Ch. 17 PPT
... 1 A small ribosomal subunit binds to a molecule of mRNA. In a prokaryotic cell, the mRNA binding site on this subunit recognizes a specific nucleotide sequence on the mRNA just upstream of the start codon. An initiator tRNA, with the anticodon UAC, base-pairs with the start codon, AUG. This tRNA car ...
... 1 A small ribosomal subunit binds to a molecule of mRNA. In a prokaryotic cell, the mRNA binding site on this subunit recognizes a specific nucleotide sequence on the mRNA just upstream of the start codon. An initiator tRNA, with the anticodon UAC, base-pairs with the start codon, AUG. This tRNA car ...
On the Nucleotide Sequence of Yeast Tyrosine Transfer RNA
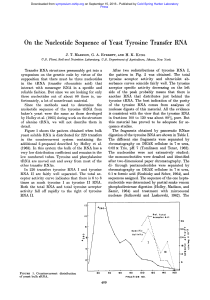
... Transfer RNA structures presumably get into a symposium on the genetic code by virtue of the supposition that there must be three nucleotides in the tRNA (transfer ribonucleic acid) that interact with messenger RNA in a specific and reliable fashion. But since we are looking for only three nucleotid ...
... Transfer RNA structures presumably get into a symposium on the genetic code by virtue of the supposition that there must be three nucleotides in the tRNA (transfer ribonucleic acid) that interact with messenger RNA in a specific and reliable fashion. But since we are looking for only three nucleotid ...
Chapter 3 Kinetic analysis of ribozyme cleavage
... ‘ribozyme’. Strictly speaking, a catalyst accelerates a multiple-turnover reaction without being changed itself. A few catalytic RNAs have this property, for example, RNase P and 23S rRNA; however, most ribozymes, in natural or evolved form, do not. For example, hairpin, HDV, VS1, and group I and gr ...
... ‘ribozyme’. Strictly speaking, a catalyst accelerates a multiple-turnover reaction without being changed itself. A few catalytic RNAs have this property, for example, RNase P and 23S rRNA; however, most ribozymes, in natural or evolved form, do not. For example, hairpin, HDV, VS1, and group I and gr ...
Foundations of Biology - Geoscience Research Institute
... The lac operon is made up of a control region and four genes: 1 LacZ - b-galactosidase - Enzyme that hydrolyzes the bond between galactose and glucose 2 LacY - Codes for a permease that lets lactose across the cell membrane 3 LacA - Transacetylase - An enzyme whose function in lactose metabolism is ...
... The lac operon is made up of a control region and four genes: 1 LacZ - b-galactosidase - Enzyme that hydrolyzes the bond between galactose and glucose 2 LacY - Codes for a permease that lets lactose across the cell membrane 3 LacA - Transacetylase - An enzyme whose function in lactose metabolism is ...
PROTEIN SYNTHESIS: TRANSLATION AND
... to the cytoplasmic sites of protein synthesis. In eukaryotes, the messengers, mRNAs, are usually synthesized as significantly larger precursor molecules that are processed prior to export from the nucleus. Eukaryotic mRNA in the cytosol has several identifying characteristics. It is almost always mo ...
... to the cytoplasmic sites of protein synthesis. In eukaryotes, the messengers, mRNAs, are usually synthesized as significantly larger precursor molecules that are processed prior to export from the nucleus. Eukaryotic mRNA in the cytosol has several identifying characteristics. It is almost always mo ...
Chapter Sixteen - Wright State University
... synthesis are. ■ Understand the general process by which proteins are made in a cell: where it happens and how it happens. ■ Understand the basic idea of the genetic code —that each amino acid is coded for by a sequence of three nucleotides (a codon). Appreciate that the human genome has about 3 bil ...
... synthesis are. ■ Understand the general process by which proteins are made in a cell: where it happens and how it happens. ■ Understand the basic idea of the genetic code —that each amino acid is coded for by a sequence of three nucleotides (a codon). Appreciate that the human genome has about 3 bil ...
lec-02-handout
... group. Prokaryotes have three types of DNA Polymerase while eukaryotes have five. Leading and lagging strands: DNA polymerase can synthesize DNA only in 5’to 3’ direction. Therefore, it synthesizes one strand (leading strand) continuously and the other strand(lagging strand) discontinuously. Each ne ...
... group. Prokaryotes have three types of DNA Polymerase while eukaryotes have five. Leading and lagging strands: DNA polymerase can synthesize DNA only in 5’to 3’ direction. Therefore, it synthesizes one strand (leading strand) continuously and the other strand(lagging strand) discontinuously. Each ne ...
Presentation 1 Guidelines
... together, a phosphate on one nucleotide forms a covalent bond with a hydroxyl group at the 3 position on another nucleotide. C7. The bases conform to the AT/GC rule of complementarity. There are two hydrogen bonds between A and T and three hydrogen bonds between G and C. The planar rings of the bas ...
... together, a phosphate on one nucleotide forms a covalent bond with a hydroxyl group at the 3 position on another nucleotide. C7. The bases conform to the AT/GC rule of complementarity. There are two hydrogen bonds between A and T and three hydrogen bonds between G and C. The planar rings of the bas ...
Lecture 1 - "Hudel" Luecke
... The universality of the Genetic Code is a result of strong evolutionary pressure: a change in a single codon would alter nearly every protein made by an organism. The universality is the basis for recombinant protein technology: mammalian mRNA sequences inserted into bacteria will be correctly expre ...
... The universality of the Genetic Code is a result of strong evolutionary pressure: a change in a single codon would alter nearly every protein made by an organism. The universality is the basis for recombinant protein technology: mammalian mRNA sequences inserted into bacteria will be correctly expre ...
Coupling transcription, splicing and mRNA export
... export machinery is co-transcriptionally loaded onto mRNAs in yeast, 30 -end formation may play a more critical role in this loading. For example, Lei and Silver [12] find that Sub2 is not required for the recruitment of Yra1 to genes that lack introns. However, proper 30 -end formation is required ...
... export machinery is co-transcriptionally loaded onto mRNAs in yeast, 30 -end formation may play a more critical role in this loading. For example, Lei and Silver [12] find that Sub2 is not required for the recruitment of Yra1 to genes that lack introns. However, proper 30 -end formation is required ...
View PDF - Molecular Systems Biology
... the biological insights provide by the analysis remain limited. We have also briefly consulted with a member of our advisory board who also acknowledged the value of the BATSeq and BATBayes methods but noted that orthogonal validation and follow up were lacking. As such, I am afraid we are not convi ...
... the biological insights provide by the analysis remain limited. We have also briefly consulted with a member of our advisory board who also acknowledged the value of the BATSeq and BATBayes methods but noted that orthogonal validation and follow up were lacking. As such, I am afraid we are not convi ...
U2Word
... 1. tRNAs are 54-100 nucleotide residues long, usually around 76. Much of the variation is in the 3 to 21 residue variable arm. Their secondary (20) structure is shaped like a “clover leaf” (by base pairing). Fig 32-9, 11 2. They contain many unusual bases - modified on primary transcript of tRNA gen ...
... 1. tRNAs are 54-100 nucleotide residues long, usually around 76. Much of the variation is in the 3 to 21 residue variable arm. Their secondary (20) structure is shaped like a “clover leaf” (by base pairing). Fig 32-9, 11 2. They contain many unusual bases - modified on primary transcript of tRNA gen ...
The structure of RNase E at the core of the RNA
... studies using X-ray spectroscopy13. In agreement with our mutagenic studies, the zinc is coordinated at the intra-domain linker through a pair of cysteine residues in the conserved CPxCxGxG motif that occurs throughout the RNase E/RNase G family13 (Fig. S1). A similar motif is found in the metal co ...
... studies using X-ray spectroscopy13. In agreement with our mutagenic studies, the zinc is coordinated at the intra-domain linker through a pair of cysteine residues in the conserved CPxCxGxG motif that occurs throughout the RNase E/RNase G family13 (Fig. S1). A similar motif is found in the metal co ...
Antisense Transcript and RNA Processing
... cells. In wild-type cells, atpB transcription reads through a downstream IR, followed by a two-step processing mechanism to yield the mature 39 end, which is coincident with the stem-loop (Stern and Kindle, 1993). In D26pAtE, the 39 IR, which is absent in D26, has been replaced by a sequence of 25 a ...
... cells. In wild-type cells, atpB transcription reads through a downstream IR, followed by a two-step processing mechanism to yield the mature 39 end, which is coincident with the stem-loop (Stern and Kindle, 1993). In D26pAtE, the 39 IR, which is absent in D26, has been replaced by a sequence of 25 a ...
tRNA
... The catalytic site forms a new peptide bond between valine and histidine. A three-aminoacid chain is now attached to the tRNA in the second binding site. The tRNA in the first site leaves, and the ribosome moves one codon over on the mRNA. Copyright © 2006 Pearson Prentice Hall, Inc. ...
... The catalytic site forms a new peptide bond between valine and histidine. A three-aminoacid chain is now attached to the tRNA in the second binding site. The tRNA in the first site leaves, and the ribosome moves one codon over on the mRNA. Copyright © 2006 Pearson Prentice Hall, Inc. ...
RNA Interference Regulates Gene Action
... dsRNA produced in the plants carrying transgenes oriented in both directions might silence RNAs more effectively than sense or antisense RNAs alone. A plasmid was constructed that contained an inverted repeat corresponding to a potato virus Y (PVY) protease gene essential for viral transmission (Fig ...
... dsRNA produced in the plants carrying transgenes oriented in both directions might silence RNAs more effectively than sense or antisense RNAs alone. A plasmid was constructed that contained an inverted repeat corresponding to a potato virus Y (PVY) protease gene essential for viral transmission (Fig ...
pdf file
... Initiation of transcription Transcription begins at the 3’ end of the gene in a region called the promoter. The promoter recruits TATA protein, a DNA binding protein, which in turn recruits other proteins. TATA binding protein Promoter DNA ...
... Initiation of transcription Transcription begins at the 3’ end of the gene in a region called the promoter. The promoter recruits TATA protein, a DNA binding protein, which in turn recruits other proteins. TATA binding protein Promoter DNA ...
Document
... Initiation of transcription Transcription begins at the 3’ end of the gene in a region called the promoter. ...
... Initiation of transcription Transcription begins at the 3’ end of the gene in a region called the promoter. ...
2.7 DNA replication, transcription and translation
... PCR is a way of producing large quantites of a specific target sequence of DNA. It is useful when only a small amount of DNA is avaliable for testing e.g. crime scene samples of blood, semen, tissue, hair, etc. ...
... PCR is a way of producing large quantites of a specific target sequence of DNA. It is useful when only a small amount of DNA is avaliable for testing e.g. crime scene samples of blood, semen, tissue, hair, etc. ...
Document
... PCR is a way of producing large quantites of a specific target sequence of DNA. It is useful when only a small amount of DNA is avaliable for testing e.g. crime scene samples of blood, semen, tissue, hair, etc. PCR occurs in a thermal cycler and involves a repeat procedure of 3 steps: 1. Denaturatio ...
... PCR is a way of producing large quantites of a specific target sequence of DNA. It is useful when only a small amount of DNA is avaliable for testing e.g. crime scene samples of blood, semen, tissue, hair, etc. PCR occurs in a thermal cycler and involves a repeat procedure of 3 steps: 1. Denaturatio ...
Exam #3 Review Exam #3 will cover from glycolysis to complex
... for phosphorylation of ADP to form ATP! ATP synthase - allows protons pumped out during production of the PMF to pass back into the cell ---> uses energy to fuel the phosphorylation of ADP to produce ATP. This is oxidative phosphorylation! • Practice: If 5 molecules of NADH are completely oxidized b ...
... for phosphorylation of ADP to form ATP! ATP synthase - allows protons pumped out during production of the PMF to pass back into the cell ---> uses energy to fuel the phosphorylation of ADP to produce ATP. This is oxidative phosphorylation! • Practice: If 5 molecules of NADH are completely oxidized b ...
Polyadenylation
Polyadenylation is the addition of a poly(A) tail to a messenger RNA The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature messenger RNA (mRNA) for translation. It, therefore, forms part of the larger process of gene expression.The process of polyadenylation begins as the transcription of a gene finishes, or terminates. The 3'-most segment of the newly made pre-mRNA is first cleaved off by a set of proteins; these proteins then synthesize the poly(A) tail at the RNA's 3' end. In some genes, these proteins may add a poly(A) tail at any one of several possible sites. Therefore, polyadenylation can produce more than one transcript from a single gene (alternative polyadenylation), similar to alternative splicing.The poly(A) tail is important for the nuclear export, translation, and stability of mRNA. The tail is shortened over time, and, when it is short enough, the mRNA is enzymatically degraded. However, in a few cell types, mRNAs with short poly(A) tails are stored for later activation by re-polyadenylation in the cytosol. In contrast, when polyadenylation occurs in bacteria, it promotes RNA degradation. This is also sometimes the case for eukaryotic non-coding RNAs.mRNA molecules in both prokaryotes and eukaryotes have polyadenylated 3'-ends, with the prokaryotic poly(A) tails generally shorter and less mRNA molecules polyadenylated.