EPIGENETICS Textbook
... – NOT methylated on active and silent genes – EXCEPTIONS: • Silencing on X chromosome • When cells differentiate • Pathological processes, e.g., inactivation of tumor suppressor genes in some cancers ...
... – NOT methylated on active and silent genes – EXCEPTIONS: • Silencing on X chromosome • When cells differentiate • Pathological processes, e.g., inactivation of tumor suppressor genes in some cancers ...
Key concepts_Regulation of transcription in
... ubiquitylation, and poly(ADP)ribosylation events. Specific enzymes exist for each of these modifications: the enzymes are highly specific for individual amino acid residues on individual histone molecules. There is also cross-talk between such modifications. Specific readers of these markings exist ...
... ubiquitylation, and poly(ADP)ribosylation events. Specific enzymes exist for each of these modifications: the enzymes are highly specific for individual amino acid residues on individual histone molecules. There is also cross-talk between such modifications. Specific readers of these markings exist ...
Analytical and Chromatography - Sigma
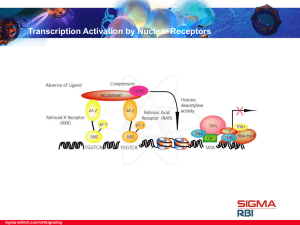
... change in the receptors causes the dissociation of the corepressors and subsequent recruitment and binding of coactivators such as Steroid Receptor Coactivator-1 (SRC-1), Glucocorticoid Receptor Interacting Protein-1 (GRIP-1), CREB Binding Protein (CBP) and CBP-Associated Factor (PCAF). This recepto ...
... change in the receptors causes the dissociation of the corepressors and subsequent recruitment and binding of coactivators such as Steroid Receptor Coactivator-1 (SRC-1), Glucocorticoid Receptor Interacting Protein-1 (GRIP-1), CREB Binding Protein (CBP) and CBP-Associated Factor (PCAF). This recepto ...
Multiple choice questions
... DNA-binding proteins Are usually monomeric Interact with DNA by ionic bonds Contain DNA-binding motifs Can regulate gene expression Can be isolated by affinity chromatography ...
... DNA-binding proteins Are usually monomeric Interact with DNA by ionic bonds Contain DNA-binding motifs Can regulate gene expression Can be isolated by affinity chromatography ...
DNA Structure
... between the nucleosome and the DNA with which it is associated. Three distinct types of change can occur Remodeling, in the strict sense, involves a change in the structure of the nucleosome, but no change in its position. Sliding, or cis-displacement, physically moves the nucleosome along the DNA T ...
... between the nucleosome and the DNA with which it is associated. Three distinct types of change can occur Remodeling, in the strict sense, involves a change in the structure of the nucleosome, but no change in its position. Sliding, or cis-displacement, physically moves the nucleosome along the DNA T ...
Epigenetics - Louisiana State University
... What is Epigenetics? • “the study of mitotically and/or meiotically heritable changes in gene function that can not be explained by changes in DNA sequences” – (Riggs et al. 1996) • originally was coined to describe “all the developmental events leading to a mature organism from a fertilized zygote ...
... What is Epigenetics? • “the study of mitotically and/or meiotically heritable changes in gene function that can not be explained by changes in DNA sequences” – (Riggs et al. 1996) • originally was coined to describe “all the developmental events leading to a mature organism from a fertilized zygote ...
Chromatin Structure and Its Effects on Transcription
... DNA/core histones when contrasted with: – Uncatalyzed DNA exposure in nucleosomes – Simple nucleosome sliding along a DNA stretch ...
... DNA/core histones when contrasted with: – Uncatalyzed DNA exposure in nucleosomes – Simple nucleosome sliding along a DNA stretch ...
Binary Switches in Gene Expression: The Histone Code
... lies outside of the DNA itself. . This system relies on packaging DNA into a DNA-histone complex called chromatin, which is the physiological substrate of all cellular processes involving the DNA. The dynamic change of the three-dimensional architecture of chromatin makes certain genes more readily ...
... lies outside of the DNA itself. . This system relies on packaging DNA into a DNA-histone complex called chromatin, which is the physiological substrate of all cellular processes involving the DNA. The dynamic change of the three-dimensional architecture of chromatin makes certain genes more readily ...
Chromatin Structure and Function
... B) Non-histone proteins 1) Chromosome structure 4 core histones found in 2) Gene regulation proteins Nucleosomes - 146 bp of DNA wrapped around an octomer of 2 (H2A, H2A, H3, H4) ...
... B) Non-histone proteins 1) Chromosome structure 4 core histones found in 2) Gene regulation proteins Nucleosomes - 146 bp of DNA wrapped around an octomer of 2 (H2A, H2A, H3, H4) ...
Acetyl-Histone H4 (Lys5) Polyclonal Antibody
... consists of four core histone proteins (H2A, H2B, H3, and H4), which undergo multiple types of post-translational modifications, including acetylation, phosphorylation, methylation, and ubiquitination (1,2). Histone acetylation occurs mainly on the amino-terminal tail domains of histones H2A (Lys5), ...
... consists of four core histone proteins (H2A, H2B, H3, and H4), which undergo multiple types of post-translational modifications, including acetylation, phosphorylation, methylation, and ubiquitination (1,2). Histone acetylation occurs mainly on the amino-terminal tail domains of histones H2A (Lys5), ...
No Slide Title
... • Gcn5p is a transcriptional activator of many genes in yeast. It is also a HAT. • PCAF (P300/CBP associated factor) is a HAT and is homologous to yeast Gcn5p. • P300 and CBP are similar proteins that interact with many transcription factors (e.g. CREB, AP1 and MyoD). • P300/CBP are needed for activ ...
... • Gcn5p is a transcriptional activator of many genes in yeast. It is also a HAT. • PCAF (P300/CBP associated factor) is a HAT and is homologous to yeast Gcn5p. • P300 and CBP are similar proteins that interact with many transcription factors (e.g. CREB, AP1 and MyoD). • P300/CBP are needed for activ ...
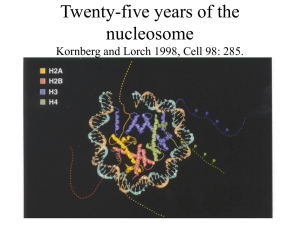
Twenty-five years of the nucleosome Kornberg and Lorch 1998, Cell
... 2. Immunocytochemistry- observe phospho-H3 throughout chromosomes during cell division Thus, this must play a role is chromosome condensation during mitosis 3. Models1. Phosphorylation + acetylation allows activation of gene expression, depending on context 2. Phospho-H3 loosens chromatin, enhancin ...
... 2. Immunocytochemistry- observe phospho-H3 throughout chromosomes during cell division Thus, this must play a role is chromosome condensation during mitosis 3. Models1. Phosphorylation + acetylation allows activation of gene expression, depending on context 2. Phospho-H3 loosens chromatin, enhancin ...
Glossary
... “chromatin”, whose structural alteration influences transcription of genes which are incorporated into/adjacent to the chromatin, thus chromatin plays important roles in gene regulation. ...
... “chromatin”, whose structural alteration influences transcription of genes which are incorporated into/adjacent to the chromatin, thus chromatin plays important roles in gene regulation. ...
Histone acetyltransferase

Histone acetyltransferases (HATs) are enzymes that acetylate conserved lysine amino acids on histone proteins by transferring an acetyl group from acetyl CoA to form ε-N-acetyllysine. DNA is wrapped around histones, and, by transferring an acetyl group to the histones, genes can be turned on and off. In general, histone acetylation increases gene expression.In general, histone acetylation is linked to transcriptional activation and associated with euchromatin. When it was first discovered, it was thought that acetylation of lysine neutralizes the positive charge normally present, thus reducing affinity between histone and (negatively charged) DNA, which renders DNA more accessible to transcription factors. Research has emerged, since, to show that lysine acetylation and other posttranslational modifications of histones generate binding sites for specific protein–protein interaction domains, such as the acetyllysine-binding bromodomain. Histone acetyltransferases can also acetylate non-histone proteins, such as nuclear receptors and other transcription factors to facilitate gene expression.