* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Directional Selection on a discrete trait
Designer baby wikipedia , lookup
Adaptive evolution in the human genome wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Koinophilia wikipedia , lookup
History of genetic engineering wikipedia , lookup
Viral phylodynamics wikipedia , lookup
Dual inheritance theory wikipedia , lookup
Human genetic variation wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
The Selfish Gene wikipedia , lookup
Selective breeding wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Genetic drift wikipedia , lookup
Sexual selection wikipedia , lookup
Natural selection wikipedia , lookup
Population genetics wikipedia , lookup
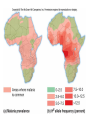
Natural Selection Natural Selection An Adaptive Mechanism Evolutionary Mechanisms Genetic Drift Migration Non-adaptive Mutations Natural Selection Adaptive Darwin Evolution acts through changes in allele frequency at each generation According to Darwin, this happens via Natural Selection (He considered Selection only, and not other evolutionary mechanisms, such as mutations, genetic drift, etc.) Sources of Genetic Variation Mutation generates genetic variation Natural Selection Natural Selection acts on genetic or epigenetic variation in a population Epigenetic modification changes expression of genes Genetic Drift can reduce genetic variation Without genetic or epigenetic variation, Natural Selection cannot OUTLINE (1) General Description of Natural Selection (2) Modes of Natural Selection (3) Selection Violating Hardy-Weinberg Equilibrium Darwin’s contribution: Population Speciation as a result of Natural Selection Too many offspring produced Limited resources and competition Variation in a population Better adapted individuals survive Survivors leave more offspring Thus, average character of population is altered Natural Selection The differential survival and/or reproduction of classes of entities that differ in one or more characteristics The only evolutionary force that is ADAPTIVE NOT random, but deterministic Darwin Population speciation Darwin Population speciation mutation Darwin Population speciation Selection favors Darwin Population speciation Greater Fitness Selection on pathogens imposed by Drugs (refer to Lecture 1) AZT AZT (Azidothymidine) is a thymidine mimic which stops reverse transcription Mutations in the reverse transcriptase gene of HIV arises such that the enzyme can recognize AZT The drugs impose Selection on the virus. Genetic variation (generated by mutations) allows the virus to respond to selection and evolve drug resistance Selection imposed by Transmission Rate on Virulence of HIV Need to keep host alive long enough to get passed on to the next host (Tradeoff between fast population growth and keeping the host alive) High Transmission rate : High Virulence (Can grow fast and jump to the next host; ok if host dies; genetic strain that grows faster will win) Low Transmission Rate : Low Virulence (More virulent strains would die with the host and get selected out; less virulent strain will win) High Transmission Rate If the virus is likely to move to a new host the faster growing (and more virulent) strain is likely to overtake the slower strains and “win” It’s ok to kill the host, since the chances of jumping to a new host is high Natural selection will favor the more virulent strain Low Transmission Rate If the virus is not likely to move to a new host the slower growing (and less virulent) strain is likely to “win” It’s not ok to kill the host, since the chances of jumping to a new host is low. If the virus kills the host, it will kill itself Natural selection will favor the less virulent strain Evolution of HIV resistance in Human Populations CCR5-Δ32 (or CCR5-D32 or CCR5 delta 32) is a mutant allele of the receptor CCR5, where the deletion of a 32 base pair segment makes the receptor nonfunctional The allele has a negative effect upon T cell function, but appears to protect against smallpox and HIV HIV has no receptor to bind to and cannot enter the cell This allele is found in 14% of Europeans HIV can impose selective pressure for CCR5-Δ32, increasing the frequency of this allele in human populations (Sullivan et al. 2001) Amy D. Sullivan et al. 2001. The coreceptor mutation CCR5Δ32 influences the dynamics of HIV epidemics and is selected for by HIV. Proc Natl Acad Sci USA. 98: 10214–10219. Frequency shift in CCR5- D32 allele Fr Modes of Selection Selection on discrete vs continuous characters When phenotypes fall into discrete classes (like Mendel’s yellow vs green peas), we can think of selection acting directly on genotypes Such discrete traits are usually coded by one or a few genes (loci) However, when phenotypes fall into a continuum (continuous traits such as those Darwin examined, beak shape, body size), you will see changes in the distributions of traits (next slide) –such traits are called “quantitative traits,” coded by many loci DIFFERENT MODES OF SELECTION (Herron & Freeman Section 9.6) Directional Selection on a discrete trait Directional Selection on a continuous trait Directional Selection: Selection changes frequency of alleles in one direction die shift in distribution Directional Selection: Selection changes frequency of alleles in one direction Mock Example Directional Selection on a discrete trait Generation 1 Generation 2 . . . Generation X AA 0.80 0.60 . . . 0.04 Aa 0.15 0.30 . . . 0.16 aa 0.05 0.10 . . . 0.75 Directional Selection on a discrete trait Directional Selection on a discrete trait How will selection act if A is dominant and selection favors red? AA aa How will selection act if A is not dominant, but Aa intermediate? AA Aa Aa aa Under which situation will the “a” allele be preserved in the population longer? How will selection act if A is dominant and selection favors red? AA Aa aa Selection will act on AA and Aa in the same manner How will selection act if A is not dominant, but Aa intermediate? AA Aa aa Selection will act more strongly on AA Under which situation will the “a” allele be preserved in the population longer? The first above, as the “a” allele is more effectively masked in the heterozygote state Directional Selection on a discrete trait Selection on Dominant vs Recessive Alleles Herron & Freeman Selection against recessive allele Set fitness of genotypes to: WAA = 1, WAa = 1, Waa = 1 – s (s = selection coefficient) Selection against dominant allele Set fitness of genotypes to: WAA = 1 - s, WAa = 1 - s, Waa = 1 (AA and Aa have same phenotype) Selection on Dominant vs Recessive Alleles What Happens? Selection against recessive allele In Allele A1, set fitness of genotypes to: WAA = 1, WAa = 1, Waa = 1 – s (s = selection coefficient) Selection against dominant allele Set fitness of genotypes to: WAA = 1 - s, WAa = 1 - s, Waa = 1 (AA and Aa have same phenotype) Selection on Dominant vs Recessive Alleles Selection against recessive allele: As recessive allele becomes rare, rate of its disappearance slows down As homozygote recessive allele becomes rare, most are in the heterozygous state and are masked from selection Selection against dominant allele: Dominant allele that is disfavored by selection is removed quickly from the population Directional Selection: Selection changes frequency of alleles in one direction Genetic diversity is reduced (knock out maladapted genotypes) Examples: Drug-resistance in tuberculosis and HIV Agriculture: artificial selection for specific traits Sexual selection for a trait (e.g. for bright plumage) Directional Selection Directional Selection during Domestication Rapid evolution under intense directional selection for large and numerous kernals Selection by humans for a few regulatory genes (affecting transcription) Reduced genetic diversity in domesticated corn Morphological differences between teosinte and maize Maize with tb1 knocked out • Has branching patterns like teosinte maize teosinte Corn Genes selected for in Corn F1 Hybrid Teosinte Major morphological difference are due to directional selection on 5 genes Genes: • Teosinte branched1 (tb1): single mutation affects branching and inflorescence • Regulator of tb1 • tga glume (outer coating) reduction on chromosome X • teosinte – ~8-12 kernels F1 hybrid 8 rows, corn 20+rows Evidence for selection in 2-4% genes, ~1200 genes Hardy Weinberg revisited Generation 1 Generation 2 Generation 3 AA Aa aa 250 750 900 500 200 90 250 50 10 (1) Is this population remaining in HW Equilibrium from generation to generation? (2) What might be going on? Hardy Weinberg revisited Generation 1 Generation 2 Generation 3 AA Aa aa 250 750 900 500 200 90 250 50 10 (1) Is this population remaining in HW Equilibrium from generation to generation? ***check if (a) alleles frequencies change across generations and (b) whether the allele frequencies do not yield the predicted genotype frequencies (2) What might be going on? AA Aa Generation 1 250 (freq = 250/1000) 500 Generation 2 750 (freq = 750/1000) 200 Generation 3 900 (freq = 900/1000) 90 First convert # of individuals to frequency of genotypes aa 250 50 10 (1) Is this population remaining in HW Equilibrium from generation to generation? NO Gen1 Freq of A (p) = 0.25 + (0.50/2) = 0.50 Freq of a (q) should be = 1 -0.50 = 0.50 Gen2 Freq of A (p) = 0.75 + (0.20/2) = 0.85 Freq of a (q) = 1-0.85 = 0.05 + (0.20/2) = 0.15 Gen3 - can do same calculations **Allele and genotype frequencies are changing across generations: freq of A is increasing across the 3 generations **Genotype frequencies deviated from expectations based on allele frequencies: freq of A is 0.5 in Gen1, so expect AA =0.25 at Gen2, (2) What might be going on? Directional selection favoring A? Hardy Weinberg revisited AA Aa aa 0.44 0.56 0 (1) Is this population in HW Equilibrium? (2) Are the HW expectations the same as the genotype frequencies above? Hint: Calculate the genotype frequencies from the # above, and then calculate allele frequencies. Use the allele frequencies to calculate HW expectations. (3) What might be going on? Hardy Weinberg revisited AA Aa aa 0.40 0.60 0 (1) Is this population in HW Equilibrium? NO (2) Are the HW expectations the same as the genotype frequencies above? NO Freq of A (p) = 0.40 + (0.60/2) = 0.70 Freq of a (q) should be = 1 - 0.70 = 0.30 Based on HW expectations, freq of aa should = 0.3 x 0.3 = 0.09 aa is missing! Negative selection acting against aa? aa might be a homozygous recessive lethal Stabilizing Selection (on discrete or continuous traits) die Selection acts against the extremes (favors intermediate trait) =purifying selection The average trait value stays the same Genetic diversity is reduced This is going on all the time, as deleterious alleles are getting selected out of the population Examples: Selection against new harmful mutations Selection for an optimal number of fingers Selection for optimal body size Example of Stabilizing Selection Selection for an optimal number of eggs Disruptive Selection (on discrete or continuous traits) Selection favors the extremes Genetic diversity is increased (help maintain genetic variation; knock out alleles that code for intermediate traits) Can lead to formation of new species Examples: Niche Partitioning (Specialization on different resources, food) Differences in habitat use Will discuss more in lecture on Speciation Sexual selection for different traits (blue birds mate with blue, red birds mate with red)—intermediate colors selected against Disruptive Selection Balancing Selection (on discrete or continuous traits) Balancing Selection is a generic term to refer to any type of selection that acts to maintain genetic variation in a population There are different mechanisms: Fluctuating selection, selection favoring heterozygote (=overdominance) Examples: Selection for heterozygotes (sickle cell anemia, and cystic fibrosis) Selection for different traits in different environments Selection for different traits at different times (fluctuating selection) Balancing Selection An Example of one mechanism: Overdominance: Selection favoring the heterozygote Generation 1 Generation 2 Generation 3 AA 0.25 0.20 0.10 Aa 0.50 0.60 0.80 aa 0.25 0.20 0.10 Genetic diversity is maintained in a population in this case because the heterozygote maintains both alleles Example: Sickle Cell anemia Balancing Selection An Example of one mechanism: Overdominance: Selection favoring the heterozygote Generation 1 Generation 2 Generation 3 AA 0.25 0.20 0.10 Aa 0.50 0.60 0.80 aa 0.25 0.20 0.10 In this case, allele frequencies (of A and a) do not change. However, the population did go out of HW equilibrium because you can no longer predict genotypic frequencies from allele frequencies For example, p = 0.5, p2 = 0.25, but in Generation 3, the observe p2 = 0.10 Balancing Selection An Example of one mechanism: Overdominance: Selection favoring the heterozygote Generation 1 Generation 2 Generation 3 AA 0.25 0.20 0.10 Aa 0.50 0.60 0.80 aa 0.25 0.20 0.10 Genetic diversity is maintained in a population in this case because the heterozygote maintains both alleles Example: Sickle Cell anemia Balancing Selection An Example of one mechanism: Overdominance: Selection favoring the heterozygote Generation 1 Generation 2 Generation 3 AA 0.25 0.20 0.10 Aa 0.50 0.60 0.80 aa 0.25 0.20 0.10 Why is this NOT stabilizing selection? In Stabilizing Selection genetic diversity decreases as the population stabilizes on a particular trait value (a quantitative trait, like body size, height, intermediate color –of a multilocus nature) Sickle cell anemia example of balancing selection Reduced ability of red blood cells to carry oxygen (mutation in HbS gene, makes defective hemoglobin protein) HAHA: homozygous, no effects of disease (sickle cell anemia) HAHs: heterozygotes, suffer mild effects HsHs: homozygous, usually die before reproduction Sickle cell anemia When Malaria is present HAHs advantage: red blood cells with abnormal hemoglobin tend to sickle when infected by parasite and are culled out 72% HAHA; 25% HAHs; and 2.2% HsHs (die young) Malaria not present AA advantage 99.999% HAHA; 0.001% HAHs; and 0% HsHs With malaria, balancing selection for the heterozygote Without malaria, directional selection for HAHA So, how does one detect Natural Selection in a population? Many types of tests The challenge is distinguishing natural selection from signatures of Genetic Drift Codon Bias In the case of amino acids Mutations in Position 1, 2 lead to Amino Acid change Mutations in Position 3 often don’t matter (1) Simplest Test: Ka/Ks Test Nonsynonymous substitution rate Synonymous substitution rate Ka Ks >1 Need coding sequence (sequence that codes proteins) Ks is used here as the “control”, proxy for neutral evolution A greater rate of nonsynonymous substitutions (Ka) than synonymous (Ks) is used as an indication of selection (Ka/Ks >1) Substitution rate: out of all the possible number of mutations, how many fixed (1) Ka/Ks Test Nonsynonymous substitution rate Synonymous substitution rate Ka Ks >1 In the absence of selection, you’d expect Ks = Ka, and for Ka/Ks = 1 A greater rate of nonsynonymous substitutions (Ka) than synonymous (Ks) is used as an indication of positive selection (Ka/Ks >1) If Ks > Ka, or Ka/Ks < 1, that would suggest purifying selection, that selection might be preserving amino acid composition Summary (1) Of the evolutionary forces that can disrupt HW Equilibrium, Natural Selection is the only one that is adaptive (2) Selection occurs through differential reproduction which is NOT random (3) Natural selection requires genetic variation upon which it can act (4) There are different ways in which selection can act (favoring the extremes, or heterozygotes, or unidirectional) Concepts Natural Selection Fitness Directional, Stabilizing, Disruptive Selection Balancing Selection Selection on dominant vs recessive alleles The following are numbers of red, pink, and white flowers in a population. Generation 1: Generation 2: Generation 3: (A1A1) Red (A1A2) Pink (A2A2) White 400 600 500 200 100 0 400 300 500 1. Is the population above in Hardy-Weinberg Equilibrium? (A) Yes (B) No (C) Yes in the first generation, but No in the third generation (D) No in the first generation, but Yes in the third generation 2. What are the frequencies of alleles and genotypes at Generation 3? (A)A1: 0.50, A2: 0.50 A1A1: 0.25, A1A2: 0.50, A2A2: 0.25 (B)A1: 0.65, A2: 0.35 A1A1: 0.60, A1A2: 0.10, A2A2: 0.30 (C) A1: 0.50, A2: 0.50 A1A1: 0.50, A1A2: 0, A2A2: 0.50 (D) A1: 0.50, A2: 0.50 A1A1: 0.25, A1A2: 0, A2A2: 0.25 3. In the previous example (from question #1), what might be going on? (A) Genetic Drift (B) Balancing Selection (C) Recombination (D) Directional Selection (E) Disruptive Selection 4. What would Darwin say? (A) Genetic drift is also an important evolutionary force that occurs through random survival and reproduction (B) Individuals pass alleles onto their offspring intact (inheritance is particulate) (C) Selection acts on genetic variation such as Mutations (D) Selection acts on differential fitness of genetically distinct offspring (E) Evolution occurs at the level of the individual Answers: 1B 2C 3E 4D