* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 高 茂 傑 (Mou-Chieh Kao)
Survey
Document related concepts
Public health genomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Point mutation wikipedia , lookup
Transcript
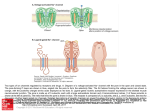
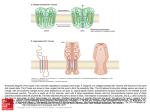
高 茂 傑 (Mou-Chieh Kao) Assistant professor 生科一館 307 室 Tel: (03)5715131 轉 42472 (office), 轉 33456 (laboratory) E-mail: [email protected] 學歷: 美國康乃爾大學博士 (Cornell University, U.S.A.) 專長: 微生物學、生化學 (Microbiology/Biochemistry) Academic Background Postdoctoral Fellow (2000-2005) - The Scripps Research Institute, La Jolla, CA, U.S.A. Ph.D. (1995-2000) - Cornell University, Ithaca, NY, U.S.A. M.S. (1995-1998) - Cornell University, Ithaca, NY, U.S.A. B.S. (1988-1992) - National Taiwan University, Taipei, Taiwan Research Interests My research interests are centered on understanding the mechanisms of complex I and developing a strategy to reestablish the functionality of complex I thus lead to the treatment of human mitochondrial diseases caused by the defects of this enzyme complex. Mammalian complex I is the most intricate membrane-bound enzyme known to date, being composed of at least 46 unlike subunits. Of these, 7 subunits are encoded by mitochondrial DNA. It is the point of entry for the major fraction of electrons that traverse the respiratory chain eventually resulting in the reduction of oxygen. Coupled to the electron transfer, protons are pumped from the matrix side to the intermembrane space of mitochondria. In contrast, the bacterial version of complex I, also called NDH-1, is located in the plasma membrane and contains only 13 or 14 different subunits. Despite the difference in the size and complexity, complex I and NDH-1 share many properties in terms of structure and function. In eukaryotes, it is not easy to manipulate mitochondrial DNA directly. In contrast, gene manipulation procedures of bacterial DNA are well advanced. Previously, my colleagues and I have used Paracoccus denitrificans and Escherichia coli NDH-1 as models to study the function-structure relations of this complex and investigated the events that are associated with pathogenic mutations of human complex I. Our discoveries from these studies demonstrate that bacterial model systems could be useful for further understanding of this complicated mammalian enzyme, and provide an alternative way to gain insight into human pathogenesis derived from malfunction of this complex. In this respect, I am interested in the following studies: In the bacterial system: Clarifying functional roles of amino acid residues of interest, especially those amino acids corresponding to residues involved in human mitochondrial diseases, in the membrane domain ND subunits of complex I Searching proton-translocating site(s) and quinone-binding site(s) Answering the question “Can Complex I pump not only protons but also sodium ions?” Determining the role of component subunits of NDH-1 in the assembly of the complex Using non-crystallization approaches to study subunit interactions and subunit arrangements in complex I Such a thorough investigation of subunit interactions and subunit arrangements should provide clues about how electron transfer and proton translocation are coordinated in this complicated complex. Along with answering these questions, it is my hope that the knowledge gained through these studies will help us to have a better understanding of the physiological-pathological dysfunction associated with human complex I diseases. In the mammalian system: Certainly, not every aspect of human diseases can be reflected in the bacterial system. Therefore, a better understanding of complex I in animal mitochondria is absolutely necessary. In this respect, I am interested in conducting studies in the developing new strategies for studies of mitochondrially encoded complex I subunits and for treatment of complex I dysfunction. Recently, human complex I, especially the mitochondrially encoded subunits, has increasing medical importance because mutations of the mtDNA have been associated with a wide variety of human neurodegenerative diseases as well as the process of aging. These seven mtDNA-encoded subunits of complex I are extremely hydrophobic and have multiple membrane-spanning regions. At present no effective remedies have been established for complex I deficiencies. Therefore, developing methods to correct complex I defects seems important. Gene therapy is by nature a powerful treatment method. However, a severe hurdle in the development of mitochondrial gene therapy is that the no practical method is currently available to transfect mammalian mitochondria with exogenous nucleic acids. In addition, the lack of such an efficient delivery system has also made the studies of the effects of mutations on the structure, assembly, and function of the mitochondrial complexes extremely difficult. One possible approach to overcome the aforementioned obstacle is to apply a so called “allotopic expression” method, in which an engineered nucleus-localized version of an mtDNA-encoded gene is introduced to express the target protein in the cytoplasm, and the expressed transgene-encoded protein is then posttranslationally imported into mitochondria. Several studies have to be conducted before the above approach can be practically applied in complex I research (or other complexes with mtDNA-encoded subunits). These studies include: Investigating factors necessary for successful gene transfer of a hydrophobic protein that is usually mitochondrial-encoded into the nucleus and developing a delivery system which has high possibility for long term expression of transgenes Considering the differences in the codon preference existing between the mitochondrial and nuclear genomes to redesign and recode the mitochondrial genes of interests for nuclear expression Changing the local hydrophobicity of the subunits to facilitate the protein transfer Surveying the use of the leader sequence as the mitochondrial targeting presequence to direct the protein to the mitochondria to achieve the correct localization Overcoming these challenges requires a solid knowledge of the mitochondrial protein transport machinery. Once the correct expression of the selected, relocated mtDNA-encoded subunit is established and the expressed subunit can restore the NADH oxidase activity in the complex I-deficient culture cells, in vivo study utilizing the murine model system can be followed. The final goal is to develop a promising strategy for the treatment of human diseases caused by complex I defects. In addition, it is anticipated that such a strategy might be applicable for therapies of dysfunction of other enzyme complexes involved in oxidative phosphorylation. Recent Publications Kao, M.-C., Nakamaru-Ogiso, Matsuno-Yagi, A., and Yagi, T. (2005). “Characterization of the membrane domain subunit NuoK (ND4L) of the NADH-quinone oxidoreductase from Escherichia coli”. Biochemistry 44:9545-9554. Kao, M.-C., Di Bernardo, S., Nakamaru-Ogiso, E., Miyoshi, H., Matsuno-Yagi, A., and Yagi, T. (2005). “Characterization of the membrane domain subunit NuoJ (ND6) of the NADH-quinone oxidoreductase from Escherichia coli by chromosomal DNA manipulation”. Biochemistry 44:3562-3571. Kao, M.-C., Di Bernardo, S., Perego, M., Nakamaru-Ogiso, E., Matsuno-Yagi, A., and Yagi, T. (2004). “Functional roles of four conserved residues in the membrane domain subunit NuoA of the proton-translocating NADH-quinone oxidoreductase from Escherichia coli”. J. Biol. Chem. 279:32360-32366. Kao, M.-C., Mastuno-Yagi, A., and Yagi, T. (2004) “Subunit proximity in the H+translocating NADH-quinone oxidoreductase probed by zero-length crosslinking”. Biochemistry 43:3750-3755. Yagi, T., Di Bernaldo, S., Nakamaru-Ogiso, E., Kao, M.-C., Seo, B.B., and MatsunoYagi, A. (2004) “NADH dehydrogenase (NADH-quinone oxidoreductase)”. In Respiration in Archaea and Bacteria. (ed: Zannoni, D.) pp15-40, Kluwer Publishings, Dordrecht. Kao, M.-C., Di Bernardo, S., Matsuno-Yagi, A., and Yagi, T. (2003) Characterization and topology of the membrane domain Nqo10 subunit of the proton-translocating NADH-quinone oxidoreductase of Paracoccus denitrificans”. Biochemistry 42:45344543. Kao, M.-C., Di Bernardo, S., Matsuno-Yagi, A., and Yagi, T. (2002) “Characterization of the membrane domain Nqo11 subunit of the proton-translocating NADH-quinone oxidoreductase of Paracoccus denitrificans”. Biochemistry 41:4377-4384. Yagi, T., Seo, B. B., Di Bernardo, S., Nakamaru-Ogiso, E., Kao, M.-C., and MatsunoYagi, A. (2001) “NADH Dehydrogenases: From Basic Science to Biomedicine”. J. Bioenerg. Biomembr. 33:233-242.