual irrespective of best fit. There are multiple scenarios of analysis, and the sponsor should assign values for EGTESTCD/EGTEST appropriately with clear documentation on what each test code represents. EGTESTCD's begin with the fragment 'QTCI' for individual-based corrected QT intervals, and 'QTCN' for population-based corrected QT intervals. For example, if the protocol calls for computing the top two best fit models, the sponsor could choose to name the top best fit model QTCIAG1 and the second best fit model QTCIAG2, in rank order. This domain provides information about the QT corrections used in the study. In that sense, it contains supporting information about tests. The name of the test about which information is being supplied is stored in QTGRPID. This domain is provided when coefficients for QT corrections other than Bazett’s or Fridericia’s are used in a study, and those corrections are derived from data collected in the study (i.e., are not historical), and the derivation was performed by a vendor, rather than the sponsor. Information on historical correction methods or derived by the sponsor could be submitted in the QT data domain, if agreed by the submitter and recipient (e.g., study sponsor and regulatory agency). For both individual and population corrections, the USUBJID will be populated. 6.2.1 Specification for ECG QT Correction The following table is based on the proposed SDTM Findings QT domain model. An additional column, BRIDG 3.0.3 Mapping, has also been included. Controlled terminology is used where it exists. Please refer to the current CDISC Controlled Terminology (http://www.cancer.gov/cancertopics/cancerlibrary/terminologyresources/cdisc). Sponsor-defined controlled terminology should be provided in the Define-XML. qt.xpt, ECG QT Correction Model Data — Findings. One record per QT correction observation per subject, Tabulation. Variable Name Variable Label STUDYID DOMAIN USUBJID QTSEQ Page 51 Draft BRIDG 3.0.3 Mapping Study Identifier DocumentIdentifier.identifier Domain PerformedObservation Abbreviation Unique Subject SubjectIdentifier.identifier Identifier Sequence Number Type Char Char Controlled Terms, Codelist or Format Role CDISC Notes Core Identifier Identifier Unique identifier for a study. Two-character abbreviation for the domain. Char Identifier implementation specific record Num identifier Identifier Identifier used to uniquely identify a subject across all Req studies for all applications or submissions involving the product. Sequence Number given to ensure uniqueness of Req subject records within a domain. May be any valid number. QT Req Req © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Variable Name Variable Label BRIDG 3.0.3 Mapping QTGRPID Group ID QTREFID QTc Reference implementation specific ID Sponsorimplementation specific Defined Identifier QTSPID Type Controlled Terms, Codelist or Format implementation specific record Char grouping variable Role Identifier Char Identifier Char Identifier QTTESTCD QTc Test or Examination Short Name DefinedObservation.nameCode Char Topic QTTEST QTc Test or Examination Name DefinedObservation.nameCode Char Synonym Qualifier QTCAT Category for QTc Subcategory for QTc Result or Finding in Original Units DefinedActivity.categoryCode Char * DefinedActivity.subcategoryCo Char de PerformedObservationResult. Char value * QTSCAT QTORRES QTSTRESC Character PerformedObservationResult. Result/Finding value in Std Format Char © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Grouping Qualifier Grouping Qualifier Result Qualifier Result Qualifier CDISC Notes Core Used to tie together a block of related records in a single domain for a subject. Example: use EGTESTCD as value for QTGRPID for easier relating of QT dataset to the EG dataset. Internal or external QT identifier. Generally not used for QT dataset. Sponsor-defined reference number. Perhaps pre-printed on the CRF as an explicit line identifier or defined in the sponsor's operational database. Generally not used for QT dataset. Short name of the measurement, test, or examination described in QTTEST. It can be used as a column name when converting a dataset from a vertical to a horizontal format. The value in QTTESTCD cannot be longer than 8 characters, nor can it start with a number (e.g., “1TEST”). QTTESTCD cannot contain characters other than letters, numbers, or underscores. Examples: QTCDESC, QTCFORM. Verbatim name of the test or examination used to obtain the measurement or finding. The value in QTTEST cannot be longer than 40 characters. Examples: QT Correction Method Description, QT Correction Formula Used to categorize QT correction model observations across subjects. Generally not used in QT dataset. A further categorization of the QT correction model. Generally not used in QT dataset. Result of the QT correction model measurement or finding as originally received or collected. Examples of expected values are 0.132 or 0.432 when the result is a coefficient or measurement, or “LINEAR” or “PARABOLIC LOG/LOG” when the result is a formula. Contains the result value for all findings, copied or derived from QTORRES in a standard format or standard units. QTSTRESC should store all results or findings in character format; if results are numeric, they should also be stored in numeric format in QTSTRESN, for example 0.132 or 0.432 when the result is a coefficient or measurement, or “LINEAR” or “PARABOLIC Perm Perm Perm Req Req Perm Perm Exp Exp Page 52 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Variable Name Variable Label BRIDG 3.0.3 Mapping Type Controlled Terms, Codelist or Format Role CDISC Notes Core 1022 1023 LOG/LOG” when the result is a formula. Used for continuous or numeric results or findings in Perm standard format; copied in numeric format from QTSTRESC. QTSTRESN should store all numeric test results or findings. QTSTAT PerformedObservation. Char (ND) Record Qualifier Used to indicate a QT correction model was not done. Perm negationIndicator Should be null if a result exists in QTORRES. QTREASND PerformedObservation. Char Record Qualifier Describes why a measurement or test was not Perm negationReason performed. Used in conjunction with QTSTAT when correction is NOT DONE. QTNAM Vendor Name Organization.name Char Record Qualifier Name or identifier of the laboratory or vendor who Perm provided the test results. QTDRVFL Derived Flag Defer to Statistical Analysis Char (NY) Record Qualifier Used to indicate a derived record. The value should be Perm Modeling. Y or null. Records which represent the average of other records, or that do not come from the CRF, or are not as originally collected or received are examples of records that would be derived for the submission datasets. If QTDRVFL=Y, then QTORRES could be null, with QTSTRESC, and (if numeric) QTSTRESN having the derived value. * Indicates variable may be subject to controlled terminology (Parenthesis indicates CDISC/ NCI codelist code value). BRIDG Mapping values in italics are comments on why no mapping exists for that variable. 1024 6.2.2 Examples for ECG QT Corrections QTSTRESN 1025 1026 1027 1028 1029 Numeric Result/Finding in Standard Units Completion Status Reason QTc Not Performed PerformedObservationResult. value Num Result Qualifier The example consists of two Findings datasets: one for the ECG measurements (eg.xpt) and one for the QTc modeling information (qt.xpt). To tie the EG and QT datasets together will also require use of RELREC (relrec.xpt) to represent a dataset-to-dataset relationship that holds true for all subjects for all values of EGTESTCD and QTGRPID. eg.xpt Row DOMAIN USUBJID EGSEQ EGCAT EGTESTCD EGTEST EGORRES EGORRESU … EG P384QT204_001 1 INTERVAL QTCIAG1 QTCI Interval, Aggregate 1 345 msec … 1 EG P384QT204_001 2 INTERVAL QTCIAG2 QTCI Interval, Aggregate 2 350 msec … 2 EG P384QT204_001 3 INTERVAL QTCNAG QTCN Interval, Aggregate 353 msec … 3 EG P384QT204_002 1 INTERVAL QTCIAG1 QTCI Interval, Aggregate 1 372 msec … 4 EG P384QT204_002 2 INTERVAL QTCIAG2 QTCI Interval, Aggregate 2 374 msec … 5 EG P384QT204_002 3 INTERVAL QTCNAG QTCN Interval, Aggregate 377 msec … 6 EG P384QT204_003 1 INTERVAL QTCIAG1 QTCI Interval, Aggregate 1 401 msec … 7 EG P384QT204_003 2 INTERVAL QTCIAG2 QTCI Interval, Aggregate 2 411 msec … 8 EG P384QT204_003 3 INTERVAL QTCNAG QTCN Interval, Aggregate 414 msec … 9 1030 Page 53 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1031 qt.xpt Row DOMAIN USUBJID QTSEQ QTGRPID QT P384QT204_001 1 QTCIAG1 1 QT P384QT204_001 2 QTCIAG1 2 QT P384QT204_001 3 QTCIAG1 3 QT P384QT204_001 4 QTCIAG2 4 QT P384QT204_001 5 QTCIAG2 5 QT P384QT204_001 6 QTCIAG2 6 QT P384QT204_001 7 QTCNAG 7 QT P384QT204_001 8 QTCNAG 8 QT P384QT204_001 9 QTCNAG 9 QT P384QT204_001 10 QTCNAG 10 QT P384QT204_001 11 QTCNAG 11 QT P384QT204_002 1 QTCIAG1 12 QT P384QT204_002 2 QTCIAG1 13 QT P384QT204_002 3 QTCIAG1 14 QT P384QT204_002 4 QTCIAG2 15 QT P384QT204_002 5 QTCIAG2 16 QT P384QT204_002 6 QTCIAG2 17 QT P384QT204_002 7 QTCNAG 18 QT P384QT204_002 8 QTCNAG 19 QT P384QT204_002 9 QTCNAG 20 QT P384QT204_002 10 QTCNAG 21 QT P384QT204_002 11 QTCNAG 22 QT P384QT204_003 1 QTCIAG1 23 QT P384QT204_003 2 QTCIAG1 24 QT P384QT204_003 3 QTCIAG1 25 QT P384QT204_003 4 QTCIAG2 26 QT P384QT204_003 5 QTCIAG2 27 QT P384QT204_003 6 QTCIAG2 28 QT P384QT204_003 7 QTCNAG 29 QT P384QT204_003 8 QTCNAG 30 QT P384QT204_003 9 QTCNAG 31 QT P384QT204_003 10 QTCNAG 32 QT P384QT204_003 11 QTCNAG 33 1032 1033 QTTESTCD QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCCOEFB QTCCOEFC QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCCOEFB QTCCOEFC QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCCOEFB QTCCOEFC QTTEST QTORRES … QT Correction Method Description PARABOLIC LOG/LOG … QT Correction Formula QTC=QT/(RR^A) … QT Correction Coefficient A 0.432 … QT Correction Method Description LINEAR … QT Correction Formula QTC=QT+(A*(1-RR)) … QT Correction Coefficient A 0.154 … QT Correction Method Description RAUTAHARJU COR … QT Correction Formula QTC=QT+A-(B*(e^(C^HR)) … QT Correction Coefficient A 0.2425 … QT Correction Coefficient B 0.434 … QT Correction Coefficient C -0.0097 … QT Correction Method Description PARABOLIC LOG/LOG … QT Correction Formula QTC=QT/(RR^A) … QT Correction Coefficient A 0.374 … QT Correction Method Description LINEAR … QT Correction Formula QTC=QT+(A*(1-RR)) … QT Correction Coefficient A 0.132 … QT Correction Method Description RAUTAHARJU COR … QT Correction Formula QTC=QT+A-(B*(e^(C^HR)) … QT Correction Coefficient A 0.2425 … QT Correction Coefficient B 0.434 … QT Correction Coefficient C -0.0097 … QT Correction Method Description PARABOLIC LOG/LOG … QT Correction Formula QTC=QT/(RR^A) … QT Correction Coefficient A 0.412 … QTc Model Type LINEAR … QT Correction Formula QTC=QT+(A*(1-RR)) … QT Correction Coefficient A 0.158 … QTc Model Type RAUTAHARJU COR … QT Correction Formula QTC=QT+A-(B*(e^(C^HR)) … QT Correction Coefficient A 0.2425 … QT Correction Coefficient B 0.434 … QT Correction Coefficient C -0.0097 … relrec.xpt Row USUBJID RDOMAIN IDVAR IDVARVAL RELTYPE RELID EG EGTESTCD MANY EGEQ1 1 QT QTGRPID MANY EGEQ1 2 1034 1035 6.3 PK Assessments 1036 1037 An appropriate PK sampling schedule should be designed to adequately characterize the PK/PD concentration/response relationship for QT/QTc prolongation. The evaluation of the concentration-QT relationship involves individual subject responses instead of averaging the QT response across subjects at each time © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 54 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1038 1039 1040 point. Concentration-QTc analysis helps in predicting QT prolongation at doses beyond those used in TQT studies and may help determine whether excessive QT prolongation observed in a few subjects is due to high drug concentration or due to differential susceptibility in these subjects. 1041 1042 1043 1044 1045 1046 This concept map displays the sequence of steps involved if performing a PK/PD analysis within a TQT study. At each time point, data on PK drug concentrations and the relevant PD parameter such as QT are obtained to allow for PK/PD modeling. SDTM standard datasets (PC and PP) will be appropriate for capturing PK data from TQT study, as well. 1047 6.4 Hemodynamics/Vital Signs 1048 1049 1050 1051 1052 1053 1054 1055 1056 1057 1058 1059 Concept Map 8: PK/PD Analysis Steps and Timing This section focuses on the assessment of response to orthostatic challenge. Orthostatic, or postural, vital signs are used to assess the body’s response to position changes when a suspected drug might have the potential for altering autonomic tone and/or directly alter vascular tone or other physiological processes influencing cardiac output. Conditions leading to hypovolemia and autonomic abnormalities may result in a sudden drop in blood pressure, known as orthostatic hypotension, and result in impaired perfusion to the upper body. Assessment of response to orthostatic challenge might be conducted for two purposes within a TQT study: 1) as a screening mechanism to screen out potential subject that display an abnormal hemodynamic response; 2) as an assessment if the TQT study is being used for the secondary purpose of the careful and well-controlled assessment of the test drug’s influence on hemodynamic parameters. Severe orthostatic hypotension may exclude a patient from participation in TQT studies. Heart rate and blood pressure at rest and under orthostatic challenge might be evaluated as screening procedures for TQT studies. Additionally, the TQT study is well suited to collect information on potential changes in these parameters with drug treatment as a secondary study objective. Please note that resting heart rate and blood pressure would routinely be assessed during the study; however, assessment of orthostatic effect is optional. Page 55 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1060 1061 1062 1063 1064 1065 1066 Concept Map 9: Assessment of Orthostasis This concept map shows that the assessment of orthostasis is made based on changes in vital sign measurements measured within 3 minutes of standing or the inability to stand because the patient is severely orthostatic symptomatic and/or symptom assessment. This concept map is based on several key publications that are outlined below. Reference limits for orthostasis are also provided; the sponsor predefines these limits generally in the statistical analysis plan. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 56 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) American Autonomic Society / American Academy of Neurology Consensus Statement, 1996 16 • Provides a definitive definition of orthostasis based on either systolic or diastolic blood pressure decrease (see reference limits in table below) o Does not suggest frequency of measurement or time of first measurement within the 3 minute interval from standing • Does not provide definition based on heart rate increase Low, 1995 17 • Defined postural orthostatic tachycardia syndrome (POTS) as an increase in heart rate ≥30 bpm or absolute value ≥120 bpm within 5 minutes of standing associated with symptoms of orthostatic intolerance. Garland, 2007 18 • POTS as an increase in heart rate ≥30 bpm or absolute value ≥120 bpm but measured at 3 and 5 minutes in standing position, for study purposes, in the absence of orthostatic hypotension (i.e. SBP decrease >20 mmHg, DBP decrease > 10 mmHg) 1067 1068 Reference Limits for Orthostasis (All Ages) Basic Assessment of Orthostasis (time= 3 min) Orthostatic hypotension Decrease in systolic blood pressure when going from 5 minutes supine to 3 minutes standing of ≥20 mmHg or the inability to stand (systolic) quickly for the measurements due to symptoms of orthostasis. Orthostatic hypotension Decrease in diastolic blood pressure when going from 5 minutes supine to 3 minutes standing of ≥10 mmHg or the inability to stand (diastolic) quickly for the measurements due to symptoms of orthostasis. Orthostatic pulse Increase in heart rate when going from 5 minutes supine to 3 minutes standing of ≥30 bpm. (tachycardia) Detailed Assessment of Orthostasis (time = 0, 1, 2, 3 min) Orthostatic hypotension Decrease in systolic blood pressure when going from 5 minutes supine to any point during standing of ≥20 mmHg or the inability to (systolic) stand quickly for the measurements due to symptoms of orthostasis. Orthostatic hypotension (diastolic) Decrease in diastolic blood pressure when going from 5 minutes supine to any point during standing of ≥10 mmHg or the inability to stand quickly for the measurement due to symptoms of orthostasis. Orthostatic pulse (tachycardia) Increase in heart rate when going from 5 minutes supine to standing of ≥30 bpm. 1069 1070 1071 1072 Page 57 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1073 1074 1075 1076 1077 1078 1079 1080 1081 1082 1083 1084 1085 1086 1087 1088 1089 1090 1091 6.4.1 Example for Orthostatic Challenge Example 1 This example shows blood pressure and heart rate data collected to assess a subject’s response to orthostatic challenge (i.e., standing up quickly). These measurements are part of a pharmacodynamic profile, which involves repeating the challenge pre-dose and at certain time points post-dose. At each time point, these vital signs are measured while the subject is supine, and then three times after they stand up. The example shows data for two time points: one before the Day 1 dose of study treatment, and one an hour after the Day 1 dose. Some variables have been replaced with an ellipsis to save space. In this study, analysis of the orthostatic challenge data utilizes an approach to “baseline” data that is more complex than can be described by flagging a single value per subject per test using the VSBLFL variable, so the flag has not been populated. Since the absence of any records with VSBLFL populated would likely to be noted by validation software, the rationale for not using the flag would be included in the Study Data Reviewer Guide. Rows 1-12: Show vital signs for the pre-dose orthostatic challenge. All the data was collected at “PREDOSE” time point. The nominal planned elapsed time between this time point and the dose is 15 minutes (VSELTM = -P15M), although actual times range over 4 minutes. The replicate number (VSREPNUM) and the actual time of the measurement (VSDTC) distinguish among the measurements during this nominal time point. The measurements at the fourth replicate are flagged with the SDTM baseline flag (BLFL) since these were the last measurements before the start of dosing. The ADaM datasets will describe how measurements are used as baseline measurements in analyses. Rows 13-24: Show vital signs data for the time point “HOUR 1.” As for the pre-dose time point, the four replicate sets of measurements within the nominal time point were made over the course of four minutes. vs.xpt Row STUDYID DOMAIN USUBJID VSSEQ VSTESTCD VSTEST ABC VS ABC-001-001 7 SYSBP Systolic Blood Pressure 1 ABC VS ABC-001-001 8 DIABP Diastolic Blood Pressure 2 VSPOS SUPINE SUPINE VSCAT RESTING RESTING VSORRES 133 77 RESTING 70 3 ABC VS ABC-001-001 9 HR Heart Rate SUPINE 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC ABC VS VS VS VS VS VS VS VS VS VS VS VS VS VS VS VS VS VS VS ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 ABC-001-001 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 SYSBP DIABP HR SYSBP DIABP HR SYSBP DIABP HR SYSBP DIABP HR SYSBP DIABP HR SYSBP DIABP HR SYSBP Systolic Blood Pressure Diastolic Blood Pressure Heart Rate Systolic Blood Pressure Diastolic Blood Pressure Heart Rate Systolic Blood Pressure Diastolic Blood Pressure Heart Rate Systolic Blood Pressure Diastolic Blood Pressure Heart Rate Systolic Blood Pressure Diastolic Blood Pressure Heart Rate Systolic Blood Pressure Diastolic Blood Pressure Heart Rate Systolic Blood Pressure STANDING STANDING STANDING STANDING STANDING STANDING STANDING STANDING STANDING SUPINE SUPINE SUPINE STANDING STANDING STANDING STANDING STANDING STANDING STANDING © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 RESTING RESTING RESTING 127 70 74 125 77 78 121 88 76 117 81 95 115 78 80 116 79 80 121 VSORRESU mmHg mmHg BEATS/MIN BEATS/MIN mmHg mmHg BEATS/MIN mmHg mmHg BEATS/MIN mmHg mmHg BEATS/MIN mmHg mmHg BEATS/MIN mmHg mmHg BEATS/MIN mmHg mmHg BEATS/MIN mmHg … VSBLFL … … … … … … … … … … … … … … … … … … … … … … Page 58 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row STUDYID DOMAIN USUBJID VSSEQ VSTESTCD VSTEST ABC VS ABC-001-001 29 DIABP Diastolic Blood Pressure 23 ABC VS ABC-001-001 30 HR Heart Rate 24 VSPOS STANDING STANDING VSCAT VSORRES 78 82 VSORRESU mmHg BEATS/MIN … VSBLFL … … 1092 Row 1 (cont) 2 (cont) 3 (cont) 4 (cont) 5 (cont) 6 (cont) 7 (cont) 8 (cont) 9 (cont) 10 (cont) 11 (cont) 12 (cont) 13 (cont) 14 (cont) 15 (cont) 16 (cont) 17 (cont) 18 (cont) 19 (cont) 20 (cont) 21 (cont) 22 (cont) 23 (cont) 24 (cont) 1093 1094 1095 1096 1097 1098 1099 1100 1101 1102 VISIT VISITNUM VSDTC VSTPT VSTPTNUM VSELTM VSTPTREF VSRFTDTC VSREPNUM DAY 1 1 2014-04-01T7:45 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T7:45 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T7:45 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T7:47 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T7:47 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T7:47 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T7:48 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 2 DAY 1 1 2014-04-01T7:48 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 2 DAY 1 1 2014-04-01T7:48 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 2 DAY 1 1 2014-04-01T7:49 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 3 DAY 1 1 2014-04-01T7:49 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 3 DAY 1 1 2014-04-01T7:49 PREDOSE 1 -P15M DAY 1 DOSE 2014-04-01T8:00 3 DAY 1 1 2014-04-01T9:00 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T9:00 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T9:00 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T9:02 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T9:02 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T9:02 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 1 DAY 1 1 2014-04-01T9:03 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 2 DAY 1 1 2014-04-01T9:03 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 2 DAY 1 1 2014-04-01T9:03 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 2 DAY 1 1 2014-04-01T9:04 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 3 DAY 1 1 2014-04-01T9:04 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 3 DAY 1 1 2014-04-01T9:04 HOUR 1 2 P1H DAY 1 DOSE 2014-04-01T8:00 3 7 Data Analysis This section contains analysis data metadata and examples of analysis datasets used for the analysis and reporting of ECG data. Caveats and disclaimers: • The analysis data metadata examples are not intended to illustrate every possible variable that might be included in the analysis dataset. Sponsors should consult the ADaMIG for further guidance on additional variables that may be added. • The Source/Derivation column is for illustration purposes only and not intended to imply universally accepted definitions or derivations of variables. Algorithms are producer-defined and dependent on trial and analysis design. • The examples are for illustration purposes only and should not be viewed as a statement of the standards themselves. In addition, the examples are intended to illustrate content and not appearance. Page 59 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1103 7.1 ADEG Example 1104 7.1.1 Analysis Data Metadata 1105 1106 1107 1108 1109 1110 1111 1112 1113 1114 1115 1116 1117 1118 1119 The ADEG dataset is based on the ADaM Basic Data Structure (BDS). Analysis Dataset Metadata for ADEG Example Dataset Name Dataset Description Dataset Location ADEG Analysis Dataset for ECG Tests adeg.xpt Dataset Structure Key Variables of Dataset One record per subject per STUDYID, USUBJID, parameter per analysis PARAMCD, APERIOD, period per analysis visit per AVISIT, ATPT analysis time point Class of Dataset Documentation BDS adeg.sas In the metadata illustrated in the table below, the parameter identifier of “*ALL*” is used for variables expected to be consistent across analysis parameters and “*DEFAULT*” is used for parameters not otherwise specified in the Parameter Identifier column for that variable. The value of PARAMCD as the parameter identifier is used for variables that have metadata dependent on the analysis parameter. In this example, the study has a crossover design schema and ECG measurements were collected as a set of 3 replicates (each replicate approximately 1 minute apart) at time points 1 hr, 2 hr, etc. on Day -1 (baseline day) and Day 1 for each period, Period 1-4. The average of each set of 3 replicates is derived as a new record in ADaM and used for analysis. A time-matched baseline is used, in which the baseline for each period is the average of the 3 replicates at each time point on the baseline day, and corresponds to the average of the 3 replicates at each post-dose time point. QTcF and QTcI are the corrected QT measurements. QTcF was provided by the vendor and included in SDTM and QTcI is derived by the sponsor. Analysis Variable Metadata for ADEG Example Dataset Name ADEG Parameter Identifier *ALL* ADEG *ALL* ADEG ADEG *ALL* *ALL* Variable Name Variable Label STUDYID Study Identifier Unique Subject Identifier USUBJID TRTSEQA TRTA Actual Sequence of Treatments Actual Treatment Variable Type text Display Format $12 Codelist / Controlled Terms Source / Derivation ADSL.STUDYID text $20 ADSL.USUBJID text Placebo-Drug A 100 mgMoxifloxacin-Drug A 1 mg, Moxifloxacin-Placebo-Drug A 1 mg-Drug 1 100 mg, $50 Drug A 1 mg-Moxifloxacin-Drug A 100 mg-Placebo, Drug A 100 mg-Drug A 1 mgPlacebo-Moxifloxacin ADSL.TRTSEQA text © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 $15 Drug A 1 mg, Drug A 100 mg, Placebo, Moxifloxacin If APERIOD=1 then TRTA=ADSL.TRT01A, If APERIOD=2 then TRTA=ADSL.TRT02A If APERIOD=3 then Page 60 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Dataset Name ADEG ADEG ADEG ADEG ADEG Parameter Identifier *DEFAULT* QTCIAG PARAMCD *ALL* *ALL* Variable Name PARAM PARAM PARAMCD APERIODC APERIOD Variable Label Parameter Parameter Parameter Code Period (C) Period Variable Type text text text text integer Display Format Codelist / Controlled Terms Heart Rate (BEATS/MIN); PR Interval, Aggregate (msec); RR $50 Interval, Aggregate (msec); QT Interval, Aggregate (msec); QTCF Interval, Aggregate (msec); …. $50 $8 QTCI Interval, Aggregate (msec) HR, PRAG, RRAG, QTAG, QTCFAG, QTCIAG $8 PERIOD 1, PERIOD 2, PERIOD 3, PERIOD 4 8 1=PERIOD 1, 2=PERIOD 2, 3=PERIOD 3, 4=PERIOD 4 Source / Derivation TRTA=ADSL.TRT03A, If APERIOD=4 then TRTA=ADSL.TRT04A For records with a corresponding record in EG, populate with the value in EGTEST concatenated with the unit in EGSTRESU in parentheses. For records created to contain the QT Individual Correction result, populate with “QTCI Interval, Aggregate (msec)” One-to-one correspondence with PARAM. For records with a corresponding record in EG, PARAMCD=EG.EGTESTCD. If PARAM=”QTCI Interval, Aggregate (msec)”, then PARAMCD=”QTCIAG” Populate based on EG.VISITNUM. If VISITNUM=2 then APERIODC=”PERIOD 1”, If VISITNUM=4 then APERIODC=”PERIOD 2”, … Numeric version of APERIODC. Populate based on EG.VISITNUM. ADEG *ALL* AVISIT Analysis Visit text ADEG *ALL* AVISITN Analysis Visit (N) integer Page 61 Draft $11 8 DAY -1, DAY 1 -1=DAY -1, If VISITNUM=2 then AVISIT=’DAY -1’, Else if VISITNUM=3 then AVISIT=’DAY 1’ …. [Note, in this example visit windowing was not performed for analysis.] Numeric version of AVISIT. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Dataset Name ADEG Parameter Identifier *ALL* Variable Name ATPT Variable Label Analysis Time Point Variable Type text Display Format Codelist / Controlled Terms $40 ADEG *ALL* ATPTN Analysis Time Point (N) integer 8 ADEG *ALL* ADT Analysis Date integer yymmdd10. ADEG *ALL* ATM Analysis Time Integer time8. ADEG *ALL* ADY Analysis Relative Day Integer 8 ADEG ADEG *ALL* *ALL* AVISDY APERDAY Analysis Visit Day Analysis Nominal Period Day Integer integer © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 8 8 1=DAY 1 1 HR, 2 HR, 3 HR, 4 HR, 6 HR, 8 HR, 10 HR, 12 HR, 24 HR 1=1 HR, 2=2 HR, 3=3 HR, 4=4 HR, 6=6 HR, 8=8 HR, 10=10 HR, 12=12 HR, 24=24 HR Source / Derivation Populate based on EG.EGTPT. [Note, in this example visit windowing was not performed for analysis.] Numeric version of ATPT. Numeric date value from EG.EGDTC. Numeric time value from EG.EGDTC. EG.EGDY Sort by APERIOD, AVISITN, ATPTN, ADY. If first AVISITN then AVISDY=ADY; Retain AVISDY until next AVISITN. The visit day remains consistent within a given AVISITN even when the actual relative day crosses into the next calendar day. Populate with the numeric relative day within the Period based on Page 62 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Dataset Name Parameter Identifier Variable Name Variable Label Variable Type Display Format Codelist / Controlled Terms Source / Derivation AVISITN. [For example, “Period 2, Day 1” is APERDAY=1] ADEG *ALL* EGREPNUM ECG Replicate Number integer 8 EG.EGREPNUM. EG.EGSTRESN. ADEG *DEFAULT* AVAL Analysis Value float best12. ADEG QTCIAG, AVAL Analysis Value float best12. HR, PRAG, RRAG, QTAG, QTCFAG, … ADEG *ALL* DTYPE Derivation Type text $15 AVERAGE ADEG *ALL* ABLFL Baseline Record Flag text $1 Y ADEG *ALL* BASE Baseline Value float best12. Page 63 Draft Also create a derived record to contain the average of the three individual replicate values for ECG measurements collected at each time point. Calculate the individual QT correction (QTCIAG) based on the SAP Section x.x. A new derived record will be created for each replicate. Also create a derived record for the average of the individual QTcI replicate measurements at each time point. DTYPE=”AVERAGE” for records containing the average of the three replicate values for each time point. Otherwise, DTYPE is null. If AVISIT=“DAY -1” and DTYPE=“AVERAGE” then ABLFL=“Y”. If AVISIT=”DAY -1” and ABLFL=”Y” then BASE=AVAL; Else if AVISIT=”DAY 1” and DTYPE=”AVERAGE” and BASETYPE=“PERIOD 1, TIMEMATCH 1 HR” then BASE=AVAL where ABLFL=“Y” and APERIODC=”PERIOD 1” and ATPT=“1 HR”; Else If AVISIT=”DAY 1” and DTYPE=”AVERAGE” and BASETYPE=“PERIOD 1, TIMEMATCH 2 HR” then BASE=AVAL where ABLFL=”Y” and © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Dataset Name ADEG ADEG 1120 1121 1122 1123 Parameter Identifier *ALL* *ALL* Variable Name CHG Variable Label Change from Baseline Variable Type float text Display Format Codelist / Controlled Terms APERIODC=”PERIOD 1” and ATPT=“2 HR”, …. If AVISIT ne ‘DAY -1’ and DTYPE=AVERAGE, then CHG=AVAL – BASE. best12. $25 PERIOD 1, TIME-MATCH 1 HR, PERIOD 1, TIME-MATCH 2 HR PERIOD 1, TIME-MATCH 3 HR PERIOD 1, TIME-MATCH 4 HR …. BASETYPE Baseline Type text $1 Y Y ADEG QTAG, RRAG QTCCIFL Individ QT Correction Coefficient Flag ADEG *ALL* ECGPCFL ECG Matching PK Flag text $1 ADEG *ALL* EGSEQ Sequence Number integer 8 ADEG *ALL* EGLEAD Lead Location Used for Measurement text $10 ADEG *ALL* EGREFID ECG Reference ID text $10 Source / Derivation Populate BASETYPE with the APERIODC, “TIME-MATCH”, and ATPT. Populate with “Y” if the QTAG or RRAG value was used to calculate the QTCCOEFx for PARAMCD=”QTCIAG” Populate with “Y” if there is a matching nominal pk sample time point for the nominal ECG time point EG.EGSEQ. Set to missing if DTYPE=“AVERAGE”or PARAMCD=”QTCIAG”. EG.EGLEAD Set to missing if DTYPE=“AVERAGE” or PARAMCD=”QTCIAG”. EG.EGREFID Set to missing if DTYPE=“AVERAGE” or PARAMCD=”QTCIAG”. 7.1.2 Analysis Dataset The following example illustrates the analysis dataset (ADEG) defined above. To save space, not all variables in the dataset are included. The SDTM dataset (EG) in Section 6.1.2 Example 1 contains the input source records for the ADaM dataset. Rows 1-3, 5-7, Show records from SDTM containing the three individual replicate values for QTAG, collected at approximately 1-minute intervals. EGREPNUM identifies the replicate number. These records support traceability of the derived average record used for analysis. EGSEQ values 9-11, 13-15: from SDTM are included and support traceability to records in SDTM EG. Rows 4, 8, 12: Shows the average of the three replicates. In this example, the average is derived in ADaM so DTYPE is assigned “AVERAGE”. If the sponsor © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 64 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Rows 9-12: Row 10: Row 16: 1124 1125 derived the average in SDTM, then include the SDTM derived flag, EGDRVFL in ADaM so average records can be identified when both individual and average records are submitted in the dataset. ABLFL=“Y” indicates these records are used as baseline values. For these time-matched baselines, BASETYPE identifies the definition of baseline and is different for each Period and time point. When BASETYPE is used for any PARAM within a dataset, it should be non-null for all records of that dataset. The AVAL at PERIOD 1 DAY -1, 1 HR (row 4), is used as the BASE for the post-baseline measurement at PERIOD 1 DAY 1, 1 HR (row 16). Similarly, the AVAL at PERIOD 1 DAY -1, 2 HR (row 8), is used as the BASE for the post-baseline measurement at PERIOD 1 DAY 1, 2 HR (row not shown). Though not illustrated, ABLFL=“Y” would also be assigned similarly for all PERIOD 1-4, DAY -1 time points. Shows difference in ADY, AVISDY for the 24 HR time point. The variable AVISDY is different from ADY. The variable AVISDY is the nominal relative study day for the Period Day, and remains consistent regardless of whether the actual relative day crosses into the next calendar day. For AVISIT=DAY -1, the AVISDY=-1 for 1 HR, 2 HR, … 24 HR, but the actual relative day ADY=1 for the 24 HR time point because it crossed into the next calendar day. Similarly, for Period 2 Day -1 (not shown), the AVISDY=3 for all time points, but ADY=4 for the 24 HR time point. Shows a record where the EGLEAD used is not the planned primary lead. The planned primary lead is LEAD II, however this measurement was taken from LEAD V. The planned primary lead is specified in the proposed terminology for Trial Summary parameters in Section 4.1.4. Shows a derived record for the average of the three post-baseline replicates at PERIOD 1 DAY 1, 1 HR. The change from baseline is calculated as CHG = AVAL – BASE. This record is flagged as having a matching PK concentration for this time point (ECGPCFL=“Y”). adeg.xpt Row STUDYID USUBJID 1 2 3 4 5 6 7 8 9 10 11 12 TRTSEQA Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Additional records for 3 HR, 4 HR, etc…. Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug STUDY01 2324-P0001 A 100 mg – Moxifloxacin Page 65 Draft TRTA PARAMCD Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG PARAM APERIODC APERIOD AVISIT AVISITN ATPT ATPTN ADT QT Interval, PERIOD 1 1 DAY -1 -1 1 HR 1 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 1 HR 1 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 1 HR 1 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 1 HR 1 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 2 HR 2 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 2 HR 2 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 2 HR 2 2014-03-22 Aggregate (msec) QT Interval, PERIOD 1 1 DAY -1 -1 2 HR 2 2014-03-22 Aggregate (msec) QT Interval, Aggregate (msec) QT Interval, Aggregate (msec) QT Interval, Aggregate (msec) QT Interval, Aggregate (msec) PERIOD 1 1 DAY -1 -1 24 HR 24 2014-03-23 PERIOD 1 1 DAY -1 -1 24 HR 24 2014-03-23 PERIOD 1 1 DAY -1 -1 24 HR 24 2014-03-23 PERIOD 1 1 DAY -1 -1 24 HR 24 2014-03-23 © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row STUDYID USUBJID 13 STUDY01 2324-P0001 14 STUDY01 2324-P0001 15 STUDY01 2324-P0001 16 STUDY01 2324-P0001 TRTSEQA Drug A 1 mg – Placebo - Drug A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug A 100 mg – Moxifloxacin Drug A 1 mg – Placebo - Drug A 100 mg – Moxifloxacin TRTA PARAMCD Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG Drug A 1 mg QTAG PARAM APERIODC APERIOD AVISIT AVISITN ATPT ATPTN ADT QT Interval, PERIOD 1 1 DAY 1 1 1 HR 1 2014-03-23 Aggregate (msec) QT Interval, PERIOD 1 1 DAY 1 1 1 HR 1 2014-03-23 Aggregate (msec) QT Interval, PERIOD 1 1 DAY 1 1 1 HR 1 2014-03-23 Aggregate (msec) QT Interval, PERIOD 1 1 DAY 1 1 1 HR 1 2014-03-23 Aggregate (msec) 1126 Row ATM ADY AVISDY APERDAY EGREPNUM AVAL 1 (cont) 10:00:21 -1 -1 -1 1 440 2 (cont) 10:01:35 -1 -1 -1 2 389 3 (cont) 10:02:14 -1 -1 -1 3 377 4 (cont) -1 -1 -1 5 (cont) 11:00:21 -1 -1 -1 1 379 6 (cont) 11:01:31 -1 -1 -1 2 402 7 (cont) 11:02:40 -1 -1 -1 3 356 8 (cont) -1 -1 -1 402 379 DTYPE AVERAGE AVERAGE ABLFL BASE CHG Y Y BASETYPE ECGPCFL EGSEQ EGLEAD EGREFID QTCCIFL PERIOD 1, TIME4 LEAD II 1234933 Y MATCH 1 HR PERIOD 1, TIMEMATCH 1 HR 8 LEAD II 1289032 Y PERIOD 1, TIMEMATCH 1 HR PERIOD 1, TIMEMATCH 1 HR PERIOD 1, TIMEMATCH 2 HR PERIOD 1, TIMEMATCH 2 HR PERIOD 1, TIMEMATCH 2 HR PERIOD 1, TIMEMATCH 2 HR 402 379 12 LEAD II 1343131 Y Y 16 LEAD II 1342003 Y 20 LEAD II 1340323 Y 24 LEAD II 1343459 Y Y Additional records for 3 HR, 4 HR, etc…. 9 (cont) 09:00:07 1 -1 -1 1 386 10 (cont) 09:01:38 1 -1 -1 2 387 11 (cont) 09:02:34 1 -1 -1 3 388 12 (cont) 1 -1 -1 13 (cont) 10:01:33 1 1 1 1 397 14 (cont) 10:02:35 1 1 1 2 403 15 (cont) 10:03:34 1 1 1 3 377 16 (cont) 1 1 1 387 392 AVERAGE AVERAGE © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Y 387 402 -10 PERIOD 1, TIMEMATCH 24 HR PERIOD 1, TIMEMATCH 24 HR PERIOD 1, TIMEMATCH 24 HR PERIOD 1, TIMEMATCH 24 HR PERIOD 1, TIMEMATCH 1 HR PERIOD 1, TIMEMATCH 1 HR PERIOD 1, TIMEMATCH 1 HR PERIOD 1, TIMEMATCH 1 HR 100 LEAD II 1347893 Y 104 LEAD V 1348832 Y 108 LEAD II 1348814 Y Y 112 LEAD II 1234933 116 LEAD II 1289032 120 LEAD II 1343131 Y Page 66 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1127 7.2 ADQT Example 1128 1129 1130 1131 1132 1133 1134 The ADQT dataset contains information about the QT correction model(s) used in the study. It is created when the QT correction formula and coefficients are derived by the sponsor or vendor, and not created for commonly used historical correction models such as Bazett or Fridericia. The model information could be for one or more specific models specified in the protocol, or one or more ‘best fit’ models from a series of models. Additional details regarding the kinds of QT correction models can be found in Section 2.3.4 and Section 6.2. If the sponsor derives the QT corrections, the calculations would usually be performed at the ADaM stage, and the model information would be submitted in ADaM ADQT and not in SDTM QT. If the vendor derives the QT corrections, the model information would be stored in the SDTM QT domain as specified in Section 6.2, and may be carried forward into ADQT, depending on the sponsor’s needs. The ADQT dataset is based on the ADaM Basic Data Structure (BDS). 1135 7.2.1 Analysis Data Metadata 1136 Analysis Dataset Metadata for ADQT Example 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 Dataset Name Dataset Description Dataset Location ADQT Analysis Dataset for ECG QTc Model Data adqt.xpt Key Variables of Dataset Dataset Structure Class of Dataset Documentation BDS adqt.sas One record per subject, STUDYID, USUBJID, per QT model, per QT AGRPID, PARAMCD model parameter In the metadata illustrated in the table below, the parameter identifier of “*ALL*” is used for variables expected to be consistent across analysis parameters, “*DEFAULT*” is used for parameters not otherwise specified in the Parameter Identifier column for that variable. The value of PARAMCD as the parameter identifier is used for variables that have metadata dependent on the analysis parameter. In this example, the ADQT dataset contains the top two ‘best fit’ QT correction models for each subject and a historical model used for all subjects. For each subject, the baseline QT/RR data was fit into twelve different protocol-specified potential models, from which the top two best fit models will be used for calculating QTcI corrections. For the historical model, the QT formula and coefficients are the same across all subjects; however, the sponsor chose to include the model information in the dataset for each subject. Analysis Variable Metadata for ADQT Example Dataset Name ADQT Parameter Identifier *ALL* ADQT *ALL* USUBJID ADQT QTCDESC PARAM ADQT QTCFORM PARAM Page 67 Draft Variable Name Variable Label STUDYID Study Identifier Unique Subject Identifier Variable Type text Display Format Codelist / Controlled Terms Source / Derivation $12 ADSL.STUDYID text $20 ADSL.USUBJID Parameter text $50 QT Correction Method Description Parameter text $50 QT Correction Formula For records created to contain the QT Correction method, populate with “QT Correction Method Description” For records created to contain the QT Correction formula, populate with “QT Correction Formula” © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Dataset Name 1148 1149 1150 1151 1152 1153 1154 1155 1156 1157 1158 1159 1160 Parameter Identifier Variable Name Variable Label Variable Type Display Format Codelist / Controlled Terms ADQT QTCCOEFx PARAM Parameter text $50 QT Correction Coefficient x ADQT *ALL* PARAMCD Parameter Code text $8 ADQT QTCCOEFx AVAL Analysis Value float best12. ADQT QTCDESC AVALC Analysis Value (C) text $50 ADQT QTCFORM AVALC Analysis Value (C) text $50 ADQT *ALL* AGRPID Group ID text $8 QTCDESC, QTCFORM, QTCCOEFx QTCIAG1, QTCIAG2, QTCNAG Source / Derivation For records created to contain the QT correction coefficient x, populate with “QT Correction Coefficient x” One-to-one correspondence with PARAM Create one or more records containing the individual or historical correction coefficient used to calculate the QTc values specified in AGRPID Set AVALC to the model description used to calculate the QTc values specified in AGRPID Set AVALC to the model formulas used to calculate the QTc values specified in AGRPID Based on the protocol, assign AGRPID to the name of the QTc parameters to be calculated for each subject 7.2.2 Analysis Dataset In this example, the variable AGRPID identifies the QT corrected parameter code (PARAMCD) in ADEG whose values are based on the model information. Variables that do not apply for the AVAL are not included in the dataset (e.g. AVISIT, ADT, ATM, etc.). Rows 1-3: Rows 4-6: Rows 7-11: Rows 12-22: Row 1 2 3 4 5 6 Shows records containing model information for the first best fit model for USUBJID=2324-P0001. This model contains one correction coefficient 0.266, which was used in the LINEAR model formula to calculate the subject’s QTc values for PARAMCD=”QTCIAG1” in ADEG. Shows records containing model information for the second best fit model for this subject. This model contains one correction coefficient 0.532, which was used in the PARABOLIC model formula to calculate the subject’s QTc values for PARAMCD=”QTCIAG2” in ADEG. Shows records containing model information for the historical model, RAUTAHARJU COR, that contains three correction coefficients. The coefficients 0.2425, 0.434, and -0.0097 are constant values across all subjects and used to calculate PARAMCD=QTCNAG in ADEG. Shows records for USUBJID=2324-P0002. The top two best fit models, LOGARITHMIC and LINEAR, and the correction coefficients are different than for USUBJID=2324-P0001. However, the historical model information is the same. STUDYID STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 USUBJID 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 PARAMCD QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA PARAM QT Correction Method Description QT Correction Formula QT Correction Coefficient A QT Correction Method Description QT Correction Formula QT Correction Coefficient A © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 AVAL AVALC LINEAR QTC=QT+(QTCCOEFA*(1-RR)) 0.266 PARABOLIC QTC=QT/(RR^QTCCOEFA) 0.532 AGRPID QTCIAG1 QTCIAG1 QTCIAG1 QTCIAG2 QTCIAG2 QTCIAG2 Page 68 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row 7 STUDYID STUDY01 USUBJID 2324-P0001 PARAMCD QTCDESC PARAM QT Correction Method Description 8 STUDY01 2324-P0001 QTCFORM QT Correction Formula 9 10 11 12 13 14 15 16 17 18 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 STUDY01 2324-P0001 2324-P0001 2324-P0001 2324-P0002 2324-P0002 2324-P0002 2324-P0002 2324-P0002 2324-P0002 2324-P0002 QTCCOEFA QTCCOEFB QTCCOEFC QTCDESC QTCFORM QTCCOEFA QTCDESC QTCFORM QTCCOEFA QTCDESC QT Correction Coefficient A QT Correction Coefficient B QT Correction Coefficient C QT Correction Method Description QT Correction Formula QT Correction Coefficient A QT Correction Method Description QT Correction Formula QT Correction Coefficient A QT Correction Method Description 2324-P0002 QTCFORM QT Correction Formula 2324-P0001 2324-P0001 2324-P0001 QTCCOEFA QTCCOEFB QTCCOEFC QT Correction Coefficient A QT Correction Coefficient B QT Correction Coefficient C 19 20 21 22 STUDY01 STUDY01 STUDY01 AVAL AVALC RAUTAHARJU COR QTC=QT+QTCCOEFA(QTCCOEFB*(e^(QTCCOEFC^HR)) 0.2425 0.434 -0.0097 LOGARITHMIC QTC=QT-QTCCOEFA*ln(RR) 0.116 LINEAR QTC=QT+(QTCCOEFA*(1-RR)) 0.387 RAUTAHARJU COR QTC=QT+QTCCOEFA(QTCCOEFB*(e^(QTCCOEFC^HR)) 0.2425 0.434 -0.0097 AGRPID QTCNAG QTCNAG QTCNAG QTCNAG QTCNAG QTCIAG1 QTCIAG1 QTCIAG1 QTCIAG2 QTCIAG2 QTCIAG2 QTCNAG QTCNAG QTCNAG QTCNAG QTCNAG 1161 7.3 Baseline Alternatives and Definition of Treatment Differences 1162 1163 1164 1165 In this section, four different baseline alternatives, including the possibility of no baseline, are described. For each baseline method, the resulting definition of treatment differences is described. Note that this is not an exhaustive list of possibilities. For example, triple-delta (∆∆∆QTc) definitions are possible where both lead-in day ECGs are collected at matched time points to the time points of collection on the treatment day and an ECG replicate set is collected immediately before treatment administration (and at the same time point on the lead-in day). 1166 7.3.1 Time-Matched Baseline; Double-Delta Treatment Difference 1167 1168 1169 1170 For time-matched baseline, the baseline for each period is the average of values at a time point on the baseline day (Day -1) that corresponds to the post-dose time point. ECGs are collected or extracted from continuous recording in replicate sets (usually 3 replicates about a minute or so apart) at each bj and Xij. The average of the replicates is presented in the table below. 1171 1172 1173 1174 For crossover design, ΔΔQTcij is computed for each subject: ΔΔQTcij = �Xij − bj � and n=time point. Page 69 Draft Drug A − �Xij − bj � Placebo where i=1, 2, … d, j=1, 2, … n; d=days postdose © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 Row USUBJID TRTA PARAMCD APERIOD AVISIT ATPT AVAL* ABLFL BASE* 1 DAY -1 1 HR b1(Drug A) Y b1(Drug A) 1 2324-P0001 Drug A 1 mg QTCFAG 1 DAY -1 2 HR b2(Drug A) Y b2(Drug A) 2 2324-P0001 Drug A 1 mg QTCFAG 1 DAY -1 3 HR b3(Drug A) Y b3(Drug A) 3 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 1 HR X11(Drug A) b1(Drug A) 4 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 2 HR X12(Drug A) b2(Drug A) 5 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 3 HR X13(Drug A) b3(Drug A) 6 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 1 HR X21(Drug A) b1(Drug A) 7 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 2 HR X22(Drug A) b2(Drug A) 8 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 3 HR X23(Drug A) b3(Drug A) 9 2324-P0001 Drug A 1 mg QTCFAG Placebo QTCFAG 2 DAY -1 1 HR b1(Placebo) Y b1(Placebo) 10 2324-P0001 Placebo QTCFAG 2 DAY -1 2 HR b2(Placebo) Y b2(Placebo) 11 2324-P0001 Placebo QTCFAG 2 DAY -1 3 HR b3(Placebo) Y b3(Placebo) 12 2324-P0001 Placebo QTCFAG 2 DAY 1 1 HR X11(Placebo) b1(Placebo) 13 2324-P0001 Placebo QTCFAG 2 DAY 1 2 HR X12(Placebo) b2(Placebo) 14 2324-P0001 Placebo QTCFAG 2 DAY 1 3 HR X13(Placebo) b3(Placebo) 15 2324-P0001 Placebo QTCFAG 2 DAY 2 1 HR X21(Placebo) b1(Placebo) 16 2324-P0001 Placebo QTCFAG 2 DAY 2 2 HR X22(Placebo) b2(Placebo) 17 2324-P0001 Placebo QTCFAG 2 DAY 2 3 HR X23(Placebo) b3(Placebo) 18 2324-P0001 *Values in these columns are mathematical expressions. See the next table for example submission values. ���������� For a parallel design, (Xij – bj) would be averaged across subjects: ΔΔQTcij = �X ıȷ − bȷ � 7.3.1.1 Analysis Dataset Example for Time-Matched Baseline CHG* (X11-b1)Drug A (X12-b2)Drug A (X13-b3)Drug A (X21-b1)Drug A (X22-b2)Drug A (X23-b3)Drug A (X11-b1)Placebo (X12-b2)Placebo (X13-b3)Placebo (X21-b1)Placebo (X22-b2)Placebo (X23-b3)Placebo Drug A BASETYPE PERIOD 1, TIME-MATCH 1 HR PERIOD 1, TIME-MATCH 2 HR PERIOD 1, TIME-MATCH 3 HR PERIOD 1, TIME-MATCH 1 HR PERIOD 1, TIME-MATCH 2 HR PERIOD 1, TIME-MATCH 3 HR PERIOD 1, TIME-MATCH 1 HR PERIOD 1, TIME-MATCH 2 HR PERIOD 1, TIME-MATCH 3 HR PERIOD 2, TIME-MATCH 1 HR PERIOD 2, TIME-MATCH 2 HR PERIOD 2, TIME-MATCH 3 HR PERIOD 2, TIME-MATCH 1 HR PERIOD 2, TIME-MATCH 2 HR PERIOD 2, TIME-MATCH 3 HR PERIOD 2, TIME-MATCH 1 HR PERIOD 2, TIME-MATCH 2 HR PERIOD 2, TIME-MATCH 3 HR ���������� − �X ıȷ − bȷ � Placebo The below table illustrates time-matched baseline and post-baseline records with example values. Rows 1-3: Row 4: Row 4, 8: Row 12, 16: Row 1 2 3 4 5 6 USUBJID 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 Shows records from SDTM containing the three individual replicate values for QTCFAG, collected at approximately 1-minute intervals. EGREPNUM identifies the replicate number. These records support traceability of the derived average record used for analysis. Shows a derived record with the average of the three replicates. ABLFL=”Y” indicates these records are used as baseline values. For these time-matched baselines, BASETYPE identifies the definition of baseline, and is different for each Period and time point. The AVAL at PERIOD 1 DAY -1, 1 HR (row 4), is used as the BASE for the postbaseline measurement at PERIOD 1 DAY 1, 1 HR (row 12). Similarly, the AVAL at PERIOD 1 DAY -1, 2 HR (row 8), is used as the BASE for the post-baseline measurement at PERIOD 1 DAY 1, 2 HR (row 16). Though not illustrated, ABLFL=”Y” would also be assigned similarly for all other PERIOD 1-4, DAY -1 time points. The change from baseline, CHG = AVAL – BASE, is calculated for the average of the replicate values. BASETYPE identifies which baseline value corresponds to the value of BASE in that row. TRTA PARAMCD APERIOD AVISIT Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY -1 ATPT EGREPNUM AVAL DTYPE ABLFL BASE CHG BASETYPE 1 HR 1 387 PERIOD 1, TIME-MATCH 1 HR 1 HR 2 357 PERIOD 1, TIME-MATCH 1 HR 1 HR 3 364 PERIOD 1, TIME-MATCH 1 HR 1 HR 369 AVERAGE Y 369 PERIOD 1, TIME-MATCH 1 HR 2 HR 1 361 PERIOD 1, TIME-MATCH 2 HR 2 HR 2 376 PERIOD 1, TIME-MATCH 2 HR © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 70 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row 7 8 9 10 11 12 13 14 15 16 1192 1193 1194 1195 1196 1197 1198 1199 1200 1201 USUBJID 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 TRTA PARAMCD APERIOD AVISIT Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY -1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 ATPT EGREPNUM AVAL DTYPE ABLFL BASE CHG BASETYPE 2 HR 3 409 PERIOD 1, TIME-MATCH 2 HR 2 HR 382 AVERAGE Y 382 PERIOD 1, TIME-MATCH 2 HR 1 HR 1 400 PERIOD 1, TIME-MATCH 1 HR 1 HR 2 388 PERIOD 1, TIME-MATCH 1 HR 1 HR 3 379 PERIOD 1, TIME-MATCH 1 HR 1 HR 389 AVERAGE 369 20 PERIOD 1, TIME-MATCH 1 HR 2 HR 1 396 PERIOD 1, TIME-MATCH 2 HR 2 HR 2 384 PERIOD 1, TIME-MATCH 2 HR 2 HR 3 402 PERIOD 1, TIME-MATCH 2 HR 2 HR 394 AVERAGE 382 12 PERIOD 1, TIME-MATCH 2 HR 7.3.2 Time-Averaged Baseline; Double-Delta Treatment Difference For time-averaged baseline, the baseline for each period is the average of all values on the baseline day (e.g. Day -1, 1 hr, 2 hr, 3 hr, 4 hr, etc.). ECGs are collected or extracted from continuous recording in replicate sets (usually 3 replicates about a minute or so apart) at each bj and Xij. The average of the replicates is presented in the table below. For crossover design, ΔΔQTcij is computed for each subject: ΔΔQTcij = �Xij − bavg � d = days postdose and n = time point. Drug A Row USUBJID TRTA PARAMCD APERIOD AVISIT ATPT AVAL* ABLFL 1 DAY -1 BASELINE bavg(Drug A) Y 1 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 1 HR X11(Drug A) 2 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 2 HR X12(Drug A) 3 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 3 HR X13(Drug A) 4 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 1 HR X21(Drug A) 5 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 2 HR X22(Drug A) 6 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 3 HR X23(Drug A) 7 2324-P0001 Drug A 1 mg QTCFAG Placebo QTCFAG 2 DAY -1 BASELINE bavg (Placebo) Y 8 2324-P0001 Placebo QTCFAG 2 DAY 1 1 HR X11(Placebo) 9 2324-P0001 Placebo QTCFAG 2 DAY 1 2 HR X12(Placebo) 10 2324-P0001 Placebo QTCFAG 2 DAY 1 3 HR X13(Placebo) 11 2324-P0001 Placebo QTCFAG 2 DAY 2 1 HR X21(Placebo) 12 2324-P0001 Placebo QTCFAG 2 DAY 2 2 HR X22(Placebo) 13 2324-P0001 Placebo QTCFAG 2 DAY 2 3 HR X23(Placebo) 14 2324-P0001 *Values in these columns are mathematical expressions. See the next table for example submission values. Page 71 Draft − �Xij − bavg � BASE* bavg(Drug A) bavg(Drug A) bavg(Drug A) bavg(Drug A) bavg(Drug A) bavg(Drug A) bavg(Drug A) bavg(Placebo) bavg(Placebo) bavg(Placebo) bavg(Placebo) bavg(Placebo) bavg(Placebo) bavg(Placebo) Placebo CHG* (X11-bavg)Drug A (X12-bavg)Drug A (X13-bavg)Drug A (X21-bavg)Drug A (X22-bavg)Drug A (X23-bavg)Drug A (X11-bavg)Placebo (X12-bavg)Placebo (X13-bavg)Placebo (X21-bavg)Placebo (X22-bavg)Placebo (X23-bavg)Placebo where bavg = ∑ bj /n; i=1, 2, … d, j=1, 2, … n; BASETYPE PERIOD 1, TIME-AVG PERIOD 1, TIME-AVG PERIOD 1, TIME-AVG PERIOD 1, TIME-AVG PERIOD 1, TIME-AVG PERIOD 1, TIME-AVG PERIOD 1, TIME-AVG PERIOD 2, TIME-AVG PERIOD 2, TIME-AVG PERIOD 2, TIME-AVG PERIOD 2, TIME-AVG PERIOD 2, TIME-AVG PERIOD 2, TIME-AVG PERIOD 2, TIME-AVG © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1202 1203 1204 1205 1206 1207 1208 1209 1210 1211 1212 1213 1214 1215 ������������� For a parallel design, (Xij – bavg) would be averaged across subjects: ΔΔQTcij = �X ıȷ − bavg � Drug A The below table illustrates time-averaged baseline and post-baseline records with example values. Rows 1-3: Rows 4, 8: Rows 9: Row 13, 17: Row 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1216 1217 1218 1219 1220 USUBJID 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 ������������� − �X ıȷ − bavg � Placebo Shows three individual replicate values for QTCFAG, collected at approximately 1-minute intervals. EGREPNUM identifies the replicate number. Shows a derived record with the average of the three replicates. The baseline for PERIOD 1 is based on the average of the replicate averages for 1 HR (row 4), 2 HR (row 8), and 3 HR, 4 HR, 6 HR, 8 HR, 10 HR, 12 HR, and 24 HR replicate averages (not shown). ABLFL=”Y” indicates this contains the baseline value for this BASETYPE. The AVAL (“370”) is the value of BASE (row 9) and the value of BASE for all rows that have the same BASETYPE (row 13, 17, and additional rows not shown). The records for BASETYPE=”PERIOD 2, TIME-AVG” are not shown. The change from baseline, CHG = AVAL – BASE, is calculated for the average of the replicate values. TRTA PARAMCD APERIOD AVISIT ATPT EGREPNUM AVAL DTYPE ABLFL BASE CHG BASETYPE Drug A 1 mg QTCFAG 1 DAY -1 1 HR 1 387 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 1 HR 2 357 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 1 HR 3 364 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 1 HR 369 AVERAGE PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 2 HR 1 361 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 2 HR 2 376 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 2 HR 3 409 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 2 HR 382 AVERAGE PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY -1 BASELINE 370 AVERAGE Y 370 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 1 400 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 2 388 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 3 379 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 389 AVERAGE 370 19 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 1 396 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 2 384 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 3 402 PERIOD 1, TIME-AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 394 AVERAGE 370 24 PERIOD 1, TIME-AVG 7.3.3 Predose Averaged Baseline; Double-Delta Treatment Difference For predose averaged baseline, ECGs are collected or extracted as replicate sets (usually 3 replicates about a minute or less apart) at predose in close temporal proximity to treatment administration (e.g., 15 min intervals and immediately before treatment administration on the same day of treatment administration) and as replicate sets (usually 3 replicates about a minute or so apart) at each Xij post dose. The average of the replicates is presented in the table below. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 72 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1221 1222 1223 1224 1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 For crossover design, ΔΔQTcij is computed for each subject: ∆∆QTcij = �Xij − b0 � days postdose, k = number of predose replicates, n = time point. Drug A − �Xij − b0 � Placebo Row USUBJID TRTA PARAMCD APERIOD AVISIT ATPT AVAL* BASE* ABLFL CHG* 1 DAY 1 BASELINE b0(Drug A) b0(Drug A) Y 1 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 1 HR X11(Drug A) b0(Drug A) (X11-b0)Drug A 2 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 2 HR X12(Drug A) b0(Drug A) (X12-b0)Drug A 3 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 1 3 HR X13(Drug A) b0(Drug A) (X13-b0)Drug A 4 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 1 HR X21(Drug A) b0(Drug A) (X21-b0)Drug A 5 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 2 HR X22(Drug A) b0(Drug A) (X22-b0)Drug A 6 2324-P0001 Drug A 1 mg QTCFAG 1 DAY 2 3 HR X23(Drug A) b0(Drug A) (X23-b0)Drug A 7 2324-P0001 Drug A 1 mg QTCFAG Placebo QTCFAG 2 DAY 1 BASELINE b0(Placebo) b0(Placebo) Y 8 2324-P0001 Placebo QTCFAG 2 DAY 1 1 HR X11(Placebo) b0(Placebo) (X11-b0)Placebo 9 2324-P0001 Placebo QTCFAG 2 DAY 1 2 HR X12(Placebo) b0(Placebo) (X12-b0)Placebo 10 2324-P0001 Placebo QTCFAG 2 DAY 1 3 HR X13(Placebo) b0(Placebo) (X13-b0)Placebo 11 2324-P0001 Placebo QTCFAG 2 DAY 2 1 HR X21(Placebo) b0(Placebo) (X21-b0)Placebo 12 2324-P0001 Placebo QTCFAG 2 DAY 2 2 HR X22(Placebo) b0(Placebo) (X22-b0)Placebo 13 2324-P0001 Placebo QTCFAG 2 DAY 2 3 HR X23(Placebo) b0(Placebo) (X23-b0)Placebo 14 2324-P0001 *Values in these columns are mathematical expressions. See the next table for example submission values. ����������� For a parallel design, (Xij – bj) would be averaged across subjects: ΔΔQTcij = �X ıȷ − b0 � Drug A where b0 = ∑ bj /k; i=1, 2, … d, j=1, 2, … n; d = BASETYPE PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG PERIOD 2, PREDOSE AVG PERIOD 2, PREDOSE AVG PERIOD 2, PREDOSE AVG PERIOD 2, PREDOSE AVG PERIOD 2, PREDOSE AVG PERIOD 2, PREDOSE AVG PERIOD 2, PREDOSE AVG ����������� − �X ıȷ − b0 � Placebo The below table is an example ADEG dataset illustrating predose averaged baseline and post-baseline records. Rows 4, 8, 12: Shows the average of three replicate QTCFAG measurements taken within one hour prior to dose which are used to calculate the baseline. Rows 13: Shows a derived record with the average of rows 4, 8, and 12. This value is the predose average baseline (ABLFL=”Y”). Rows 17, 21: The change from baseline, CHG = AVAL – BASE, is calculated for the average of the replicate values. Row 1 2 3 4 5 6 7 8 USUBJID 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 Page 73 Draft TRTA PARAMCD APERIOD AVISIT Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 Drug A 1 mg QTCFAG 1 DAY 1 ATPT -45 MIN -45 MIN -45 MIN -45 MIN -30 MIN -30 MIN -30 MIN -30 MIN EGREPNUM AVAL DTYPE BASE ABLFL CHG BASETYPE 1 380 PERIOD 1, PREDOSE AVG 2 381 PERIOD 1, PREDOSE AVG 3 382 PERIOD 1, PREDOSE AVG 381 AVERAGE PERIOD 1, PREDOSE AVG 1 378 PERIOD 1, PREDOSE AVG 2 379 PERIOD 1, PREDOSE AVG 3 380 PERIOD 1, PREDOSE AVG 379 AVERAGE PERIOD 1, PREDOSE AVG © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Row 9 10 11 12 13 14 15 16 17 18 19 20 21 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 USUBJID 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 2324-P0001 TRTA PARAMCD APERIOD AVISIT ATPT EGREPNUM AVAL DTYPE BASE ABLFL CHG BASETYPE Drug A 1 mg QTCFAG 1 DAY 1 -15 MIN 1 395 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 -15 MIN 2 393 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 -15 MIN 3 394 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 -15 MIN 394 AVERAGE PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 BASELINE 385 AVERAGE 385 Y PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 1 400 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 2 388 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 3 379 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 1 HR 389 AVERAGE 385 4 PERIOD 1, PREDOSE AVG PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 1 396 Drug A 1 mg QTCFAG 1 DAY 1 2 HR 2 384 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 3 402 PERIOD 1, PREDOSE AVG Drug A 1 mg QTCFAG 1 DAY 1 2 HR 394 AVERAGE 385 9 PERIOD 1, PREDOSE AVG 7.3.4 No Baseline; Single-Delta Treatment Difference ECGs are collected or extracted from continuous recordings in replicate sets (usually 3 replicates about a minute or so apart) at each Xij. The average of the replicates is presented in the table below. This definition of a treatment difference is rare and is not recommended, particularly not recommended for parallel studies. For crossover design, ΔQTcij is computed for each subject: ∆QTcij = Xij(Drug A) − Xij(Placebo) where i=1, 2, … d, j=1, 2, … n; d=days postdose and n=time point. USUBJID TRTA PARAMCD APERIOD 2324-P0001 Drug A 1 mg QTCFAG 1 2324-P0001 Drug A 1 mg QTCFAG 1 2324-P0001 Drug A 1 mg QTCFAG 1 2324-P0001 Drug A 1 mg QTCFAG 1 2324-P0001 Drug A 1 mg QTCFAG 1 2324-P0001 Drug A 1 mg QTCFAG 1 2324-P0001 Placebo QTCFAG 2 2324-P0001 Placebo QTCFAG 2 2324-P0001 Placebo QTCFAG 2 2324-P0001 Placebo QTCFAG 2 2324-P0001 Placebo QTCFAG 2 2324-P0001 Placebo QTCFAG 2 *Values in this column are mathematical expressions. AVISIT DAY 1 DAY 1 DAY 1 DAY 2 DAY 2 DAY 2 DAY 1 DAY 1 DAY 1 DAY 2 DAY 2 DAY 2 ATPT 1 HR 2 HR 3 HR 1 HR 2 HR 3 HR 1 HR 2 HR 3 HR 1 HR 2 HR 3 HR AVAL* X11(Drug A) X12(Drug A) X13(Drug A) X21(Drug A) X22(Drug A) X23(Drug A) X11(Placebo) X12(Placebo) X13(Placebo) X21(Placebo) X22(Placebo) X23(Placebo) X ij(Drug A) − � X ij(Placebo) For a parallel design, the Xijs would be averaged across subjects: ∆QTcij = � © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 74 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1247 Appendices 1248 Appendix A: Project Proposal 1249 1250 1251 1252 1253 1254 1255 1256 1257 1258 The project began in Q2 2013, with a target completion of the TAUG-QT in Q3 2014. 1259 Appendix B: CFAST Organizations 1260 1261 1262 1263 1264 1265 1266 1267 1268 1269 1270 1271 1272 1273 1274 1275 1276 1277 1278 1279 1280 1281 1282 1283 1284 1285 1286 1287 1288 1289 1290 1291 1292 1293 1294 1295 CFAST is proposing development of v1.0 of the CDISC QT therapeutic area data standard. This standard will build on the existing SDTM to facilitate the collection and use of data relevant to QT clinical trials. The workgroup proposes developing a CDISC therapeutic area user guide, including concept maps, examples and controlled terminology. It is clear that the existing CDISC standards are sufficient to cover the data standard requirements. The user guide is expected to focus on how these existing standards are to be used in the QT setting via examples and guidelines. Some investigation into the current ECG terminology and FDA subject level analysis dataset requirements related to QT may be required. CDISC CDISC is a global, open, multidisciplinary, non-profit organization that has established standards to support the acquisition, exchange, submission and archive of clinical research data and metadata. The CDISC mission is to develop and support global, platform-independent data standards that enable information system interoperability to improve medical research and related areas of healthcare. CDISC standards are vendor-neutral, platformindependent and freely available via the CDISC website. Critical Path Institute An independent, non-profit organization established in 2005 with public and private philanthropic support from the Arizona community, Science Foundation Arizona, and the U.S. Food and Drug Administration, C-Path’s mission is to improve human health and well-being by developing new technologies and methods to accelerate the development and review of medical products. An international leader in forming collaborations, C-Path has established global, public-private partnerships that currently include 1,000+ scientists from government regulatory agencies, academia, patient advocacy organizations, and dozens of major pharmaceutical companies. Association of Clinical Research Organizations The Association of Clinical Research Organizations (ACRO) represents the world's leading clinical research organizations. ACRO members provide specialized services that are integral to the development of drugs, biologics and medical devices. ACRO advances clinical outsourcing to improve the quality, efficiency and safety of biomedical research. Each year, ACRO’s members conduct thousands of clinical trials and provide related drug development services in more than 115 countries while ensuring the safety of nearly 2 million research participants. Innovative Medicines Initiative The Innovative Medicines Initiative (IMI) is Europe's largest public-private partnership aiming to improve the drug development process by supporting a more efficient discovery and development of better and safer medicines for patients. IMI supports collaborative research projects and builds networks of industrial and academic experts in Europe that will boost innovation in healthcare. Acting as a neutral third party in creating innovative partnerships, IMI aims to build a more collaborative ecosystem for pharmaceutical research and development. IMI supports research projects in the areas of safety and efficacy, knowledge management and education and training. National Cancer Institute Enterprise Vocabulary Services Since 1997, NCI Enterprise Vocabulary Services (EVS) has provided terminology content, tools, and services to accurately code, analyze and share cancer and biomedical research, clinical and public health information. EVS works with many partners to develop, license and publish terminology, jointly develop software tools, and support harmonization and shared standards. EVS provides the foundational layer for NCI's informatics infrastructure, and plays an important role in federal and international standards efforts. Page 75 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1296 1297 1298 1299 1300 1301 1302 1303 1304 1305 1306 1307 1308 1309 U.S. Food and Drug Administration The FDA is an agency of the United States Department of Health and Human Services. FDA is responsible for protecting the public health by assuring the safety, efficacy and security of human and veterinary drugs, biological products, medical devices, our nation’s food supply, cosmetics, and products that emit radiation. FDA is also responsible for advancing the public health by helping to speed innovations that make medicines more effective, safer, and more affordable and by helping the public get the accurate, science-based information they need to use medicines and foods to maintain and improve their health. 1310 Appendix C: Workgroup 1311 TransCelerate BioPharma, Inc. Launched in September 2012, TransCelerate BioPharma, Inc. aims to simplify and accelerate the delivery of innovative medicines to patients. The TCB mission is to collaborate across the global biopharmaceutical R&D community in order to identify, prioritize, design and facilitate implementation of solutions designed to drive the efficient, effective and high quality delivery of new medicines. Name Rhonda Facile, Team Leader Charles Beasley Cathy Bezek Natalie Boone Katie Carothers Marty Cisneroz Julie Evans Bala Hosmane Donna Kowalski Jordan Li Erin Muhlbradt Jaya Natarajan John Owen Dianne Reeves Pamela Rinaldi Klaus Romero Darcy Wold Diane Wold Fred Wood Sherry Wunder Institution/Organization CDISC Eli Lilly and Company Astellas Pharma Global Development, Inc. Astellas Pharma Global Development, Inc. CDISC C-Path CDISC AbbVie Astellas Pharma Global Development, Inc. NIH NCI EVS Janssen Research & Development Janssen Research & Development NCI EVS Boehringer Ingelheim C-Path CDISC GlaxoSmithKline Accenture Janssen Research & Development Ron Fitzmartin FDA © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 76 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1312 Appendix D: Glossary and Abbreviations Term/Abbreviation ADaM BRIDG CDASH CDISC CDSA CFAST C-Path Collected Controlled Terminology CRF CRO CV CV SAC Domain ECG/EKG eCRF FDA Foundational Standards GLS HR MedDRA mm msec mV Patient NCI EVS NIH QTc QTcB QTcF SDS SDTM SDTMIG SHARE Subject TCB UG UML 1313 1314 Definition Analysis Data Model Biomedical Research Integrated Domain Group Clinical Data Acquisition Standards Harmonization Project Clinical Data Interchange Standards Consortium Clinical Diagnostic Services Associate Coalition for Accelerating Standards and Therapies Critical Path Institute “Collected” refers to information that is recorded and/or transmitted to the sponsor. This includes data entered by the site on CRFs/eCRFs as well as vendor data such as core lab data. This term is a synonym for “captured”. A finite set of values that represent the only allowed values for a data item. These values may be codes, text, or numeric. A codelist is one type of controlled terminology. Case Report Form (sometimes called a Case Record Form). A printed, optical, or electronic document designed to record all required information to be reported to the sponsor for each trial subject. Clinical research organization Cardiovascular Cardiovascular Safety Advisory Committee A collection of observations with a topic-specific commonality about a subject. Electrocardiogram Electronic Case Report Form U.S. Food and Drug Administration “Foundational standards” is the term used to refer to the suite of CDISC standards that describe the clinical study protocol (Protocol), design (Study Design), data collection (CDASH), laboratory work (Lab), analysis (ADaM), and data tabulation (SDTM and SEND). See http://www.cdisc.org/ for more information on each of these clinical data standards. Generic laboratory system Heart rate Medical Dictionary for Regulatory Activities. Global standard medical terminology designed to supersede other terminologies (such as COSTART and ICD9) used in the medical product development process. millimeter millisecond microvolt A recipient of medical attention. National Cancer Institute (NCI) Enterprise Vocabulary Services National Institutes of Health QT interval corrected for heart rate QT interval corrected for heart rate using Bazett’s formula QT interval corrected for heart rate using Fridericia’s formula Submission Data Standards. Also the name of the team that created the SDTM and SDTMIG Study Data Tabulation Model SDTM Implementation Guide (for Human Clinical Trials) CDISC’s metadata repository that is currently under development. A participant in a study. TransCelerate BioPharma Inc. User Guide Unified Modeling Language See also Section 2.2, Explanation of Common Terms. Page 77 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) 1315 Appendix E: References 1. International Conference on Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use. E14: The Clinical Evaluation of QT/QTc Interval Prolongation and Proarrhythmic Potential for Non-antiarrhythmic Drugs. ICH. May 12, 2005. Available at: http://www.ich.org/fileadmin/Public_Web_Site/ICH_Products/Guidelines/Efficacy/E14/E14_Guideline.pdf. Accessed April 11, 2014. 2. Committee for Medicinal Products for Human Use. ICH topic E14: the clinical evaluation of QT/QTc interval prolongation and proarrhythmic potential for non-antiarrhythmic drugs questions and answers (R1)(EMA/CHMP/ICH/310133/2008). EMA. May 2012. Available at: http://www.ema.europa.eu/docs/en_GB/document_library/Scientific_guideline/2009/09/WC500002878.pdf. Accessed March 18, 2014. 3. Committee for Medicinal Products for Human Use. ICH topic E14: the clinical evaluation of QT/QTc interval prolongation and proarrhythmic potential for non-antiarrhythmic drugs questions and answers (R2) (EMA/CHMP/ICH/310111/2008). ICH. March 21, 2014. Available at: http://www.ich.org/fileadmin/Public_Web_Site/ICH_Products/Guidelines/Efficacy/E14/E14_QAs_R2_Step4.pdf. Accessed July 24, 2014. 4. Garnett CE, Zhu H, Malik M, et al. Methodologies to characterize the QT/corrected QT interval in the presence of drug-induced heart rate changes or other autonomic effects. Am Heart J. 2012;163(6):912-930. doi: 10.1016/j.ahj.2012.02.023. 5. Darpo B. The thorough QT/QTc study 4 years after the implementation of the ICH E14 guidance. Br J Pharmacol. 2010;159(1):49-57. doi: 10.1111/j.1476-5381.2009.00487.x. 6. Couderc JP, McNitt S, Hyrien O, et al. Improving the detection of subtle I(Kr)-inhibition: assessing electrocardiographic abnormalities of repolarization induced by moxifloxacin. Drug Saf. 2008;31(3):249-260. 7. Shah RR, Hondeghem LM. Refining detection of drug-induced proarrhythmia: QT interval and TRIaD. Heart Rhythm. 2005;2(7):758-772. doi: 10.1016/j.hrthm.2005.03.023. 8. United States Food and Drug Administration. Guidance for Industry: E14 Clinical Evaluation of QT/QTc Interval Prolongation and Proarrhythmic Potential for Non-Antiarrhythmic Drugs. Questions and Answers (R1). U.S. Food and Drug Administration. October 2012. Available at: http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/ucm073161.pdf. Accessed March 15, 2014. 9. Morganroth J. Design and conduct of the thorough phase I ECG trial for new bioactive drugs. In: Morganroth J, Gussak I, eds. Cardiac Safety of Noncardiac Drugs. Totowa, NJ: Humana Press; 2005. 10. Couderc JP, Xiaojuan X, Zareba W, Moss AJ. Assessment of the stability of the individual-based correction of QT interval for heart rate. Ann Noninvasive Electrocardiol. 2005;10(1):25-34. doi: 10.1111/j.1542474X.2005.00593.x. 11. Dmitrienko A, Beasely C, Mitchell M. Design and Analysis of Thorough QT Studies. BioPharmaceutical Network. April 29, 2008. Available at: http://www.biopharmnet.com/doc/2008_04_29_report.pdf. Accessed July 6, 2014. 12. Tsong Y. On the designs of thorough QT/QTc clinical trials. J Biopharm Stat. 2013;23(1):43-56. doi: 10.1080/10543406.2013.735762. 13. Salvi V, Karnad DR, Panicker GK, Kothari S. Update on the evaluation of a new drug for effects on cardiac repolarization in humans: issues in early drug development. Br J Pharmacol. 2010;159(1):34-48. doi: 10.1111/j.1476-5381.2009.00427.x. 14. Judson RS, Salisbury BA, Reed CR, Ackerman MJ. Pharmacogenetic issues in thorough QT trials. Mol Diagn Ther. 2006;10(3):153-162. 15. van Noord C, Eijgelsheim M, Stricker BH. Drug- and non-drug-associated QT interval prolongation. Br J Clin Pharmacol. 2010;70(1):16-23. doi: 10.1111/j.1365-2125.2010.03660.x. 16. Kaufmann H. Consensus statement on the definition of orthostatic hypotension, pure autonomic failure and multiple system atrophy. Clin Auton Res. 1996;6(2):125-126. 17. Low PA, Opfer-Gehrking TL, Textor SC, et al. Postural tachycardia syndrome (POTS). Neurology. 1995;45(4 © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 78 Draft CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Suppl 5):S19-S25. 18. Garland EM, Raj SR, Black BK, Harris PA, Robertson D. The hemodynamic and neurohumoral phenotype of postural tachycardia syndrome. Neurology. 2007;69(8):790-798. doi: 10.1212/01.wnl.0000267663.05398.40. 19. Malik M. Problems of heart rate correction in assessment of drug-induced QT interval prolongation. J Cardiovasc Electrophysiol. 2001;12(4):411-20. 1317 1318 1319 Appendix E1: Figures The following figures have been reproduced, with permission: Figure 3 Creative Commons Attribution 2.5 (CC-BY 2.5) license, 2007 Diagram of the Heart’s Heuser J. Electrical conduction system of the heart. Based on: Lynch PJ, Jaffe CC. Conduction System Heart anterior view coronal section. http://commons.wikimedia.org/wiki/File:RLS_12blauLeg.png Figure 4 Public domain. Sequence of Heart http://en.ecgpedia.org/wiki/File:Conduction_ap.svg. Excitation and the Associated ECG Waveforms Figure 6 (Left) Public domain. Standard Electrode/Lead http://en.wikipedia.org/wiki/File:Limb_leads.svg Configuration for a 12(Right) Creative Commons CC0 1.0 Universal (CC0 1.0) Public Domain Dedication Lead/Waveform ECG http://en.wikipedia.org/wiki/File:Precordial_leads_in_ECG.png 1320 1321 All other figures in this document were created or donated by the team, or originated within CDISC. 1322 Appendix E2: Further Reading 1323 1324 1325 1326 1327 1328 1329 1330 1331 1332 1333 1334 1335 1336 1337 1338 1339 1340 The following works are of interest to this document, but not actively referenced within it. • • • • • Florian JA, Tornoe CW, Brundage R, Parekh A, Garnett CE. Population pharmacokinetic and concentration-QTc models for moxifloxacin: pooled analysis of 20 thorough QT studies. J Clin Pharmacol. 2011;51(8):11521162. doi: 10.1177/0091270010381498. Garnett CE, Beasley N, Bhattaram VA, et al. Concentration-QT relationships play a key role in the evaluation of proarrhythmic risk during regulatory review. J Clin Pharmacol. 2008;48(1):13-18. doi: 10.1177/0091270007307881. ICH Topic S 7 B The nonclinical Evaluation of the Potential for delayed Ventricular Repolarization (QT Interval Prolongation) by Human Pharmaceuticals. (2005, November). European Medicines Agency. Available at: http://www.ema.europa.eu/docs/en_GB/document_library/Scientific_guideline/2009/09/WC500002841.pdf. Accessed 28 March 2014. Low PA, Sandroni P, Joyner M, et al. Postural tachycardia syndrome (POTS). J Cardiovasc Electrophysiol. 2009;20(3):352-358. Tornoe CW, Garnett CE, Wang Y, Florian J, Li M, Gobburu JV. Creation of a knowledge management system for QT analyses. J Clin Pharmacol. 2011;51(7):1035-1042. doi: 10.1177/0091270010378408. Page 79 Draft © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 CDISC Therapeutic Area Data Standards: User Guide for QT (Version 1.0 Draft) Appendix F: Representations and Warranties, Limitations of Liability, and Disclaimers CDISC Patent Disclaimers It is possible that implementation of and compliance with this standard may require use of subject matter covered by patent rights. By publication of this standard, no position is taken with respect to the existence or validity of any claim or of any patent rights in connection therewith. CDISC, including the CDISC Board of Directors, shall not be responsible for identifying patent claims for which a license may be required in order to implement this standard or for conducting inquiries into the legal validity or scope of those patents or patent claims that are brought to its attention. Representations and Warranties “CDISC grants open public use of this User Guide (or Final Standards) under CDISC’s copyright.” Each Participant in the development of this standard shall be deemed to represent, warrant, and covenant, at the time of a Contribution by such Participant (or by its Representative), that to the best of its knowledge and ability: (a) it holds or has the right to grant all relevant licenses to any of its Contributions in all jurisdictions or territories in which it holds relevant intellectual property rights; (b) there are no limits to the Participant’s ability to make the grants, acknowledgments, and agreements herein; and (c) the Contribution does not subject any Contribution, Draft Standard, Final Standard, or implementations thereof, in whole or in part, to licensing obligations with additional restrictions or requirements inconsistent with those set forth in this Policy, or that would require any such Contribution, Final Standard, or implementation, in whole or in part, to be either: (i) disclosed or distributed in source code form; (ii) licensed for the purpose of making derivative works (other than as set forth in Section 4.2 of the CDISC Intellectual Property Policy (“the Policy”)); or (iii) distributed at no charge, except as set forth in Sections 3, 5.1, and 4.2 of the Policy. If a Participant has knowledge that a Contribution made by any Participant or any other party may subject any Contribution, Draft Standard, Final Standard, or implementation, in whole or in part, to one or more of the licensing obligations listed in Section 9.3, such Participant shall give prompt notice of the same to the CDISC President who shall promptly notify all Participants. No Other Warranties/Disclaimers. ALL PARTICIPANTS ACKNOWLEDGE THAT, EXCEPT AS PROVIDED UNDER SECTION 9.3 OF THE CDISC INTELLECTUAL PROPERTY POLICY, ALL DRAFT STANDARDS AND FINAL STANDARDS, AND ALL CONTRIBUTIONS TO FINAL STANDARDS AND DRAFT STANDARDS, ARE PROVIDED “AS IS” WITH NO WARRANTIES WHATSOEVER, WHETHER EXPRESS, IMPLIED, STATUTORY, OR OTHERWISE, AND THE PARTICIPANTS, REPRESENTATIVES, THE CDISC PRESIDENT, THE CDISC BOARD OF DIRECTORS, AND CDISC EXPRESSLY DISCLAIM ANY WARRANTY OF MERCHANTABILITY, NONINFRINGEMENT, FITNESS FOR ANY PARTICULAR OR INTENDED PURPOSE, OR ANY OTHER WARRANTY OTHERWISE ARISING OUT OF ANY PROPOSAL, FINAL STANDARDS OR DRAFT STANDARDS, OR CONTRIBUTION. Limitation of Liability IN NO EVENT WILL CDISC OR ANY OF ITS CONSTITUENT PARTS (INCLUDING, BUT NOT LIMITED TO, THE CDISC BOARD OF DIRECTORS, THE CDISC PRESIDENT, CDISC STAFF, AND CDISC MEMBERS) BE LIABLE TO ANY OTHER PERSON OR ENTITY FOR ANY LOSS OF PROFITS, LOSS OF USE, DIRECT, INDIRECT, INCIDENTAL, CONSEQUENTIAL, OR SPECIAL DAMAGES, WHETHER UNDER CONTRACT, TORT, WARRANTY, OR OTHERWISE, ARISING IN ANY WAY OUT OF THIS POLICY OR ANY RELATED AGREEMENT, WHETHER OR NOT SUCH PARTY HAD ADVANCE NOTICE OF THE POSSIBILITY OF SUCH DAMAGES. Note: The CDISC Intellectual Property Policy can be found at http://www.cdisc.org/about/bylaws_pdfs/CDISCIPPolicy-FINAL.pdf © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved July 31, 2014 Page 80 Draft Supplementary Material SDTMIG Draft Domain: ECG QT Correction Model Data (QT) 6 Domain Models Based on the General Observation Classes 6.3 Findings ECG QT Correction Model Data (QT) QT – Description/Overview for ECG QT Correction Model Data Domain Model Data describing the description, correction formula and the coefficients of the correction formula used in correction of QT values. CDISC controlled terminology handles standard correction factors such as Bazett’s and Fredericia’s; however, due to the large and growing number of correction methods used, will not develop controlled terminology for those alternative correction factors. Therefore, this new findings domain was proposed to store the correction formula information. The ECG QT Correction Model Data domain is a draft domain at the time of this publication, but it fully conforms to the SDTM findings structure for versions 1.0 through 1.4, and could be used ould be used by sponsors under this name as a custom domain. No CDISC controlled terminology definition exists for the domain yet. QT – Specification for ECG QT Correction Model Data Domain Model qt.xpt, ECG QT Correction Model Data — Findings. One record per QT correction observation per subject, Tabulation. Controlled Role CDISC Notes Variable Name Variable Label Type Terms, Codelist or Format STUDYID Study Identifier Char Identifier Unique identifier for a study. DOMAIN Domain Abbreviation Char QT Identifier Two-character abbreviation for the domain. USUBJID Unique Subject Identifier Char Identifier Identifier used to uniquely identify a subject across all studies for all applications or submissions involving the product. QTSEQ Sequence Number Num Identifier Sequence Number given to ensure uniqueness of subject records within a domain. May be any valid number. QTGRPID Group ID Char Identifier Used to tie together a block of related records in a single domain for a subject. Example: use EGTESTCD as value for QTGRPID for easier relating of QT dataset to the EG dataset. QTREFID QTc Reference ID Char Identifier Internal or external QT identifier.Generally not used for QT dataset. QTSPID Sponsor-Defined Identifier Char Identifier Sponsor-defined reference number. Perhaps pre-printed on the CRF as an explicit line identifier or defined in the sponsor's operational database. Generally not used for QT dataset. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft Core Req Req Req Req Perm Perm Perm QT - Page 1 July 7, 2014 SDTMIG Draft Domain: ECG QT Correction Model Data (QT) Variable Name QTTESTCD Variable Label QTc Test or Examination Short Name Type Controlled Terms, Codelist or Format Char Role Core Short name of the measurement, test, or examination described in QTTEST. It can be used as a column name when converting a dataset from a vertical to a horizontal format. The value in QTTESTCD cannot be longer than 8 characters, nor can it start with a number (e.g., “1TEST”). QTTESTCD cannot contain characters other than letters, numbers, or underscores. Examples: QTCDESC, QTCFORM. QTTEST QTc Test or Examination Char Synonym Verbatim name of the test or examination used to obtain the measurement Name Qualifier or finding. The value in QTTEST cannot be longer than 40 characters. Examples: QT Correction Method Description, QT Correction Formula QTCAT Category for QTc Char * Grouping Used to categorize QT correction model observations across subjects. Qualifier Generally not used in QT dataset. QTSCAT Subcategory for QTc Char * Grouping A further categorization of the QT correction model. Generally not used Qualifier in QT dataset. QTORRES Result or Finding in Char Result Result of the QT correction model measurement or finding as originally Original Units Qualifier received or collected. Examples of expected values are 0.132 or 0.432 when the result is a coefficient or measurement, or “LINEAR” or “PARABOLIC LOG/LOG” when the result is a formula. QTSTRESC Character Result/Finding Char Result Contains the result value for all findings, copied or derived from in Std Format Qualifier QTORRES in a standard format or standard units. QTSTRESC should store all results or findings in character format; if results are numeric, they should also be stored in numeric format in QTSTRESN, for example 0.132 or 0.432 when the result is a coefficient or measurement, or “LINEAR” or “PARABOLIC LOG/LOG” when the result is a formula. QTSTRESN Numeric Result/Finding in Num Result Used for continuous or numeric results or findings in standard format; Standard Units Qualifier copied in numeric format from QTSTRESC. QTSTRESN should store all numeric test results or findings. QTSTAT Completion Status Char (ND) Record Used to indicate a QT correction model was not done. Should be null if a Qualifier result exists in QTORRES. QTREASND Reason QTc Not Char Record Describes why a measurement or test was not performed. Used in Performed Qualifier conjunction with QTSTAT when correction is NOT DONE. QTNAM Vendor Name Char Record Name or identifier of the laboratory or vendor who provided the test Qualifier results. QTDRVFL Derived Flag Char (NY) Record Used to indicate a derived record. The value should be Y or null. Records Qualifier which represent the average of other records, or that do not come from the CRF, or are not as originally collected or received are examples of records that would be derived for the submission datasets. If QTDRVFL=Y, then QTORRES could be null, with QTSTRESC, and (if numeric) QTSTRESN having the derived value. * Indicates variable may be subject to controlled terminology (Parenthesis indicates CDISC/ NCI codelist code value) © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft Topic CDISC Notes Req Req Perm Perm Exp Exp Perm Perm Perm Perm Perm QT - Page 2 July 7, 2014 SDTMIG Draft Domain: ECG QT Correction Model Data (QT) QT – Assumptions for ECG QT Correction Model Data Domain Model 1. The possible kinds of QT corrections are: a. 2. 3. 4. 5. 6. 7. 8. Historical Population QTc: Derived by fitting a model to data from a historical population of subjects. The formula for correcting QT, including its coefficient(s), is based on a formula which is known before the study starts. For example Bazett’s and Fridericia’s corrections are historical correction methods. b. Population QTc: Derived by fitting a model to data from the population of subjects in this study. The formula for correcting QT is known before the study starts, but the coefficient(s) in the correction formula are derived from data collected in the study. c. Multi-model population QTc: Derived by fitting several models to data for the study populating, then identifying one or more “best” models and using a correction method based on the best model(s). The candidate models, the method for choosing the best model(s) and, for methods that use multiple best models, the way in which the best models are combined, are known before the study starts. The formula for correcting QT, as well as its coefficient(s), are derived from data collected in the study. d. Individual QTc: Derived by fitting a model to data for each study subject individually. The formula for correcting QT is known before the study starts, but the coefficient(s) in the formula are derived from data collected from the individual subject. e. Multi-model Individual QTc: Derived by fitting several models to data for each subject, then identifying one or more “best fit” models for the subject and using a correction method based on the best fit model(s). The candidate models, the method for choosing the best model(s) and, for methods that use multiple best models, the way in which the best models are combined, are known before the study starts. The formula for correcting QT, as well as its coefficient(s), are derived from data collected from the individual subject. For each QT correction method used in the study, values of EGTESTCD and EGTEST which are assigned at the study level. CDISC Controlled Terminology includes EGTESTCD and EGTEST values for QT corrected by Bazett’s method and QT corrected by Fredericia’s method. CDISC Controlled Terminology does not plan to include any other corrected QT tests in CDISC Controlled Terminology in the near future. This is because of the large and growing number of QT corrections in use, and the fact that there is not scientific consensus on a best fit QT correction method. A protocol may specify that several different QT correction models be assessed and that only the best fit model for the individual be used for calculating the individual’s corrected QT intervals. Another alternative is that the protocol specifies multiple models be computed for each individual irrespective of best fit. There are multiple scenarios of analysis, and the sponsor should assign values for EGTESTCD/EGTEST appropriately with clear documentation on what each test code represents. For example, if the protocol calls for computing the top two best fit models, the sponsor could choose to name the top best fit model QTCIAG1 and the second best fit model QTCIAG2, in rank order. This domain provides information about the QT corrections used in the study. In that sense it contains findings about tests. The name of the test about which information is being supplied is stored in QTGRPID This domain is provided when coefficients for QT corrections other than Bazett’s or Fredericia’s are used in a study, and those corrections are derived from data collected in the study (i.e., are not historical), and the derivation was performed by a vendor, rather than the sponsor. Information on historical correction methods or derived by the sponsor could be submitted in the QT data domain, if agreed by the submitter and recipient (e.g., study sponsor and regulatory agency). For both individual and population corrections the USUBJID will be populated. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft QT - Page 3 July 7, 2014 SDTMIG Draft Domain: ECG QT Correction Model Data (QT) QT – Examples for ECG QT Correction Model Data Domain Model Example 1 This example consists of two Findings datasets: one for the ECG measurements (eg.xpt) and one for the QTc modeling information (qt.xpt). To tie the EG and QT datasets together will also require use of RELREC (relrec.xpt) to represent a dataset-to-dataset relationship that holds true for all subjects for all values of EGSEQ and QTGRPID. Rows 1-9: Shows calculated QTc values for three subjects. Each subject has three calculated QTc values. For each subject the first two corrections are based on an individual correction and the third for a population based correction. eg.xpt Row 1 2 3 4 5 6 7 8 9 STUDYID DOMAIN USUBJID EGSEQ EGCAT EGTESTCD EGTEST EGORRES EGORRESU STUDY01 EG 1 1 INTERVAL QTCIAG1 QTCI Interval, Aggregate 1 345 msec STUDY01 EG 1 2 INTERVAL QTCIAG2 QTCI Interval, Aggregate 2 350 msec STUDY01 EG 1 3 INTERVAL QTCNAG QTCN Interval, Aggregate 353 msec STUDY01 EG 2 1 INTERVAL QTCIAG1 QTCI Interval, Aggregate 1 372 msec STUDY01 EG 2 2 INTERVAL QTCIAG2 QTCI Interval, Aggregate 2 374 msec STUDY01 EG 2 3 INTERVAL QTCNAG QTCN Interval, Aggregate 377 msec STUDY01 EG 3 1 INTERVAL QTCIAG1 QTCI Interval, Aggregate 1 401 msec STUDY01 EG 3 2 INTERVAL QTCIAG2 QTCI Interval, Aggregate 2 411 msec STUDY01 EG 3 3 INTERVAL QTCNAG QTCN Interval, Aggregate 414 msec The example below contains the QT correction model information used for the three subjects mentioned in the eg.xpt file above. Rows 1-3: Row 1: Row 2: Row 3: Shows the correction method information for QTCIAG1. Shows a description of the QT Correction method. Shows the formula that was used to calculate the QT correction for that subject. Shows the QT correction co-efficient that was used in the formula for row 2. Note that this is an example of an individual correction model therefore the co-efficient used differs between subjects. Also note that for this example there is only 1 co-efficient used in the formula. Rows 4-6: Shows the same information used to calculate the linear correction model QTc value. Note that this is an example of an individual correction model therefore the coefficient used differs between subjects. Rows 7-11: Shows the same information used to calculate the Rautaharju correction model information. Note that this is a population based correction model and therefore the three co-efficients that are used in this calculation are the same for each subject. Also note that this model shows an example of how to represent three co-efficients used in a formula. Rows 12-22: Shows the same correction information used for subject 2. Rows 23-33: Shows the same correction information used for subject 3. qt.xpt Row 1 2 3 4 5 6 7 8 STUDYID DOMAIN USUBJID QTSEQ QTGRPID STUDY01 QT 1 1 QTCIAG1 STUDY01 QT 1 2 QTCIAG1 STUDY01 QT 1 3 QTCIAG1 STUDY01 QT 1 4 QTCIAG2 STUDY01 QT 1 5 QTCIAG2 STUDY01 QT 1 6 QTCIAG2 STUDY01 QT 1 7 QTCNAG STUDY01 QT 1 8 QTCNAG QTTESTCD QTTEST QTORRES QTCDESC QT Correction Method Description PARABOLIC LOG/LOG QTCFORM QT Correction Formula QTC=QT/(RR^A) QTCCOEFn QT Correction Coefficient 1 0.432 QTCDESC QT Correction Method Description LINEAR QTCFORM QT Correction Formula QTC=QT+(A*(1-RR)) QTCCOEFn QT Correction Coefficient 1 0.154 QTCDESC QT Correction Method Description RAUTAHARJU COR QTCFORM QT Correction Formula QTC=QT+A-(B*(e^(C^HR)) © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft QT - Page 4 July 7, 2014 SDTMIG Draft Domain: ECG QT Correction Model Data (QT) Row 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 STUDYID DOMAIN USUBJID QTSEQ QTGRPID STUDY01 QT 1 9 QTCNAG STUDY01 QT 1 10 QTCNAG STUDY01 QT 1 11 QTCNAG STUDY01 QT 2 1 QTCIAG1 STUDY01 QT 2 2 QTCIAG1 STUDY01 QT 2 3 QTCIAG1 STUDY01 QT 2 4 QTCIAG2 STUDY01 QT 2 5 QTCIAG2 STUDY01 QT 2 6 QTCIAG2 STUDY01 QT 2 7 QTCNAG STUDY01 QT 2 8 QTCNAG STUDY01 QT 2 9 QTCNAG STUDY01 QT 2 10 QTCNAG STUDY01 QT 2 11 QTCNAG STUDY01 QT 3 1 QTCIAG1 STUDY01 QT 3 2 QTCIAG1 STUDY01 QT 3 3 QTCIAG1 STUDY01 QT 3 4 QTCIAG2 STUDY01 QT 3 5 QTCIAG2 STUDY01 QT 3 6 QTCIAG2 STUDY01 QT 3 7 QTCNAG STUDY01 QT 3 8 QTCNAG STUDY01 QT 3 9 QTCNAG STUDY01 QT 3 10 QTCNAG STUDY01 QT 3 11 QTCNAG Rows 1-2: QTTESTCD QTCCOEFn QTCCOEFn QTCCOEFn QTCDESC QTCFORM QTCCOEFn QTCDESC QTCFORM QTCCOEFn QTCDESC QTCFORM QTCCOEFn QTCCOEFn QTCCOEFn QTCDESC QTCFORM QTCCOEFn QTCDESC QTCFORM QTCCOEFn QTCDESC QTCFORM QTCCOEFn QTCCOEFn QTCCOEFn QTTEST QTORRES QT Correction Coefficient 1 0.2425 QT Correction Coefficient 2 0.434 Correction Coefficient 3 -0.0097 QT Correction Method Description PARABOLIC LOG/LOG QT Correction Formula QTC=QT/(RR^A) QT Correction Coefficient 1 0.374 QT Correction Method Description LINEAR QT Correction Formula QTC=QT+(A*(1-RR)) QT Correction Coefficient 1 0.132 QT Correction Method Description RAUTAHARJU COR QT Correction Formula QTC=QT+A-(B*(e^(C^HR)) QT Correction Coefficient 1 0.2425 QT Correction Coefficient 2 0.434 Coefficient 3 -0.0097 QT Correction Method Description PARABOLIC LOG/LOG QT Correction Formula QTC=QT/(RR^A) QT Correction Coefficient 1 0.412 QTc Model Type LINEAR QT Correction Formula QTC=QT+(A*(1-RR)) QT Correction Coefficient 1 0.158 QTc Model Type RAUTAHARJU COR QT Correction Formula QTC=QT+A-(B*(e^(C^HR)) QT Correction Coefficient 1 0.2425 QT Correction Coefficient 2 0.434 Coefficient 3 -0.0097 Shows the RELREC information used to join the data in the eg.xpt dataset and the qt.xpt dataset. relrec.xpt Row 1 2 3 4 5 6 STUDYID RDOMAIN USUBJID IDVAR IDVARVAL RELTYPE RELID STUDY01 EG 1 EGTESTCD MANY EGEQ1 STUDY01 QT 1 QTGRPID MANY EGEQ1 STUDY01 EG 2 EGTESTCD MANY EGEQ1 STUDY01 QT 2 QTGRPID MANY EGEQ1 STUDY01 EG 3 EGTESTCD MANY EGEQ1 STUDY01 QT 3 QTGRPID MANY EGEQ1 © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft QT - Page 5 July 7, 2014 SDTMIG-MD Draft Dataset Specification: Device-Device Relationships (RELDEV) 4 Device Domains 4.8 Device-Device Relationships Some devices have components and it is important to record those relationships. In some cases relationship between a device and a component device may be physical and relatively permanent. In other cases, the top-level device may be a logical device, consisting of a devices used together to perform a test or intervention. reldev.xpt, Device-Device Relationships — Version x.x. One record per relationship parameter Variable Name Variable Label Controlled Definition Type Terms, Codelist or Format Char Unique identifier for a study. Char Sponsor-defined identifier for a device. STUDYID SPDEVID Study Identifier Sponsor Device Identifier PARENT Parent Sponsor Device Identifier Char DEVLVL Device Level Num PARMCD Relationship Parameter Short Name Relationship Parameter Char Relationship Parameter Value Char PARM VAL Char Implementation Notes Core Unique identifier for a study. It must be unique for each tracked unit of the device under study, and can be at whatever level of granularity the device should be identified (e.g., model or serial number, or combination of identifiers) as defined in DI. Must be a value of SPDEVID as defined in DI. Req Req The sponsor-defined identifier for the Exp “parent” device, the device of which the device named in SPDEVID is a component. Represents the hierarchy level of the device Should be a positive integer. The device at the top level has a Req named in SPDEVID, where 1 is the highest level of 1. A direct component of a level 1 device has a level of level. 2, etc. Short name for the relationship parameter. Perm A parameter that describes properties of the Examples: Quantity, Connection Type, Connection Location relationship between the component device named in SPDEVID and the parent device named in PARENT. The value of the parameter described in PARM. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft Perm Perm Page 1 April 4, 2014 SDTMIG-MD Draft Dataset Specification: Device-Device Relationships (RELDEV) 4.8.1 Assumptions for Device-Device Relationships Dataset 1. 2. 3. 4. 5. 6. The Device-Device Relationships Dataset is a relationship dataset that links device components to “parent” devices. As a relationship dataset, this does not have a Topic variable. As a relationship dataset, this dataset does not include variables such as DOMAIN and SEQ which are present in all general observation class domains. RDOMAIN is not needed, since it is expected that all records in this domain will link back to records in DI. PARENT is not populated for a device at the Level 1. PARM, PARMCD, and VAL are populated only if there are properties of the relationship that need to be represented. If the relationship between a device and a component has multiple properties, then the dataset will contain a record for each of these as a parameter. 4.8.2 Examples for Device-Device Relationships Dataset Example 1: This example shows the types of devices used to obtain ECG data for a particular study. Although multiple ECG machines were used, they were all of the same type, so identifiers were assigned to the type of machine, rather than to individual devices. There were two separate devices used to obtain the ECG data: an ECG machine that recorded data from the subject, and a second device that performed further processing of the ECG data from the first machine, and also allowed manual over-reading of the ECG data. Because these two machines were used together to produce the results in the ECG domain, the combination of the two devices is treated as a composite device, and the combination is given its own unique sponsor ID. Row 1: Rows 2-5: Rows 6-10: Shows the sponsor device identifier for the composite ECG Device. This is the device whose identifier appears in the EG dataset where subject data are represented. Further information about what this device consisted of is provided in the Device-Device Relationships dataset, below. Show the characteristics of the ECG machine, which has been given an SPDEVID value of "2 Show the characteristics of the machine which performs further processing of the ECG data from the first machine. This device has been given an SPDEVID value of “3.” di.xpt Row STUDYID DOMAIN SPDEVID DISEQ DIPARMCD DIPARM DIVAL ABC-123 DI 1 1 TYPE Device Type Composite ECG Device 1 ABC-123 DI 2 1 TYPE Device Type ECG Machine 2 ABC-123 DI 2 2 MANUF Manufacturer Acme 3 ABC-123 DI 2 3 MODEL Model XYZ 2000 4 ABC-123 DI 3 1 TYPE Device Type ECG Analyzer 5 ABC-123 DI 3 2 MANUF Manufacturer Acme 6 ABC-123 DI 3 3 MODEL Model ECG Wizard 7 The relationships between the two ECG machines and their combination is a simple one, which requires no additional parameters to describe the relationship, so the permissible variables PARMCD, PARM, and VAL are not included in the Device-Device Relationship dataset. © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft Page 2 April 4, 2014 SDTMIG-MD Draft Dataset Specification: Device-Device Relationships (RELDEV) Row 1: Row 2: Row 3: Shows that the device with SPDEVID = 1 has no parent and thus is at Level 1. Shows that the device with SPDEVID = 2 is a part of the composite device with SPDEVID = 1. Since the device with SPDEVID = 1 is at Level 1, this device is at Level 2. Shows that the device with SPDEVID = 3 is also a part of the composite device with SPDEVID = 1 and thus is also at Level 2. reldev.xpt Row STUDYID SPDEVID PARENT LEVEL ABC 1 1 ABC 2 1 2 2 ABC 3 1 2 3 Example 2 This example shows the components of a cardiac pacing device. There is information about the nature of the relationship between a device and its components that is needed, so the Device-Device Relationship Dataset includes the parameters PARMCD, PARM, and VAL. Row 1: Row 2-3: Row 4: Shows that the device with SPDEVID = LSKKDH23 has no parent and thus is at Level 1. Show that the device with SPDEVID = 237YALU is a part of the composite device with SPDEVID = LSKKDH23. There are two records for this relationship because there are two properties of the relationship of this component to its parent that are considered to be important in describing the relationship, and have been recorded. Row 3 shows that ten of the type of device with SPDEVID = 237YALU are components of the parent device. Row 4 shows that the connections of these components to the parent have “Connection Type” = “Active.” Shows that the device with SPDEVID = 29384LHS is also a part of the composite device with SPDEVID = LSKKDH23 and thus is also at Level 2. This relationship has one property, “Connection Location,” which is “Slot 3.” reldev.xpt Row STUDYID SPDEVID PARENT LEVEL PARMCD PARM VAL ABC LSKKDH23 1 1 ABC 237YALU LSKKDH23 2 QTY Quantity 10 2 ABC 237YALU LSKKDH23 2 CONNTYPE Connection Type Active 3 ABC 29384LHS LSKKDH23 2 CONNLOC Connection Location Slot 3 4 © 2014 Clinical Data Interchange Standards Consortium, Inc. All rights reserved Draft Page 3 April 4, 2014