* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download - ChemAxon
Survey
Document related concepts
Entity–attribute–value model wikipedia , lookup
Microsoft SQL Server wikipedia , lookup
Extensible Storage Engine wikipedia , lookup
Open Database Connectivity wikipedia , lookup
Ingres (database) wikipedia , lookup
Oracle Database wikipedia , lookup
Microsoft Jet Database Engine wikipedia , lookup
Concurrency control wikipedia , lookup
Versant Object Database wikipedia , lookup
Relational model wikipedia , lookup
Clusterpoint wikipedia , lookup
Transcript
JChem Base chemical database
Szilárd Dóránt
1
May, 2005
Slide ‹#›
Contents
Introduction
Structural overview
Compatibility
Administration
JChem tables
Fingerprints
Structural search
Jchem Base chemical database — May 2005
Structure cache
Standardization
Search options
JSP example
API examples
Performance
Future plans
2
Slide ‹#›
Introduction
JChem Base provides high performance Java
based tools for the storage, search and
retrieval of chemical structures and associated
data.
These components can be integrated into webbased or standalone applications in association
with other ChemAxon tools.
Jchem Base chemical database — May 2005
3
Slide ‹#›
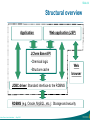
Structural overview
Application
Web application (JSP)
JChem Base API:
•Chemical logic
•Structure cache
Web
browser
JDBC driver: Standard interface to the RDBMS
RDBMS (e.g. Oracle, MySQL, etc.) : Storage and security
Jchem Base chemical database — May 2005
4
Slide ‹#›
Compatibility and integration
File formats:
• SMILES
• MDL molfile
(v2000 and v3000)
• MDL SDF
• RXN
• RDF
• MRV
Integration:
• 100% Java
• extensive API
• JChem Cartridge for
Oracle
Jchem Base chemical database — May 2005
Database engines:
• Oracle
• MySQL
• MS SQL Server
• PostgreSQL
• MS Access
• DB2
• etc.
Operating systems:
• Windows
• Linux
• Mac OS X
• Solaris
• etc.
5
Slide ‹#›
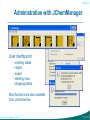
Administration with JChemManager
User interface for
• creating tables
• import
• export
• deleting rows
• dropping tables
Most functions are also available
from command-line.
Jchem Base chemical database — May 2005
6
Slide ‹#›
The property table
The property table stores information about JChem
structure tables, including:
• Fingerprint parameters
• Custom standardization rules
• Recent changes (to optimize cache updates)
• Other table options and information
• Database-related licence keys
More than one property table can be used, each
property table represents a particular JChem
environment.
Jchem Base chemical database — May 2005
7
Slide ‹#›
The structure of JChem tables
Column name
Explanation
cd_id
unique numeric identifier in the table
cd_structure
the imported structure in the original format, without
modifications (except for the removal of data fields)
cd_smiles
the standardized structure in ChemAxon Extended Smiles
(cxsmiles) format, used by the search process
cd_formula
the formula of the standardized structure
cd_molweight
the molecular weight of the standardized structure
cd_hash
hash code used for duplicate filtering (PERFECT search)
cd_flags
can store row specific option, e.g. overriding the chiral
flag
cd_timestamp
the date and time of the insertion of the row
cd_fp…
fingerprint columns
[user fields]
custom data fields can be added by the user
Jchem Base chemical database — May 2005
8
Slide ‹#›
Chemical Hashed Fingerprints
• Chemical Hashed Fingerprints encode structural
patterns in bit strings
• If structure A is a substructure of structure B, every
bit in B’s fingerprint will be set that is set in structure
A’s fingerprint:
A& B A
• Tanimoto similarity of hashed fingerprints can be
used for diversity analysis and similarity search:
Tsim X , Y
Jchem Base chemical database — May 2005
BitCount X & Y
BitCount X BitCount Y BitCount X & Y
9
Slide ‹#›
Structural search in database
Two stage method provides optimal performance:
1. Rapid pre-screening reduces the number of
possible hit candidates
-
Chemical Hashed Fingerprints are used for
substructure and superstructure searches
Hash code is used for duplicate filtering
(usually during compound registration)
2. Graph search algorithm is used to determine
the final hit list
Jchem Base chemical database — May 2005
10
Slide ‹#›
Structure Cache
• Contains Fingerprints for screening and ChemAxon Extended
SMILES for ABAS
• Instant access to the structures for the search process
• Reduced load on the database server
• Incremental update ensures minimum overhead after changes
in the table
• Small memory footprint due to
– SMILES compression
– Optimized storage technique
• Approximately 100MB memory needed for 1 million typical
drug-like structures (using 512 bit long fingerprints)
Jchem Base chemical database — May 2005
11
Slide ‹#›
Standardization
• Default standardization
includes:
– Hydrogen removal
– Aromatization
• Custom standardization
can be specified for
each table by specifying
an XML configuration file
at table creation or in the
“Regenerate” dialog of
JChem Manager (jcman)
Jchem Base chemical database — May 2005
12
Slide ‹#›
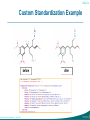
Custom Standardization Example
before
Jchem Base chemical database — May 2005
after
13
Slide ‹#›
Database search options
• Maximum search time / number of hits
• SQL SELECT statement for pre-filtering
• Ordering of results
• Result table
• Inverse hit list
• Chemical Terms filter constraint
Jchem Base chemical database — May 2005
14
Slide ‹#›
JSP example application
•
Open source, customizable
•
Features:
– Substructure, Superstructure,
Exact and Similarity search
– Molecular Descriptor similarity
search with descriptor coloring
– Substructure hit alignment and
coloring, inverse hit list
– Chemical Terms filter
– Import / Export
– Export of hits
– Insert / Modify / Delete
structures
Jchem Base chemical database — May 2005
15
Slide ‹#›
API example : connecting to a database
ConnectionHandler ch = new chemaxon.jchem.db.ConnectionHandler();
ch.setDriver(“oracle.jdbc.driver.OracleDriver”);
ch.setUrl(“jdbc:oracle:thin:@localhost:1521:mydb”);
ch.setPropertyTable(“JChemProperties”);
ch.setLoginName(“scott”);
ch.setPassword("tiger");
ch.connect();
// the java.sql.Connection object is available if needed:
Connection con=ch.getConnection();
…
// closing the connection:
ch.close();
Jchem Base chemical database — May 2005
16
Slide ‹#›
API example : database import
Importer importer = new chemaxon.jchem.db.Importer();
importer.setConnectionHandler(conh);
importer.setInput(“sample.sdf”);
// importer.setInput(is);
// alternatively a stream can also be specified
importer.setTableName(“SCOTT.STRUCTURES”);
importer.setHaltOnError(false);
importer.setDuplicateImportAllowed(false);
//can filter duplicates
// specifying SDFile field - table field pairs:
String fieldPairs = “DB_Field1=SDF_Field1; DB_Field2=SDF_Field2”;
importer.setFieldConnections(fieldPairs);
int importedCount = importer.importMols();
System.out.println( “Imported” + importedCount + “structures” );
Jchem Base chemical database — May 2005
17
Slide ‹#›
API example : database export
Exporter exporter = new chemaxon.jchem.db.Exporter();
exporter.setConnectionHandler(conh);
exporter.setTableName(“structures”);
//data fields to be exported with the structure:
exporter.setFieldList(“cd_id cd_formula name comments”);
String fileName=“output.sdf”;
OutputStream os=new FileOutputStream(fileName);
exporter.setOutputStream(os);
exporter.setFormat(“sdf”);
int exportedCount = exporter.writeAll();
System.out.println(“Exported ” + exportedCount + “structures”);
Jchem Base chemical database — May 2005
18
Slide ‹#›
API example : database search
JChemSearch searcher = new chemaxon.jchem.db.JChemSearch();
searcher.setConnectionHandler(ch);
searcher.setSearchType(JChemSearch.SUBSTRUCTURE)
searcher.setQueryStructure(“c1ccccc1”);
searcher.setStructureTable(“SCOTT.STRUCTURES”);
// a query that returns cd_id values can be used for prefiltering:
Searcher.setFilterQuery(
“SELECT cd_id FROM structures, biodata WHERE ”
+ “structures.cd_id = biodata.cd_id AND biodata.toxicity < 0.3” );
searcher.setWaitingForResult(true);
// otherwise runs in a separate thread
searcher.setStructureCaching(true);
// caching speeds up the search
searcher.run();
// getting the results as cd_id values:
int[] results=searcher.getResults();
Jchem Base chemical database — May 2005
19
Slide ‹#›
API example : inserting a structure
// ConnectionHandler, mode, table name and data field names:
UpdateHandler uh = new chemaxon.jchem.db.UpdateHandler(
ch, UpdateHandler.INSERT, “structures”, “comment, stock”);
uh.setValueForFixColumns(“c1ccccc1”); // the structure
// specifying data field values:
uh.setStructureValueForAdditionalColumn(1, “some text”);
uh.setStructureValueForAdditionalColumn(2, new Double(8.5));
uh.setDuplicateFiltering(true); // filtering duplicate structures
int id=uh.execute(true); // getting back the cd_id of the inserted structure
if ( id > 0 ) {
System.out.println(“Inserted, cd_id value : ” + id);
} else {
System.out.println(“Already exists with cd_id value : ” + (-id));
}
// storing update information, the database connection remains open :
uh.close();
Jchem Base chemical database — May 2005
20
Slide ‹#›
Performance (1)
Compound registration:
Substructure search
in a table of 3 million
compounds:
Number of
compounds
Elapsed time
Duplicates not checked
Duplicates checked
10,000
32s
45s
100,000
4min 11s
6min 20s
200,000
8min 17s
12min 26s
Query
Number of hits
Search time (s)
12
0.1
936
0.9
0
1.2
49740
10.7
Server parameters: Windows XP; 1 CPU: Intel P4 3.0GHz; 2GB RAM; Oracle 9i
Jchem Base chemical database — May 2005
21
Slide ‹#›
Performance (2)
Similarity search:
Tanimoto >0.8
Query
Number of hits
Search time (s)
24
1.5
156
1.3
336
1.3
Server parameters: Windows XP; 1 CPU: Intel P4 3.0GHz; 2GB RAM; Oracle 9i
Jchem Base chemical database — May 2005
22
Slide ‹#›
Future plans
• Additional layer: JChem Server (later also as grid)
• Structural keys as optional extension to current
fingerprints
• Tables for storing query structures
• Tables for storing general (Markush) structures
• Partial clean option for hit alignment
• Installer
• etc.
Jchem Base chemical database — May 2005
23
Slide ‹#›
Summary
ChemAxon’s JChem Base toolkit
provides sophisticated methods to deal
with chemical structures and associated
data.
The usage of fingerprints and structure
cache provide high search performance.
Jchem Base chemical database — May 2005
24
Slide ‹#›
Links
• JChem home page:
– www.jchem.com
• Live demos:
– www.jchem.com/examples
• API documentation:
– www.jchem.com/doc/api
• Brochure:
– www.chemaxon.com/brochures/JChemBase.pdf
Jchem Base chemical database — May 2005
25
Slide ‹#›
Thank you for your attention
Máramaros köz 3/a
Budapest, 1037
Hungary
[email protected]
www.chemaxon.com
Jchem Base chemical database — May 2005
26