Word file - UC Davis
... 4) Cytochrome P450 enzymes form a super-family of haem-containing oxygenases that are found in all kingdoms of life. These proteins have very similar structures but show extraordinary diversity in their reaction chemistry. Let us consider these three examples: (A) the human CYP46A1, an enzyme that c ...
... 4) Cytochrome P450 enzymes form a super-family of haem-containing oxygenases that are found in all kingdoms of life. These proteins have very similar structures but show extraordinary diversity in their reaction chemistry. Let us consider these three examples: (A) the human CYP46A1, an enzyme that c ...
Spectrophotometric methods for determination of proteins
... Qualitative test refers to descriptions or distinctions based on some quality or characteristic rather than on some quantity or measured value. It can be a form of analysis that yields the identity of a compound. ...
... Qualitative test refers to descriptions or distinctions based on some quality or characteristic rather than on some quantity or measured value. It can be a form of analysis that yields the identity of a compound. ...
General Reference - Methods Enzymol. 182 "Guide to Protein
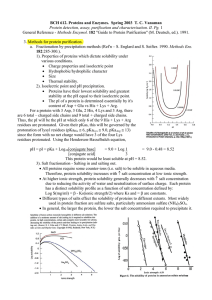
... x Proteins have their lowest solubility and greatest stability at the pH equal to their isoelectric point. x The pI of a protein is determined essentially by it's content of Asp + Glu vs His + Lys + Arg. For a protein with 3 Asp, 3 Glu, 2 His, 4 Lys and 3 Arg, there are 6 total – charged side chains ...
... x Proteins have their lowest solubility and greatest stability at the pH equal to their isoelectric point. x The pI of a protein is determined essentially by it's content of Asp + Glu vs His + Lys + Arg. For a protein with 3 Asp, 3 Glu, 2 His, 4 Lys and 3 Arg, there are 6 total – charged side chains ...
Steps in Protein Sequencing Separate Fragments and Sequence
... – A phylogenetic tree has been developed just from comparing sequences of cytochrome c from many organisms. (See Figure 5.29) ...
... – A phylogenetic tree has been developed just from comparing sequences of cytochrome c from many organisms. (See Figure 5.29) ...
Basic Local Alignment Search Tool (BLAST) IMBB 19, May 2015
... ü StaCsCcal measures like E-‐value. P-‐Value and bit score ü Percentage idenCty (% of idenCcal residues between sequences) ü The length of sequence stretch that is similar 2. Homology; homologs diverse f ...
... ü StaCsCcal measures like E-‐value. P-‐Value and bit score ü Percentage idenCty (% of idenCcal residues between sequences) ü The length of sequence stretch that is similar 2. Homology; homologs diverse f ...
Document
... Select known 3D structures of lipase (for example). Compare the target sequence with the known lipase structure by magic fit of Swiss-pdb Viewer. Use center the molecule on one atom tool bar to focus on the region of binding site. Compare the above two structures. ...
... Select known 3D structures of lipase (for example). Compare the target sequence with the known lipase structure by magic fit of Swiss-pdb Viewer. Use center the molecule on one atom tool bar to focus on the region of binding site. Compare the above two structures. ...
n - IBIVU
... A sheet consists of two or more hydrogen bonded strands. The two neighboring strands may be parallel if they are aligned in the same direction from one terminus (N or C) to the other, or anti-parallel if they are aligned in the opposite direction. ...
... A sheet consists of two or more hydrogen bonded strands. The two neighboring strands may be parallel if they are aligned in the same direction from one terminus (N or C) to the other, or anti-parallel if they are aligned in the opposite direction. ...
CSCE590/822 Data Mining Principles and Applications
... Given the amino acid sequence of a protein, what’s its shape in threedimensional space? ...
... Given the amino acid sequence of a protein, what’s its shape in threedimensional space? ...
- computer science publication server
... The rapid change in sensitivity and speci city at a threshold of 23 is due to lifting o the noise oor; a similar behavior is observed for pair-wise sequence comparisons. The evaluation on the SCOP1+SP data set allows us to investigate two di erent aspects of our method. First, we should be able to ...
... The rapid change in sensitivity and speci city at a threshold of 23 is due to lifting o the noise oor; a similar behavior is observed for pair-wise sequence comparisons. The evaluation on the SCOP1+SP data set allows us to investigate two di erent aspects of our method. First, we should be able to ...
Experimentally solving protein structures and protein
... There are through-bond interactions and through-space interactions. The latter usually being a consequence of the so-called nuclear Overhauser effect (NOE). Experiments of the nuclear-Overhauser variety may establish distances between atoms. These distances are subjected to a technique called Distan ...
... There are through-bond interactions and through-space interactions. The latter usually being a consequence of the so-called nuclear Overhauser effect (NOE). Experiments of the nuclear-Overhauser variety may establish distances between atoms. These distances are subjected to a technique called Distan ...
Chapter 4 - Evangel University
... Oxygen Binding of Hemoglobin (Hb) • A _________ of two -chains (141 amino acids each) and two -chains (153 amino acids each); 22 • Each chain has 1 heme group; hemoglobin can bind up to 4 molecules of O2 • Binding of O2 exhibited by _________ ___________; when one O2 is bound, it becomes easier ...
... Oxygen Binding of Hemoglobin (Hb) • A _________ of two -chains (141 amino acids each) and two -chains (153 amino acids each); 22 • Each chain has 1 heme group; hemoglobin can bind up to 4 molecules of O2 • Binding of O2 exhibited by _________ ___________; when one O2 is bound, it becomes easier ...
IOSR Journal of Computer Engineering (IOSR-JCE)
... Sequence alignment is a process of finding matching between DNA and Protein sequences in character-to-character level. The main role of sequences comparisons algorithms is to detect homologous regions between compared sequences. Comparing sequences may reveal or predict functional, structural, and e ...
... Sequence alignment is a process of finding matching between DNA and Protein sequences in character-to-character level. The main role of sequences comparisons algorithms is to detect homologous regions between compared sequences. Comparing sequences may reveal or predict functional, structural, and e ...
BIPASS: Bioinformatics pipelines alternative splicing services
... specific dimensions and information known about splicing events to (1) identify the signals controlling an encoded gene (here 'signal' refers to a transcription factor as illustrated in Figure 6) and (2) identify the proteins the mutated gene is likely to produce. Moreover, (3) we will infer the pro ...
... specific dimensions and information known about splicing events to (1) identify the signals controlling an encoded gene (here 'signal' refers to a transcription factor as illustrated in Figure 6) and (2) identify the proteins the mutated gene is likely to produce. Moreover, (3) we will infer the pro ...
Lecture 3 – Secondary Structure - LCQB
... • A secondary structure element is a contiguous segment of a protein sequence that presents a particular 3D geometry • Protein secondary structure prediction can be a first step toward tertiary structure prediction • PSSP algorithms historically rely on amino acid preferences for certain types of se ...
... • A secondary structure element is a contiguous segment of a protein sequence that presents a particular 3D geometry • Protein secondary structure prediction can be a first step toward tertiary structure prediction • PSSP algorithms historically rely on amino acid preferences for certain types of se ...
Secondary Structures and Properties of Fibrous Proteins
... 10). Proteins display motions (returning to the idea that life is dynamic!) 11). Quaternary structures describe the association between polypeptide chains. 12). Quaternary associations can be “open” or “closed” 13). Quaternary structures are stable (an interplay between entropy and chemical interact ...
... 10). Proteins display motions (returning to the idea that life is dynamic!) 11). Quaternary structures describe the association between polypeptide chains. 12). Quaternary associations can be “open” or “closed” 13). Quaternary structures are stable (an interplay between entropy and chemical interact ...
BIOGRAPHICAL SKETCH Abhijeet Kapoor Postdoctoral Research
... transitions in Ras proteins with the goal to identify specific structural features controlling the intrinsic conformational transitions, and complement the results using all-atom simulations. First, I developed the coarse-grained model that successfully folded nineteen proteins to their native state ...
... transitions in Ras proteins with the goal to identify specific structural features controlling the intrinsic conformational transitions, and complement the results using all-atom simulations. First, I developed the coarse-grained model that successfully folded nineteen proteins to their native state ...
Tasks Monday January 21st 2006
... click the 'Vista browser' option. Add all available organisms for comparison. What can you conclude? Which organisms could also be used to identify these individual elements and which are not very informative? What is the estimated size of each conserved region? Close the VISTA browser window and se ...
... click the 'Vista browser' option. Add all available organisms for comparison. What can you conclude? Which organisms could also be used to identify these individual elements and which are not very informative? What is the estimated size of each conserved region? Close the VISTA browser window and se ...
Protein Sequence - University of California, Davis
... 2. Will I be able to find structural or functional relatives? Is the protein similar to one that has been sequenced before? 1. How similar? 2. What does the similarity mean? Can I predict the function of the gene product, or is the predicted function consistent with what I know about the protein? Ca ...
... 2. Will I be able to find structural or functional relatives? Is the protein similar to one that has been sequenced before? 1. How similar? 2. What does the similarity mean? Can I predict the function of the gene product, or is the predicted function consistent with what I know about the protein? Ca ...
Introduction to 3D-Structure Visualization and Homology Modeling
... Template identification: Blast, PSI-Blast PDB: database of protein structures ...
... Template identification: Blast, PSI-Blast PDB: database of protein structures ...
The MOLECULES of LIFE
... can fold after translation by a different organism or even a cell-free (in vitro) translation system. ...
... can fold after translation by a different organism or even a cell-free (in vitro) translation system. ...
No Slide Title
... • Group of residues with high contact density, number of contacts within domains is higher than the number of contacts between domains. • A stable unit of protein structure that can fold autonomously • A rigid body linked to other domains by flexible linkers • A portion of the protein that can be ac ...
... • Group of residues with high contact density, number of contacts within domains is higher than the number of contacts between domains. • A stable unit of protein structure that can fold autonomously • A rigid body linked to other domains by flexible linkers • A portion of the protein that can be ac ...
Document
... There are through-bond interactions and through-space interactions. The latter usually being a consequence of the so-called nuclear Overhauser effect (NOE). Experiments of the nuclear-Overhauser variety may establish distances between atoms. • These distances are subjected to a technique called Dist ...
... There are through-bond interactions and through-space interactions. The latter usually being a consequence of the so-called nuclear Overhauser effect (NOE). Experiments of the nuclear-Overhauser variety may establish distances between atoms. • These distances are subjected to a technique called Dist ...
SQUADS #4
... denatured and then allowed to renature in a way that allows them to have their lowest-energy shapes. Which of the following statements about the proteins is most consistent with the information presented in the passage? A. If Scientist 1 is correct, all of the proteins will have their active shapes. ...
... denatured and then allowed to renature in a way that allows them to have their lowest-energy shapes. Which of the following statements about the proteins is most consistent with the information presented in the passage? A. If Scientist 1 is correct, all of the proteins will have their active shapes. ...
Test 2
... 2x2x2x2 16 different angles each of which can very over a range You can make same argument for a linear array of sugars; there will be 4 joins between sugars, and in each join you can vary the angle of one sugar or the other to get roughtly the same numebr of conformations The real difference comes ...
... 2x2x2x2 16 different angles each of which can very over a range You can make same argument for a linear array of sugars; there will be 4 joins between sugars, and in each join you can vary the angle of one sugar or the other to get roughtly the same numebr of conformations The real difference comes ...
Structural alignment

Structural alignment attempts to establish homology between two or more polymer structures based on their shape and three-dimensional conformation. This process is usually applied to protein tertiary structures but can also be used for large RNA molecules. In contrast to simple structural superposition, where at least some equivalent residues of the two structures are known, structural alignment requires no a priori knowledge of equivalent positions. Structural alignment is a valuable tool for the comparison of proteins with low sequence similarity, where evolutionary relationships between proteins cannot be easily detected by standard sequence alignment techniques. Structural alignment can therefore be used to imply evolutionary relationships between proteins that share very little common sequence. However, caution should be used in using the results as evidence for shared evolutionary ancestry because of the possible confounding effects of convergent evolution by which multiple unrelated amino acid sequences converge on a common tertiary structure.Structural alignments can compare two sequences or multiple sequences. Because these alignments rely on information about all the query sequences' three-dimensional conformations, the method can only be used on sequences where these structures are known. These are usually found by X-ray crystallography or NMR spectroscopy. It is possible to perform a structural alignment on structures produced by structure prediction methods. Indeed, evaluating such predictions often requires a structural alignment between the model and the true known structure to assess the model's quality. Structural alignments are especially useful in analyzing data from structural genomics and proteomics efforts, and they can be used as comparison points to evaluate alignments produced by purely sequence-based bioinformatics methods.The outputs of a structural alignment are a superposition of the atomic coordinate sets and a minimal root mean square deviation (RMSD) between the structures. The RMSD of two aligned structures indicates their divergence from one another. Structural alignment can be complicated by the existence of multiple protein domains within one or more of the input structures, because changes in relative orientation of the domains between two structures to be aligned can artificially inflate the RMSD.