Molecular Clocks
... The rate of this reaction is preset by the sequence and structure of the peptide or protein and associated peptides or proteins as genetically specified in the DNA. The rate may also be modified by changes in protein structure and solvent conditions in-vivo. This rate can be set to have a half-time ...
... The rate of this reaction is preset by the sequence and structure of the peptide or protein and associated peptides or proteins as genetically specified in the DNA. The rate may also be modified by changes in protein structure and solvent conditions in-vivo. This rate can be set to have a half-time ...
Promega Notes 39: Stabilized TMB Substrate for Horseradish
... Immunodetection on Western blots using TMB Stabilized Substrate and CN was performed to compare the detection sensitivities obtainable with these color development substrates. Samples of serially diluted beta-galactosidase (Worthington) were run on an 8% SDS-polyacrylamide minigel and electroblotted ...
... Immunodetection on Western blots using TMB Stabilized Substrate and CN was performed to compare the detection sensitivities obtainable with these color development substrates. Samples of serially diluted beta-galactosidase (Worthington) were run on an 8% SDS-polyacrylamide minigel and electroblotted ...
Endoplasmic Reticulum-Associated Degradation of Ricin A Chain
... Proteins that fail to fold in the endoplasmic reticulum (ER) or cannot find a pattern for assembly are often disposed of by a process named ER-associated degradation (ERAD), which involves transport of the substrate protein across the ER membrane (dislocation) followed by rapid proteasome-mediated p ...
... Proteins that fail to fold in the endoplasmic reticulum (ER) or cannot find a pattern for assembly are often disposed of by a process named ER-associated degradation (ERAD), which involves transport of the substrate protein across the ER membrane (dislocation) followed by rapid proteasome-mediated p ...
Proteasome inhibition induces reversible impairments in protein
... proteasome inhibitors to impair neuronal protein synthesis occurs prior to neuron death, at concentrations that also inhibit proteasome activity, and demonstrate a relationship between the concentrations of inhibitor necessary to impair protein synthesis and the LD50 for the different inhibitors. Si ...
... proteasome inhibitors to impair neuronal protein synthesis occurs prior to neuron death, at concentrations that also inhibit proteasome activity, and demonstrate a relationship between the concentrations of inhibitor necessary to impair protein synthesis and the LD50 for the different inhibitors. Si ...
Fibrous proteins and collagen
... types and organization are dictated by the structural role collagen plays in a particular organ. In some tissues, collagen may be dispersed as a gel that gives support to the structure, as in the extracellular matrix or the vitreous humor of the eye. In other tissues, collagen may be bundled in tigh ...
... types and organization are dictated by the structural role collagen plays in a particular organ. In some tissues, collagen may be dispersed as a gel that gives support to the structure, as in the extracellular matrix or the vitreous humor of the eye. In other tissues, collagen may be bundled in tigh ...
Ubiquitination and Auxin Signaling: A Degrading Story
... zyme, E1. Activated ubiquitin then is passed to one of a family of E2 ubiquitin-conjugating enzymes, from which it is conjugated to a Lys residue of the target protein, often with the assistance of an E3 enzyme, ubiquitin protein ligase. This process is reiterated such that one or two additional ubi ...
... zyme, E1. Activated ubiquitin then is passed to one of a family of E2 ubiquitin-conjugating enzymes, from which it is conjugated to a Lys residue of the target protein, often with the assistance of an E3 enzyme, ubiquitin protein ligase. This process is reiterated such that one or two additional ubi ...
REVIEWS - Haseloff Lab
... Box 1 | Ubiquitin and the ubiquitin–26S proteasome system In the initial ATP-consuming reaction, an E1 (or ubiquitin-activating enzyme) first activates the ubiquitin moiety by forming a high energy thioester bond between an E1 Cys residue and the carboxy-terminal Gly of ubiquitin (see the figure, pa ...
... Box 1 | Ubiquitin and the ubiquitin–26S proteasome system In the initial ATP-consuming reaction, an E1 (or ubiquitin-activating enzyme) first activates the ubiquitin moiety by forming a high energy thioester bond between an E1 Cys residue and the carboxy-terminal Gly of ubiquitin (see the figure, pa ...
Modification-specific proteomics: Strategies for characterization of
... for identifying candidate substrate proteins for certain types of PTM, such as tyrosine phosphorylation. However, the small size of the structural motifs of other common PTMs (for example, protein methylation and acetylation) makes it difficult to generate pan-specific antibodies, which recognize PT ...
... for identifying candidate substrate proteins for certain types of PTM, such as tyrosine phosphorylation. However, the small size of the structural motifs of other common PTMs (for example, protein methylation and acetylation) makes it difficult to generate pan-specific antibodies, which recognize PT ...
Full Text - Labs / Projects - Fred Hutchinson Cancer Research Center
... The isolation and characterization of the Hairy-binding proteins Groucho, dSir2, and dCtBP have suggested that Hairy represses transcription through a chromatin-remodeling mechanism. The conserved C-terminal WRPW residues of Hairy are required for the recruitment of the corepressor protein Groucho ( ...
... The isolation and characterization of the Hairy-binding proteins Groucho, dSir2, and dCtBP have suggested that Hairy represses transcription through a chromatin-remodeling mechanism. The conserved C-terminal WRPW residues of Hairy are required for the recruitment of the corepressor protein Groucho ( ...
The Sec61p Complex Mediates the Integration of a Membrane
... of 85 amino acids (85-mer), translation in the presence of membranes generated a band that migrated more slowly than that seen in the absence of membranes (Figure 1B, lane 19 versus 17; marked by an open star). This band corresponds to glycosylated chains because it was not seen when a competitor pe ...
... of 85 amino acids (85-mer), translation in the presence of membranes generated a band that migrated more slowly than that seen in the absence of membranes (Figure 1B, lane 19 versus 17; marked by an open star). This band corresponds to glycosylated chains because it was not seen when a competitor pe ...
Functional Regulation of MyD88-Activated Interferon Regulatory
... MyD88 pathway, where it interacts with both MyD88 and the E3 ubiquitin ligase, TRAF6. To understand the role of these interactions in the regulation of IRF-5, we examined the role of ubiquitination and showed that IRF-5 is subjected to TRAF6-mediated K63-linked ubiquitination, which is important for ...
... MyD88 pathway, where it interacts with both MyD88 and the E3 ubiquitin ligase, TRAF6. To understand the role of these interactions in the regulation of IRF-5, we examined the role of ubiquitination and showed that IRF-5 is subjected to TRAF6-mediated K63-linked ubiquitination, which is important for ...
Hitting the Target: Emerging Technologies in the Search for Kinase
... conductor. Aside from containing sequences necessary for three-dimensional folding into domains, many proteins have evolved motifs that allow posttranslational modifications that modulate function. By far the largest group of enzymes that catalyze regulatory posttranslational modifications is the fa ...
... conductor. Aside from containing sequences necessary for three-dimensional folding into domains, many proteins have evolved motifs that allow posttranslational modifications that modulate function. By far the largest group of enzymes that catalyze regulatory posttranslational modifications is the fa ...
Developmentally regulated, alternative splicing of the Rpn10 gene
... The 26S proteasome is a multisubunit proteindestroying machinery that degrades ubiquitin-tagged proteins. To date only a single species of Rpn10, which possibly functions as a multiubiquitin chain-binding subunit, has been identi®ed in various organisms. Here we report that mouse Rpn10 mRNAs occur i ...
... The 26S proteasome is a multisubunit proteindestroying machinery that degrades ubiquitin-tagged proteins. To date only a single species of Rpn10, which possibly functions as a multiubiquitin chain-binding subunit, has been identi®ed in various organisms. Here we report that mouse Rpn10 mRNAs occur i ...
Protein stabilization: a common consequence of mutations
... phenotype, and intriguingly, these are clustered in a hotspot region in the amino terminal transactivation domain (Henriksson and Luscher, 1996; Hoang et al., 1995; Smith-Sorensen et al., 1996). As a consequence of oncogenic activation, the normal signalling mechanisms which regulate cellular protoo ...
... phenotype, and intriguingly, these are clustered in a hotspot region in the amino terminal transactivation domain (Henriksson and Luscher, 1996; Hoang et al., 1995; Smith-Sorensen et al., 1996). As a consequence of oncogenic activation, the normal signalling mechanisms which regulate cellular protoo ...
Proteases and proteolytic cleavage of storage
... whereas the smaller more hydrophobic one (mol. wt. 20 kDa) has a strongly basic pi and is named /-chain. Both chains are linked by a disulphide bridge between cysteine residues at highly conserved positions in the a(amino acid residue 87) and /-chain (amino acid residue 7). The amino acid sequences ...
... whereas the smaller more hydrophobic one (mol. wt. 20 kDa) has a strongly basic pi and is named /-chain. Both chains are linked by a disulphide bridge between cysteine residues at highly conserved positions in the a(amino acid residue 87) and /-chain (amino acid residue 7). The amino acid sequences ...
instructions on the annotation of pdf files
... is not due to differences in domain lengths because it is observed also for proteins with domains of similar size (Table S1). For example, the values of the ratio nACO(Nt) < ACO(Ct)/ nACO(Ct) < ACO(Nt) for all the two-domain proteins in CATH and SCOP, when those with a difference of more than ten am ...
... is not due to differences in domain lengths because it is observed also for proteins with domains of similar size (Table S1). For example, the values of the ratio nACO(Nt) < ACO(Ct)/ nACO(Ct) < ACO(Nt) for all the two-domain proteins in CATH and SCOP, when those with a difference of more than ten am ...
COLLAGEN - Rihs.com.pk
... After hydroxylation and glycosylationPro α-chains are converted to Pro-collagen. Pro-collagen has a central region of triple helix and its ends have non-helical regions of amino and carboxyl terminal extensions . These extensions are called Propeptides. ...
... After hydroxylation and glycosylationPro α-chains are converted to Pro-collagen. Pro-collagen has a central region of triple helix and its ends have non-helical regions of amino and carboxyl terminal extensions . These extensions are called Propeptides. ...
collagen - MBBS Students Club
... After hydroxylation and glycosylationPro α-chains are converted to Pro-collagen. Pro-collagen has a central region of triple helix and its ends have non-helical regions of amino and carboxyl terminal extensions . These extensions are called Propeptides. ...
... After hydroxylation and glycosylationPro α-chains are converted to Pro-collagen. Pro-collagen has a central region of triple helix and its ends have non-helical regions of amino and carboxyl terminal extensions . These extensions are called Propeptides. ...
Digestive Enzymes Plus
... matter; they are essential to maintain all the body functions working properly. Each enzyme has a specific function in the body that no other enzyme can carry out. Digestive enzymes are specifically involved in the process of digestion. They are considered proteolytic enzymes since they only work on ...
... matter; they are essential to maintain all the body functions working properly. Each enzyme has a specific function in the body that no other enzyme can carry out. Digestive enzymes are specifically involved in the process of digestion. They are considered proteolytic enzymes since they only work on ...
Contribution of molecular chaperones to protein folding in the
... molecular chaperones are: Hsp110, Hsp100, Hsp90, Hsp70, Hsp60 (chaperonins) and sHsp (small Hsp, with molecular masses between 15 and 30 kDa) ...
... molecular chaperones are: Hsp110, Hsp100, Hsp90, Hsp70, Hsp60 (chaperonins) and sHsp (small Hsp, with molecular masses between 15 and 30 kDa) ...
Analysis of protein carbonylation
... interaction with pre-formed reactive carbonyl species). Data in Table 3 are sufficient to illustrate the variety ...
... interaction with pre-formed reactive carbonyl species). Data in Table 3 are sufficient to illustrate the variety ...
Biological significance of structural differences between two highly
... The structure of Uev1D30 was solved by molecular replacement using a poly-alanine model of Mms2 (PDB accession number: 1J74). The density for the core 140 residues of Uev1D30 was well ordered (Fig. 1B) and adopted a structure very similar to Mms2 [8] with an RMSD of 0.5 Å. This was expected given th ...
... The structure of Uev1D30 was solved by molecular replacement using a poly-alanine model of Mms2 (PDB accession number: 1J74). The density for the core 140 residues of Uev1D30 was well ordered (Fig. 1B) and adopted a structure very similar to Mms2 [8] with an RMSD of 0.5 Å. This was expected given th ...
1 Causality, Transfer Entropy and Allosteric
... Ubiquitin, (PDB code 1UBQ), is a 76 amino acid protein. It consists of 8 distinct secondary structures that actively take part in interactions with a large number of proteins. Ubiquitin, although not known as an allosteric protein itself, communicates with several other proteins to send information ...
... Ubiquitin, (PDB code 1UBQ), is a 76 amino acid protein. It consists of 8 distinct secondary structures that actively take part in interactions with a large number of proteins. Ubiquitin, although not known as an allosteric protein itself, communicates with several other proteins to send information ...
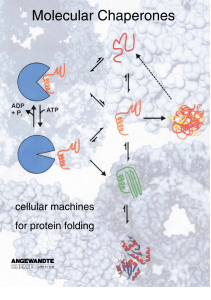
Molecular Chaperones - Cellular Machines for Protein Folding
... The GroE proteins of the bacterium E. coli are the most extensively studied molecular chaperones.[21±24] The groEL and groES genes encode proteins of 57 kDa and 10 kDa size, respectively, which are both required for the viability of E. coli.[25] Thus, at least one essential E. coli protein cannot fo ...
... The GroE proteins of the bacterium E. coli are the most extensively studied molecular chaperones.[21±24] The groEL and groES genes encode proteins of 57 kDa and 10 kDa size, respectively, which are both required for the viability of E. coli.[25] Thus, at least one essential E. coli protein cannot fo ...
Ubiquitin

Ubiquitin is a small (8.5 kDa) regulatory protein that has been found in almost all tissues (ubiquitously) of eukaryotic organisms. It was discovered in 1975 by Goldstein and further characterized throughout the 1970s and 1980s. There are four genes in the human genome that produce ubiquitin: UBB, UBC, UBA52 and RPS27A.The addition of ubiquitin to a substrate protein is called ubiquitination or ubiquitylation. Ubiquitination can affect proteins in many ways: It can signal for their degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitination is carried out in three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade binds ubiquitin to lysine residues on the protein substrate via an isopeptide bond or to the amino group of the protein's N-terminus via a peptide bond.The protein modifications can be either a single ubiquitin protein (monoubiquitination) or a chain of ubiquitin (polyubiquitination). The ubiquitination bonds are always formed with one of the seven lysine residues from the ubiquitin molecule. These 'linking' lysines are represented by a ""K"" (which is the one-letter amino acid notation of lysine) and a number, referring to its position in the ubiquitin molecule. First, a ubiquitin molecule is bonded by its C-terminus to a specific lysine residue (e.g. K48, K29, K63,...) on the target protein. Poly-ubiquitination occurs when the C-terminus of another ubiquitin, will be linked again to a lysine residue (for example again K48 or K29) on the previously added ubiquitin molecule, forming a chain. This process repeats several times, leading to the addition of several ubiquitins. Only poly-ubiquitination on defined lysines, mostly on K48 and K29, is related to degradation with the proteasome (referred to as the ""molecular kiss of death""), while other polyubiquitinations (e.g. on K63, K11, K6) and monoubiquitinations may regulate processes such as endocytic trafficking, inflammation, translation and DNA repair.Lysine 48-linked chains have been much-studied. They are the forms of chains that signal proteins to the proteasome, which destroys and recycles proteins. This discovery won the Nobel Prize for chemistry in 2004.