* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Cortical Parcellations of the Macaque Monkey
Survey
Document related concepts
Lateralization of brain function wikipedia , lookup
Node of Ranvier wikipedia , lookup
Eyeblink conditioning wikipedia , lookup
Aging brain wikipedia , lookup
Emotional lateralization wikipedia , lookup
Neuroesthetics wikipedia , lookup
Surface wave detection by animals wikipedia , lookup
Feature detection (nervous system) wikipedia , lookup
Human brain wikipedia , lookup
Cognitive neuroscience of music wikipedia , lookup
Neuroeconomics wikipedia , lookup
Neural correlates of consciousness wikipedia , lookup
Cortical cooling wikipedia , lookup
Inferior temporal gyrus wikipedia , lookup
Transcript
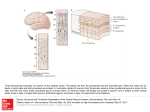
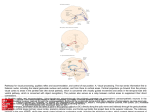
Cerebral Cortex October 2012;22:2227– 2240 doi:10.1093/cercor/bhr290 Advance Access publication November 2, 2011 Cortical Parcellations of the Macaque Monkey Analyzed on Surface-Based Atlases David C. Van Essen, Matthew F. Glasser, Donna L. Dierker and John Harwell Department of Anatomy & Neurobiology, Washington University School of Medicine, St. Louis, MO 63110, USA Address correspondence to David C. Van Essen. Email: [email protected]. Surface-based atlases provide a valuable way to analyze and visualize the functional organization of cerebral cortex. Surfacebased registration (SBR) is a primary method for aligning individual hemispheres to a surface-based atlas. We used landmark-constrained SBR to register many published parcellation schemes to the macaque F99 surface-based atlas. This enables objective comparison of both similarities and differences across parcellations. Cortical areas in the macaque vary in surface area by more than 2 orders of magnitude. Based on a composite parcellation derived from 3 major sources, the total number of macaque neocortical and transitional cortical areas is estimated to be about 130--140 in each hemisphere. Keywords: architectonic, areas, maps, registration, retinotopy Introduction Primate cerebral cortex contains a complex mosaic of areas that differ in architecture, connectivity, topographic organization, and/or functional characteristics (Felleman and Van Essen 1991; Kaas 2005). The macaque monkey is the most intensively studied nonhuman primate, yet despite a century’s effort, an accurate, consensus cortical parcellation is lacking for most of macaque cortex. A primary reason is that differences between neighboring areas are often subtle when assessed by any of the available methods. Moreover, many individual areas show internal heterogeneity in architecture, connectivity, and/or functional characteristics. Another confound is that the size and shape of each cortical area can vary 2-fold or more across individuals (Van Essen et al. 1984; Maunsell and Van Essen 1987). Finally, the complexity and variability of cortical convolutions are an impediment to visualizing results and making accurate comparisons across individuals. Many of the challenges posed by cortical convolutions and their variability can be addressed using surface reconstructions of individual hemispheres combined with surface-based registration (SBR) to an atlas (Van Essen and Dierker 2007). Landmark-SBR (Van Essen et al. 1998, 2001, 2005; Van Essen 2005) is a specific type of SBR that uses explicitly delineated landmark contours to constrain the registration from a source (individual) to a target (atlas) surface. Here, we introduce the landmark vector difference (LVD) algorithm for Landmark-SBR, which achieves accurate alignment even when it entails large local deformations. The LVD method is useful for registering individuals to an atlas and for inter-atlas registration within a species or between species (e.g., for macaque--human comparisons). We applied this approach to an analysis of cortical parcellations in the macaque. Data from numerous published studies were registered to the macaque F99 atlas (Van Essen 2002a; Van Essen and Dierker 2007) using the LVD algorithm or older Landmark-SBR methods. This facilitates identification of discrepancies as well as commonalities among different parcellation The Author 2011. Published by Oxford University Press. All rights reserved. For permissions, please e-mail: [email protected] schemes. Informative examples illustrated herein include areas of orbital and medial prefrontal cortex (OMPFC), the middle temporal (MT) area and nearby areas in the superior temporal sulcus (STS), and areas in medial and posterior parietal cortex. Another objective of this study was to estimate the total number of distinct cortical areas in the macaque. This number represents a fundamental aspect of brain organization in any mammalian species, yet has proven difficult to resolve even in intensively studied species like the macaque. Published parcellations that span most or all the cortical sheet span a wide range in the total number of areas. Here, we generated a composite parcellation in which the areas identified in each region are derived from a parcellation considered to be especially reliable in that region. We also consider how to assess the ‘‘granularity’’ of cortical parcellation in regions where the confounds of internal heterogeneity and subtlety of areal boundaries are particularly challenging (see Discussion). All atlas data sets are freely available and are useful for a variety of analyses and comparisons within and across species. Materials and Methods Surface-based visualization and analysis were carried out using Caret software (v5.616 and earlier versions) (Van Essen et al. 2001; Caret, http://brainvis.wustl.edu/wiki/index.php/Caret:About). Macaque F99 Atlas The macaque single-subject F99 atlas (Fig. 1A,C) includes cortical midthickness (layer 4) surfaces for the left and right hemispheres, derived from segmentation of a high-resolution (0.5 mm) T1-weighted MR scan of an individual adult rhesus macaque (Van Essen 2002a, 2002b). The midthickness representation (top panels) is advantageous because each unit of surface area is associated with a similar cortical volume for both gyral and sulcal regions (Van Essen and Maunsell 1980; Van Essen et al. 2001). Revised demarcations were made of the F99 atlas noncortical ‘‘medial wall’’ boundaries (black in Fig. 1C) with the aid of published atlases (Paxinos and Franklin 2000; Saleem and Logothetis 2007). The analysis of cortical surface area included all of neocortex plus several transitional regions (subicular complex) and pyriform cortex. The hippocampus proper (CA1-4) was excluded because it was not accurately segmented. To bring the left and right hemisphere surfaces into correspondence, 26 landmark contours were drawn in corresponding gyral or sulcal locations on the F99 native-mesh surfaces. These were projected to the ‘‘spherical standard’’ (distortion-reduced and geographically aligned) surfaces of each hemisphere. The right and mirror-flipped left hemisphere spherical landmarks were averaged and used as a target for registration of each hemisphere using the landmark pin and relax (LPR) algorithm (Van Essen 2005). This yielded standard-mesh surface representations (Saad et al. 2004) containing 73 730 nodes that are identified as being on the ‘‘74k_f99’’ mesh, reflecting both the number of surface nodes and the F99 target atlas. The left and right hemispheres are in good geographic correspondence on the 74k_f99 mesh, as assessed by visual inspection. More specifically, highlighting any given node in one hemisphere identifies a node in the other hemisphere that is very similar in its position relative to nearby gyral and sulcal landmarks, usually within 1--3 mm. This assessment was Figure 1. Landmarks for SBR displayed on the F99 atlas and an individual subject right hemisphere. (A) Lateral views of the F99 atlas midthickness (top), inflated (middle), and spherical (bottom) surfaces. Fifteen landmark contours are projected to the surface. (B) Corresponding lateral views of an individual macaque (Case CH from Kolster et al. 2009) generated using the FreeSurfer processing pipeline followed by a semiautomated pipeline in Caret. After conversion to Caret format, the individual-subject surface was inflated, mapped to a sphere, processed by multiresolution morphing to reduce distortions and rotated to a spherical standard orientation. A standard set of template landmark contours (originally generated on a different hemisphere and stored as contour points relative to that hemisphere’s spherical standard surface) was projected to the spherical standard surface and viewed on other configurations (pre-edit landmarks on the inflated surface). Manual editing yielded post-edit contours shown in the other panels. (C) Medial views of the atlas surface. Revised medial wall borders, compared with earlier version of the F99 atlas (Van Essen and Dierker 2007) include the boundary between neocortex or transitional cortex and basal forebrain, piriform cortex, and other non-neocortical structures. (D) Medial views of the individual surface. supported by computing a difference map between cortical folding (mean curvature) for the left versus the right hemisphere. This difference map confirmed that creases of high folding are generally in good agreement (within 2 mm). In a few locations, the mismatch between folding maps was larger, but these were places where genuine hemispheric differences in local folding patterns were evident. Versions of the left and right F99 midthickness surfaces were generated whose dimensions and surface area matched that of an average adult macaque. First, the F99 midthickness surfaces were registered to the ‘‘D99’’ atlas volume of Saleem and Logothetis (2007) using an F99-to-D99 affine transformation matrix generated by McLaren et al. (2009). In generating their 112RM-SL population-average adult rhesus MR volume, McLaren et al. (2009) registered each of 112 individual-subject MR scan to the D99 atlas volume by an affine transformation (after an initial rigid-body alignment). The scale factors used for the affine transformations were averaged separately for the 52 male and 32 female brains and for the x, y, and z dimensions. Applying the inverse of these scale factors to the F99 surfaces in D99 space yielded F99 atlas surfaces that represent actual rhesus macaque brain size averaged across sexes. FreeSurfer-to-F99 Registration Pipeline A semiautomated process was implemented to convert FreeSurfergenerated macaque surfaces into Caret file formats and to register the individual hemispheres to the F99 atlas. The FreeSurfer cortical segmentation process generates pial and white (gray-white boundary) surface representations that can be averaged to form a midthickness surface, shown for an exemplar individual hemisphere (Fig. 1B,D). Comparison of the F99 atlas (Fig. 1A,C) and the individual (Fig. 1B,D) midthickness surfaces (top row) reveals many shape features in common but numerous differences in detail. A set of 15 template landmark contours is shown on the midthickness (top) and inflated (middle) and spherical (bottom) configurations of the atlas surface (Fig. 1A,C). Corresponding template landmarks were projected to the individual subject’s surface (‘‘preedit’’ in Fig. 1B). These template landmarks were edited manually, yielding ‘‘postedit’’ landmark contours that represent corresponding geographic (gyral and sulcal) features in the source (individual) and target (atlas) surfaces. These landmarks 2228 Macaque Surface-Based Cortical Parcellations d Van Essen et al. were used to constrain registration of the individual sphere to the atlas sphere using the LVD algorithm described in the next section. An ‘‘FSto-F99’’ tutorial document and associated data set illustrating this process is available (http://sumsdb.wustl.edu/sums/directory.do?id=5255961). LVD Registration The LVD method of Landmark-SBR is described in detail because of its broad utility (see Introduction) and because understanding the process can aid in cases that require customized adjustment of registration parameters. Although similar in overarching objective to the diffeomorphic landmark-based spherical registration algorithm of Glaunes et al. (2004) and the elastic energy algorithm of Joshi et al. (2007) and Pantazis et al. (2010), it is very different in its algorithmic approach (see Discussion). The LVD algorithm deforms a source (individual) spherical mesh until its landmarks match the target (atlas) landmarks, while minimizing local distortions in regions between neighboring landmarks. Alignment is achieved in an interactive coarse-to-fine sequence involving multiple ‘‘stages’’ and multiple ‘‘cycles’’ within each stage. In each cycle, the vector difference between the current location of source and corresponding target landmark points provides the driving force for the deformation. The vector difference is distributed across nearby surface nodes by spatial smoothing of the initial deformation field. Key steps in the process are illustrated in Figure 2 (same individual and atlas surfaces as in Fig. 1); see also http://brainvis.wustl.edu/wiki/index.php/ Caret:Operations/SurfaceBasedRegistration. An initial registration step is to resample landmark contours so that corresponding source and target landmarks each contain the same number of points (Fig. 2A,B). Each source landmark point is then intercalated into the nearest tile of the standard-mesh source sphere. This yields a landmark-intercalated source tessellation whose topology reflects the newly incorporated nodes. This is illustrated in Figure 2C for a small surface patch in the calcarine sulcus (red nodes are the intercalated landmark points). Similarly, a landmark-intercalated target tessellation is generated by intercalating each target landmark point into the nearest tile of the target sphere (Fig. 2D). At the outset of each cycle, the differences in 3D coordinates of corresponding landmark nodes in the source and target spheres are Figure 2. Key stages in LVD registration, using an exemplar individual (source) hemisphere registered to the F99 atlas target. (A, B) Landmark contours on source (case CH of Kolster et al. 2009) and target (F99 atlas) spheres after resampling to match the number of contour points in corresponding source and target landmarks. (C) Expanded views of surface mesh in the calcarine sulcus, showing landmark nodes (red) intercalated into the 74 k node spherical mesh. The surface was extensively smoothed and projected back to a sphere, which caused compression of tiles in the vicinity of the 12 nodes that have only 5 (instead of 6) neighbors. This was important in order to avoid smoothing-induced distortions at intermediate stages of the deformation process. (D) Expanded views of target nodes intercalated into an equivalent mesh at the locations of the resampled target landmarks. Red arrows in C and D indicate displacement vectors computed for the highlighted landmark nodes (black). (E) Y-component of the smoothed displacement field. Yellow--orange signifies nodes to be displaced to the left (positive y), blue signifies nodes to be displaced to the right. (F) Deformed surface mesh at the end of stage 1, which includes 2 cycles of displacement, smoothing, and morphing, resulting in displacement of the mesh about one-third of the distance to the target. To transfer these results to the native-mesh surface in preparation for the next stage, each node from the native-mesh source sphere is projected to the initial landmark-intercalated source sphere (panel C) then unprojected based on its location in the deformed landmark-intercalated source sphere (panel F). (The projection and unprojection steps use barycentric coordinates to specify the position of nodes in one mesh relative to the nearest tile of the other mesh—see Fig. 3E). This deformed native-mesh sphere becomes the input source sphere for the next stage of the LVD process. (G) A fresh surface mesh at the beginning of stage 2. Landmark nodes preserve their location from the end of stage 1 (panel F) but are intercalated into an undistorted spherical mesh (as in panel C but with shifted location of landmark nodes). (H) Alignment of source calcarine sulcus landmarks after 5 stages of LVD registration. The similarity of landmark node positions in panels D and H signifies successful registration. computed (red arrows for highlighted nodes in Fig. 2B) and assigned as vector x, y, and z components for each intercalated landmark node. This defines a displacement field on the source surface in which only the intercalated landmark nodes have nonzero values; the displacement field is stored as a Caret ‘‘surface-shape’’ file with separate fields for the x, y, and z components. This sharply peaked displacement field is smoothed by repeatedly averaging each node’s vector component value with that of its neighbors. The number of smoothing steps is specified separately for each cycle and stage; it generally entails many iterations for early coarse registration stages and fewer iterations for the later fine-tuning stages. In panels C--E, the smoothed displacement field for the y-axis (horizontal) component (shown in Fig. 2E) diverges, pointing leftward (orange) on the left and rightward (blue) on the right; there is also a large upward component (z-axis) (not shown). Surface nodes are displaced in proportion to the magnitude and direction of this displacement field. The mesh is then projected back to a spherical configuration, smoothed, and subjected to spherical morphing (Drury et al. 1996) to reduce local distortions, while holding the landmark nodes in their displaced locations. After 2 cycles of displacement and landmark-constrained smoothing and morphing, source landmark nodes are shifted substantially toward the target nodes (i.e., arrows in 2F are ~30% shorter than in 2C). To bring the landmark nodes progressively closer to their targets without creating unacceptably large local distortions, additional stages are carried out in which the deformed landmark points are converted back to contours (strings of contour points) and intercalated from their displaced positions into a fresh (undistorted) standard-mesh sphere (Fig. 2G). In this exemplar case, 5 stages of displacement, smoothing, and morphing (Fig. 2C--F), brought the deformed source landmark nodes (Fig. 2H) acceptably close to their desired target locations (Fig. 2D). The deformed native-mesh source sphere from the final stage is used to create a ‘‘deformation map’’ that encodes the precise relationship between source and target meshes. This is illustrated in Figure 3 using low-resolution views (top row) and a zoomed surface patch on the prelunate gyrus (bottom row). A highlighted node on the ‘‘native’’ midthickness surface mesh (Fig. 3A) is also highlighted in the ‘‘preregistration’’ spherical surface (Fig. 3B) and on the deformed ‘‘postregistration’’ native-mesh sphere (Fig. 3C), with landmarks now in register with the atlas target landmarks (Fig. 3D). This establishes a precise spatial relationship between each node on the atlas sphere and a surface tile in the deformed native mesh sphere as shown in a highly expanded patch in Figure 3E. The deformation map file encodes these relationships using barycentric coordinates to describe the position of each target node within the nearest tile of the deformed source spherical mesh. In the resampled midthickness surface, each atlas node has an appropriately interpolated coordinate location based on the coordinates of the associated tile in the native-mesh midthickness surface. The resampled surface preserves the shape of the nativemesh midthickness surface but has a more regular tessellation pattern (lower panel) and is slightly smoother as a result of the interpolation process (Fig. 3F). Thus, the deformation map encodes both the 2D spatial displacement between source and target across the surface, and the resampling that allows native mesh surfaces with disparate numbers of vertices to be brought onto a standard mesh. The ‘‘source-to-target’’ individual-to-atlas deformation map enables rapid conversion of data from the individual to the 74k_f99 atlas mesh. For example, the map of cortical folding (top, Fig. 3F) appears as a slightly smoothed version of the native-mesh folding map (Fig. 3A). In the reverse direction, a target-to-source deformation map specifies the location of each target node in relation to the 3 nodes of the deformed source surface tile it overlies. This allows any data of interest on the Cerebral Cortex October 2012, V 22 N 10 2229 Figure 3. Individual midthickness surface represented using native mesh and atlas mesh tessellations. (A) Native-mesh midthickness surface (top) and expanded to reveal the native mesh with one node highlighted (bottom panel). (B) The native-mesh spherical surface with landmarks, centered on the highlighted node (top panel) and expanded to show the preregistration spherical mesh (bottom panel). (C) The deformed native-mesh sphere, with landmarks precisely aligned to those of with the target spherical. (D) Target (atlas) spherical landmarks. (E) Target spherical nodes overlaid on deformed source spherical mesh. A source-to-target deformation map specifies the barycentric coordinates of each target node (e.g., red node) relative to the nearest tile in the deformed native-mesh sphere (e.g., blue nodes). (F) A resampled (74k_f99) individual subject midthickness surface is generated by positioning each target node based on its barycentric coordinates relative to the native-mesh midthickness surface. atlas to be mapped to the individual’s surface mesh. These maps are ‘‘quasi-invertible,’’ in the sense that transformation of data from one mesh to another, then back to the first preserves the original representation except for degradation relating to differences in the mesh density (e.g., a coarser representation arises if data are mapped from a fine mesh to a coarse mesh and back) and minimal smoothing. Resampling of data can be done using either the source node nearest to the target node or a weighted average of the source nodes defining the tile within which the target node lies. Assessing Geographic Correspondences and Distortions The LVD Landmark-SBR process establishes node-to-node correspondences between the atlas surface and the resampled (74k_f99) version of the individual surface. The fidelity of registration can be assessed by visual inspection of highlighted nodes in relation to nearby geographic features. For example, Figure 4A shows 10 highlighted nodes (black squares) on the individual (top) and atlas (bottom) midthickness surfaces. In general, the node locations are in good correspondence (within ~1--2 mm) relative to nearby gyral and sulcal features. However, imperfect correspondences occur in places. For example, node 8, which is near the tip of the inferior occipital sulcus (IOS) in the atlas surface but is ventral to the IOS tip by about 4 mm in the individual surface. This is attributable to differences in the trajectories of the IOS in these 2 hemispheres. Several types of spatial distortion impact the quality of the registration process, Figure 4B shows a distortion map based on the ratio of surface areas for each standard-mesh atlas tile and the corresponding standardmesh individual tile, each measured on the midthickness surfaces. The distortions are locally quite variable but are moderate in magnitude overall (only 5% of tiles having distortion more than 2-fold of the mean). This overall distortion arises from 2 main sources. One occurs in the mapping of 3D midthickness (or other anatomical) surfaces to a sphere. Local distortions are concentrated in regions associated with intrinsic (Gaussian) curvature, that is, concavities and convexities that are not simple folds or creases (Van Essen and Drury 1997; Van Essen 2005). The sphere versus midthickness distortion maps (Fig. 4C) show broad regional similarities for the individual and atlas surfaces, including compression of the frontal and occipital poles (dark blue regions) and 2230 Macaque Surface-Based Cortical Parcellations d Van Essen et al. expansion of the Sylvian fissure (red regions) that are characteristic of both hemispheres. However, many local bumps and dents are idiosyncratic to each hemisphere, so the maps differ in many local details. Another source of distortion arises from the local deformations that occur during spherical registration. Figure 4D shows an areal distortion map for pre- versus post-registration spheres, with the landmark contours overlaid. The distortions are generally modest in magnitude (99.7% within 2-fold of the average) and are concentrated near landmark terminations. One reason is that the correspondences near the terminations of sulci and gyri are the most difficult to estimate (see Fig. 1). Also, the LVD registration algorithm does not completely eliminate distortions in the region between adjacent landmarks. Landmarks translated over a large distance tend to have residual compression in front of the direction of translation and residual expansion in the region immediately behind. LVD Parameter Specification The LVD algorithm can achieve accurate registration of landmarks (within 1 mm) even when large local deformations are required, as long as an adequate number of stages with appropriate parameters at each stage are used to compensate for the initial misalignment. Numerous free parameters can be adjusted to achieve the desired degree of alignment and to reduce local distortions. A standard set of parameters suitable for registering individual macaque hemispheres is available and can be read in automatically (see SumsDB data set associated with Fig. 2 and the FS-to-F99 tutorial). Any topological defects (crossovers) occurring at any stage of a deformation are reported and can be used as a guide in adjusting parameters, the surfaces generated at intermediate stages are also available. The parameters used successfully for any one registration can be read in and used for other registrations that have similar requirements. Registration quality is impacted by the number and distribution of landmarks, based on the number of local features considered reliable, the shape differences between the source and target surfaces and the desired degree of alignment in different regions. Computation time depends heavily on the number of landmark contours, the number of Figure 5. Preparatory steps for Landmark-SBR of a partial hemisphere reconstruction. (A) Anterior view of 3D anatomical surface of macaque case om43, with landmarks drawn along gyral and sulcal landmarks and along perimeter of prefrontal cortex. (B) Inflated surface generated using a combination of smoothing iterations and expansion from the center of gravity. A cut along the principal sulcus (gap at top) reduced distortions in the cortical flat map (not shown). (C) A partial sphere generated by 1) projecting all nodes in the inflated surface to a fixed radius, 2) reducing distortions relative to the 3D anatomical surface using spherical morphing, 3) applying a spherical compression step that shifts all nodes in the front portion to subtend a solid angle approximately matching that of the target atlas landmarks, and 4) rotating the compressed sphere into approximate alignment with the atlas. (D) Corresponding landmarks on the F99 atlas anatomical surface. (E) Target landmarks on the F99 atlas sphere. Figure 4. Geographic correspondences and areal distortion maps for different stages of spherical registration using LVD Landmark-SBR. (A) Highlighted nodes (black squares) indicated corresponding geographic locations in the individual (top) and atlas (bottom) midthickness surfaces, each shaded by is own map of cortical folding. (B) Ratio between the surface area of each atlas midthickness tile and the corresponding tile in the registered and resampled individual surface. (C) Distortion map between individual sphere and its 3D midthickness surface (top) and between F99 atlas sphere and its 3D midthickness surface (bottom). The individual sphere versus midthickness distortion map is based on the Caret-generated (multiresolution morphing) sphere, whose distortions differ from the FreeSurfer process. (D) Distortion map and between atlas and individual spheres, showing the local distortions needed to align source and target landmarks. processing stages and cycles per stage, and the density of the surface mesh used for spherical registration. For the cases used in this study, it typically took about an hour on a MacBook Pro (2.53 GHz Intel CPU, 8 GB RAM). Surfaces Reconstructed from Contours Some architectonic parcellations had been mapped to surfaces reconstructed from contours of histological sections. The PHT (Paxinos and Franklin 2000) and MMF11 (Markov et al. 2011) atlases were available as complete hemisphere reconstructions based on contours drawn along the estimated trajectory of layer 4 in images of each section loaded into Caret. Landmark-SBR registration from these atlas surfaces to the F99 atlas was similar to that described above for Freesurfer-to-F99, except that a larger number of landmarks were used in order to better compensate for the local irregularities in the reconstructed surfaces. The current version of the PHT00 atlas supplants an older version that had been registered using the LPR algorithm and fewer landmarks. Customized preprocessing was required for cortical surfaces that included only a partial hemisphere (e.g., only prefrontal cortex). Figure 5 illustrates preprocessing steps used to prepare a partial hemisphere of macaque prefrontal cortex (Ferry et al. 2000) for Landmark-SBR. Landmark contours were delineated along geographically corresponding locations in the individual (Fig. 5A) and atlas (Fig. 5D) surfaces. This example included 15 gyral and sulcal landmark contours, of which 9 ran along the perimeter of the partial-hemisphere surface. The individual surface (Fig. 5A) was inflated (Fig. 5B) and mapped to a partial sphere (Fig. 5C) whose landmarks were rotated and scaled to approximately align with the corresponding landmarks on the target sphere (Fig. 5E). Registration of Cortical Flat Maps Many parcellations were registered initially to the 79-0 atlas (Drury et al. 1996) and then to the F99 atlas using an older landmarkconstrained diffeomorphic (fluid-based) algorithm applicable to 2D flat maps—the landmark flat-fluid (LFF) algorithm (Joshi 1997; Van Essen et al. 2001). The input parcellations were represented on computerized surface reconstructions or on manually drawn flat maps (see Table 2). Data Availability Data sets associated with this study are freely available in the SumsDB database (http://sumsdb.wustl.edu/sums/directory.do?id=8286148&dir_ name=MACAQUE_ATLAS_CC11). Associated with each figure in this study is a ‘‘scene’’ file that allows regeneration of all the constituent figure panels. To facilitate cross-platform data migration, most surfacerelated files are in GIFTI format (http://www.nitrc.org/projects/gifti). Data sets can be viewed online using WebCaret without downloading data or software and can be downloaded for offline analysis using Caret or other GIFTI-compliant software. Results Macaque Cortical Surface Area and Variability Table 1 shows data on cortical surface area and its variability in the macaque. The macaque F99 atlas has an average surface Cerebral Cortex October 2012, V 22 N 10 2231 area of 122 cm2 per hemisphere in its native dimensions. After normalization to the average dimensions for adult rhesus macaques (male and female combined; see Materials and Methods), surface area averages 105 cm2 per hemisphere. Sex differences and individual variability in macaque brain dimensions were estimated using scale factors that relate each of 112 individual brains to the 112RM-SL population-average atlas volume (see Materials and Methods). Average brain size in males exceeds that in females by 17% in volume (11% in surface area). Estimated brain volume was not correlated for with age in either males (r2 = 0.009, range = 3--36 years) or females (r2 = 0.05, range = 4--29 years). similarities and differences among parcellations at any given location are difficult to resolve in this figure. When viewing the same data sets in Caret, the assignments at any location can be immediately ascertained by clicking on a surface node of interest and viewing a list of areas in the ‘‘Identify node’’ window (Fig. 6L,M). For example, the white highlighted node in each panel (white arrow) has relatively consistent assignments, listed in Figure 6L: it is identified as MT in 6 parcellations, and the other assignments (areas 19, OAa, and TA) reflect older parcellations in which area MT had not been identified. In contrast, the nearby black highlighted node (black arrow) has a distinct areal identity in all 6 parcellations where an assignment was made (Fig. 6M: area PA, TP, 22, STP, PBc, or TA). In places where parcellations disagree, a few involve differences are only in name, that is, different names used for what is fundamentally the same well-defined area (e.g., areas V1, 17, and OC are synonymous). At the other extreme, areal boundaries in one parcellation can show minimal correlation with those in another parcellation. Intermediate situations are by far the most common, in which areal boundaries are correlated but imperfectly on the atlas map. Lack of correlation can arise from several factors, including 1) differences in the size (2-fold or more), shape, or location of areas in the contributing hemispheres (biological variability); 2) inaccuracies in delineating areal boundaries owing to methodological bias or error; or 3) inaccuracies or bias in the registration process. Another major factor relates to the distinction between ‘‘lumping’’ and ‘‘splitting.’’ For example, one parcellation may identify a single area (e.g., area 3) in a region where another parcellation identifies 2 (e.g., areas 3a and 3b), resulting in areal boundaries that are strongly correlated along part but not all of their trajectories. The following sections discuss several cortical regions where cross-parcellation comparisons benefit from having data registered to the F99 atlas and from having access to individual hemisphere surfaces that contributed to the atlas. These regions include orbital and OMPFC; the STS, especially area MT; and parietooccipital cortex. We also illustrate a composite map based on multiple parcellations spanning most of cerebral neocortex. Macaque Parcellations Table 2 provides information on 15 published cortical parcellations mapped to the F99 atlas, including 5 newly incorporated or revised maps, plus 9 previously registered parcellations (Van Essen 2004). The parcellations are based on architectonic studies, retinotopy using neurophysiology or functional magnetic resonance imaging (fMRI), connectivity, or a combination of the above. Figure 6 displays all 15 parcellations on the very inflated atlas surface. Six parcellations (Fig. 6A--F) each encompass most or all of cerebral neocortex and transitional cortex. The others cover only a minority of cortex; panels H and I each combine 3 nonoverlapping parcellations. For the 6 pan-hemispheric parcellations, the number of identified areas ranges from 26 (BB47) to 161 (PHT00) (Table 2, column 4). Specific Table 1 Macaque neocortical surface area Macaque F99 atlas Surface area (cm2) Left Right Average Individuala Normalized Sex Scaling factorsb 125 119 122 107 103 105 Male (n 5 52) Female (n 5 30) Combined 1.04 ± 0.12 (0.77--1.37) 1.22 ± 0.13 (1.03--1.65) 1.11 a Surface areas of the original mesh (~30 000 nodes each for the left and right hemisphere) were about 3% larger than the standard-mesh values (because resampling slightly smooths the surface) but were not a discernibly better fit to cortical shape when overlaid on the MR volume. b Average 3D scaling factors for individuals relative to the 112RM-SL average MR, based on the product of the x, y, and z scaling factors. Orbital and Medial Prefrontal Cortex The F99 atlas includes 8 parcellations having cortical areas in the OMPFC region. Figure 7 compares 2 of the most widely Table 2 Macaque cortical parcellations registered to the F99 atlas (in the order illustrated in Fig. 6) Study Abbreviation Lobe(s) Number of areas Number of cases Modality Input (source) surface Registration algorithm Lewis and Van Essen (2000) Felleman and Van Essen (1991) Paxinos and Franklin (2000) Markov et al. (2011) Brodmann (1905) von Bonin and Bailey (1947) Ungerleider and Desimone (1986) Seltzer and Pandya (1978, 1980, 1986) Galletti et al. (1999) Preuss and Goldman-Rakic (1991) Preuss and Goldman-Rakic (1991) Lyon and Kaas (2002) Bayliss et al. (1987) Kolster et al. (2009) Ferry et al. (2000) LV00 FV91 PHT00 MMF11 B05 BB47 UD86 SP78-80-86 GFG99 PGR91a PGR91b LK02 BRL87 KMA09 FOA00 All All All All All All O, T, P P, T P P F O T O, P, F F 88 80 161 78 28 26 17 13 1 5 38 11 9 12 20 5 1 1 1 1 1 1 1 1 1 1 1 1 4 5 Architectonic Multiple Architectonic Architectonic Architectonic Architectonic Retinotopic Physiology Architectonic Retinotopic physiology Architectonic Architectonic Connectivity Architectonic Retinotopic fMRI Architectonic Caret flat map Manual flat map 3D reconstruction 3D surface Manual flat map Manual flat map Manual flat map Manual flat map 3D partial hemisphere Manual flat map Manual flat map Manual flat map Manual flat map FreeSurfer surface 3D partial hemisphere LFF to F99 LFF LVD (30) LVD (24) LFF via 79-0 LFF via 79-0 LFF via 79-0 LFF LVD (9) LFF via 79-0 LFF via 79-0 LFF via 79-0 LFF via 79-0 LVD (16) LVD-ph (15) Note: LVD; the number of landmarks is indicated in parentheses LVD-ph: partial-hemisphere, registered via LVD LFF (Joshi 1997; Van Essen et al. 2001) via 79-0 signifies initial registration to the Case 79-0 atlas (Drury et al. 1996), followed by registration to F99. 2232 Macaque Surface-Based Cortical Parcellations d Van Essen et al. Figure 6. Cortical areas from 15 parcellations mapped to the F99 atlas surface. (A--K) Areas for one or more nonoverlapping parcellations, presented in the same sequence as in Table 1. (L, M) Areas identified for the white (panel L) and black (panel M) nodes highlighted in panels A--J. Each area is identified using a standard format: \area-name[_\parcellation[. used, the FOA00 parcellation of Ferry et al. (2000) and the PHT00 parcellation of Paxinos and Franklin (2000). For the FOA00 parcellation, OMPFC areas were delineated histologically in 5 hemispheres using criteria based on cytoarchitecture, myeloarchitecture, and 3 types of chemoarchitecture (Carmichael and Price 1994). Areas were mapped to surface reconstructions of prefrontal cortex in each hemisphere (Fig. 7A) and then registered to the F99 atlas (Fig. 7B; see Materials and Methods). A probabilistic map (Fig. 7C, n = 5) shows that intersubject alignment was reasonably good for all 3 Cerebral Cortex October 2012, V 22 N 10 2233 Figure 7. Comparing OMPFC areas in the FOA00 and PHT00 parcellations. (A) Ventral view of orbital cortex an individual subject (case om43 of Ferry et al. 2000). (B) Case om43 areas registered to the F99 atlas. (C) Probabilistic areas 13a, Iai, and 11l (n 5 5 hemispheres). (D) Composite map of FOA00 areas on ventral (top) and medial (bottom) views. (E) FOA00 composite borders overlaid on PHT00 areas registered to F99 atlas. (F) PHT00 areas on ventral and medial views of the reconstructed PHT00 atlas surface (Paxinos and Franklin 2000). illustrated areas (13a, Iai, and 11l). Composite boundaries drawn using the complete probabilistic map show the most likely location and extent of each area on ventral and medial atlas views (Fig. 7D). The PHT00 parcellation was based on cytoarchitecture and chemoarchitecture (cholinesterase) of a single atlas hemisphere reconstructed from section contours (Fig. 7F; see Materials and Methods). Side-by-side comparison of the FOA00 and PHT00 parcellations (Fig. 7D,F) shows many similarities but also many differences. Figure 7E allows a more direct comparison by showing the PHT00 areas registered to the F99 atlas, with the FOA00 borders overlaid. A direct comparison reveals that some areas are reasonably well aligned (e.g., PHT00 13M, 13L, and 11L vs. FOA00 13m, 13l, and 11l), whereas other areas (e.g., 25) are in substantial disagreement; most areas lie in between (some overlap but extensive nonoverlap). Inspection of both parcellations on the original hemispheres (Fig. 7A,F) indicate that the mismatches generally cannot be attributed to inaccurate registration to the atlas for either parcellation. Area MT and the Posterior Bank of the STS Area MT is one of the most intensively studied extrastriate area in the macaque, yet its location and extent vary markedly in the 7 parcellations mapped to the F99 atlas. Macaque MT was originally defined using 4 independent criteria that revealed 2234 Macaque Surface-Based Cortical Parcellations d Van Essen et al. coextensive boundaries: it is heavily myelinated, contains a complete visuotopic map, has a high incidence of directionselective cells, and receives a strong projection from area V1 originating in layer 4B (Van Essen et al. 1981; Maunsell and Van Essen 1983). Using these criteria (especially myeloarchitecture), MT was mapped to the medial half of the posterior bank of the STS, in a region typically ~3--4 mm wide, ~1-cm long, and with an average surface area of 33 mm2 (Van Essen et al. 1981; Maunsell and Van Essen 1987). In Figure 8A, the white squares outline the posterior bank of the STS, and red shows the extent of myeloarchitectonic MT in the FV91 parcellation, in good agreement with the aforementioned location and extent. (It is more elongated in the figure than in the actual atlas midthickness surface because the inflation process compresses the surface anisotropically in this region.) Figure 8B shows MT and the other FV91 areas displayed on the very inflated atlas surface, with white squares again outlining the posterior bank of the STS. Figure 8C--H shows the diversity in position, shape, and size of area MT (red) in 6 other studies mapped to the atlas, presented in order of decreasing similarity to MT of FV91 (black contour in each panel). The LV00 (Lewis and Van Essen) MT map (Fig. 8C), based mainly on myeloarchitecture, is centered on the same location but is narrower and longer than FV91 MT (black contour). The UD86 (Ungerleider and Desimone 1986) MT map (Fig. 8D) is similar in size and shape to FV91 MT but is Figure 8. Comparison of macaque area MT size and location across multiple studies. (A) Area MT (red) from FV91 (Felleman and Van Essen 1991), based on myeloarchitectonic MT on inflated F99 atlas, with posterior bank of the STS highlighted (white squares). MT is several-fold longer than wide when visualized in 3D space (see Fig. 5, Van Essen et al. 1981) or on the midthickness surface, but it appears more highly elongated in Figure 8 owing to anisotropies that occur during surface inflation. (B) FV91 visual areas with black contour outlining MT on the very inflated atlas surface. (C--H) Cortical areas from 6 other parcellation schemes, with MT as read and FV91 MT outlined (black contour). (C) LV00 5 Lewis and Van Essen (2000). (D) UD86 5 Ungerleider and Desimone (1986). (E) LK02 5 Lyon and Kaas (2002). (F) PHT00—Paxinos and Franklin (2000). (G) MMF11 5 Markov et al. (2011). (H) Probabilistic area MT from KMA09 5 Kolster et al. (2009, n 5 4). (I) STS parcellation of KMA09. (J, K) KMA09 parcellations from the left and right hemispheres of case CH (panel K) and case TO (panel J), with the composite borders of MT and MTp outlined. MTp was identified as a distinct visuotopic map labeled as ‘‘[2]’’ by Kolster et al. (2009, their Figs 2, 4, 6, and 7) and designated as MTp by Kolster et al. (2010, their Fig. 1) in a reinterpretation of the same data set. oriented differently so that the 2 maps overlap only about 50%. The LK02 (Lyon and Kaas 2002) MT map (Fig. 8E) is smaller and lies mostly lateral to FV91 MT. The PHT00 (Paxinos and Franklin 2000) (Fig. 8F) and MMF11 (Markov et al. 2011) (Fig. 8G) maps are both based mainly on cytoarchitecture and are more than twice as large; they extend to the dorsolateral lip of the STS and also into the anterior bank of the STS. Kolster et al. (2009) mapped the central visual field representation of MT and other visual areas using retinotopic fMRI in 4 hemispheres that were mapped to the F99 atlas. A probabilistic map of their MT (Fig. 8H) overlaps partially with FV91 MT but is centered in the lateral half of the STS. The missing peripheral field representation should be more dorsal (up on the map) based on the observed retinotopy. (Their visual stimuli extended to 12 degrees, but their areal boundaries covered only the central ~6 degrees—cf. their Fig. 2) In the composite areal parcellation (Fig. 8I), FV91 MT overlaps more extensively with KMA09 areas MTp (pink), MSTv (blue), and FST (brown) than with their MT. In maps of the individual inflated hemispheres (Fig. 8J,K), FV91 MT overlaps substantially with KMA09 MT in only one hemisphere (TO right) and generally shows greatest overlap with their MSTv. Kolster et al. (2009, their Fig. 6, case CH right) noted a good correspondence between their MT and that of Ungerleider and Desimone (1986) at one slice level, consistent with our maps (Fig. 8D,J). However, the discrepancy for the population-average MT as a whole is puzzling. One possibility is that the KMA09 retinotopic data might be consistent with a revised parcellation in which MT would include cortex they assigned to MTp and perhaps part of MSTv. Comparisons across parcellations for the other extrastriate visual areas displayed in Figure 8 reveals numerous additional differences and discrepancies. For some parcellations, even the early extrastriate areas V2 (yellow), V3d and V3v (green), and V4 (blue) differ substantially in terms of size and location. These differences are likely to reflect a combination of methodological and biological factors (see Discussion). Parietooccipital Areas Figure 9 shows parcellations of parietooccipital cortex from 6 studies, with the LV00 borders shown as an overlay on all Cerebral Cortex October 2012, V 22 N 10 2235 Figure 9. Comparison of macaque parietal regions from 6 studies displayed on the very inflated F99 atlas from medial (top) and dorsal (bottom) views, all with LV00 borders overlaid. (A) Cortical areas in the macaque parietooccipital cortex from the LV00 architectonic parcellations. White arrow points to the gap between area V1 and the corpus callosum. (B) FV91 areas. Arrow points to FV91 prostriate area (blue). (C) PHT00 parcellation. Arrow points to PHT00 prostriate area (PS, blue). (D) Area V6 (Galletti et al., 1999) with LV00 borders overlaid. (E) Parietal areas from Preuss and Goldman-Rakic (1991) and Lyon and Kaas (2002), with LV00 borders overlaid. panels to facilitate comparisons. Three parcellations—LV00 (Fig. 8A), FV91 (Fig. 9B), and PHT00 (Fig. 9C)—each span most of this region. Several observations warrant comment, especially in relation to interspecies comparisons (Van Essen et al. 2011). Macaque V1 extends nearly to the anterior tip of the calcarine sulcus, where it adjoins a narrow strip of prostriate cortex (white arrows in Fig. 8A--C) that separates it from the subiculum and the termination of cortex at the corpus callosum. In this anterior region, dorsal and ventral halves of area V2 (yellow) are separated by the prostriate area (blue) and also by area 29 (in the FV91 parcellation). (See also Van Essen et al. 1982 for supporting evidence based on interhemispheric connections.) In contrast, human V1 (area 17) terminates much farther posterior (near the junction of the POS with the CaS), and there is a wider expanse of cortex between it and the corpus callosum (Van Essen et al. 2011). Retinotopically mapped area V6 of GFG99 (Galletti et al. 1999; pink in Fig. 8D) includes a complete map that emphasizes the visual periphery. It adjoins the peripheral visual field representation of area V2. (Indeed, it overlaps partially with V2 in all 3 illustrated parcellations.) Its retinotopic organization is similar to that of human hV6. However, hV6 is widely separated from V2 (Pitzalis et al. 2006; Van Essen et al. 2011), so it is unclear whether they are genuinely homologous to one another. In the intraparietal sulcus (top-bottom row, Fig. 9A--C), the 5 parcellations mapped to the atlas include areas that are generally elongated parallel to the sulcus. The LV00 parcellation of the lateral IPS includes LIPv (dark orange), similar to PHT00 area POaI and PGR91 area LIP, and LIPd (light orange), similar to PHT00 area POaE and PGR91 area 7am. In the fundus and medial IPS, the LV00 parcellation includes VIPl (light green) and VIPm (dark green), which are nearly coextensive with PGR91 2236 Macaque Surface-Based Cortical Parcellations d Van Essen et al. area VIP and overlap partially with FV91 area VIP and with PHT00 areas DIP and PEa. Size Range and Total Number of Macaque Cortical Areas The parcellations discussed in this study span a wide range in terms of their average size and their total number within the region covered (Table 2). The PHT00 parcellation is the most fine-grained and includes a total of 161 parcels. However, this is likely to be a substantial overestimate because some of the entries represent transitional regions between 2 well-defined areas (e.g., area 2/1 in between area 2 and area 1). In addition, some of the smaller areas have not been validated in other architectonic studies or using other methods. We generated a composite pan-hemispheric architectonic map (Fig. 10) that entailed merging of 3 independent parcellations (LV00, FOA00, and PHT00). The rationale is that each of these parcellations is likely to be more accurate than the others in some regions but is incomplete or arguably less accurate in other regions. In any one region, only one parcellation was used. The composite map includes 129 distinct areas of neocortex and transitional cortex. These are primarily from the LV00 parcellation in dorsal, lateral, and posterior cortex, primarily from the FOA00 parcellation in orbital cortex and from all 3 parcellations in medial cortex. This entails subjective judgment to select the parcellation for any given region, and the composite should not be considered ‘‘ground truth’’ or a ‘‘gold standard’’ (see Discussion). These 129 parcels have an average area of 74 mm2 and encompass total surface area of 96 cm2, which is 93% of total right hemisphere cortical surface area (103 cm2). As indicated in the area-size histogram (Fig. 10E), these areas vary greatly in size. The total range exceeds 100-fold range between the 2 largest areas (V1, V2) and 9 areas that are all smaller than 10 mm2. A large majority of areas (96) are within an 8-fold range in the middle (20--160 mm2). If the remaining 7% of Figure 10. (A) Map of 129 areas from the composite LV-FOA-PHT parcellation shown in lateral, medial, dorsal, and ventral views. Unassigned regions (gray with asterisks, 7% of total surface area) either lacked a well-defined areal assignment in any of the 3 contributing parcellations or were assigned to a cortical area in one parcellation (e.g., V2v of PHT00) that was considered less accurate than the corresponding area of a different parcellation (e.g., V2v of LV00). (B) Histogram of areal sizes in the composite parcellation, calculated using the midthickness surface for the right hemisphere normalized to average rhesus macaque brain size (cf. Table 1). unassigned cortex (gray regions with asterisks in Fig. 10A) contains areas similar in size to the average, then macaque cortex (neocortex plus transitional) contains an estimated 140 total cortical areas based on this composite parcellation. Discussion In this study, we used a new method of SBR to expand and improve upon the set of cortical parcellations registered to the macaque F99 atlas. Our analysis suggests that macaque cortex contains about 130--140 areas that span more than 2 orders of magnitude in average surface area. Comparisons across different parcellations reveal considerable diversity in the estimated location and extent of many areas, even ones such as MT that have been intensively studied for decades. Four general issues warrant additional discussion: 1) surface-based atlases and registration; 2) evaluating and reconciling cortical parcellations; and 3) estimating the total number of cortical areas in the macaque; and 4) interspecies comparisons. Registration Algorithms and Atlases A primary objective of registering data to a cortical atlas is to maximize intersubject alignment of cortical parcellations and functional maps. The Landmark-SBR algorithms used in the present study constrain registration using structural (geographic) features as a surrogate for underlying functional organization. We evaluated the fidelity of registration based on the alignment of geographic features themselves and also based on the consistency of areal alignment in probabilistic maps (Fig. 7C and also Fig. 4 in Van Essen et al. 2001). The consistency of folding patterns in the macaque enables accurate delineation of corresponding geographic landmarks in individual and atlas surfaces. For the Landmark-SBR approach described here, geographic correspondences in the regions between landmarks are typically accurate within about 2 mm based on visual inspection (cf. Fig. 4A). Because areal boundaries have relatively consistent locations relative to folds in the macaque, Landmark-SBR also achieves reasonably good intersubject alignment of cortical areas, as shown for probabilistic architectonic areas (Ferry et al. 2000; Lewis and Van Essen 2000) and retinotopic visual areas (Kolster et al. 2009). The residual variability present in these probabilistic maps arises from multiple factors, including: 1) inaccuracies or different criteria for identifying areal boundaries in individual subjects, 2) interindividual variability in areal size, 3) variability in the location of areal boundaries relative to the folds used as landmarks, and 4) local distortions associated with the Landmark-SBR method. While it is difficult to quantify the relative impact of each factor for the macaque data sets analyzed here, distortions and biases attributable to the registration process are likely to be modest relative to the other factors. Inaccuracies in mapping of areal borders and/or criterion differences in identifying areal boundaries may be the dominant factors in accounting for misalignment of many areas (see below). Some parcellations were mapped to the F99 atlas using older data sets that had initially been mapped using older algorithms (the LFF method for flat maps) and in some cases an older atlas (Case 79-0) as an initial target. These algorithmic differences might impact comparisons between parcellations to a limited degree, but they are unlikely to be a major confound. Only modest deformations are needed to map from one macaque surface to another, and the density of landmarks used to constrain the registration was adequate to prevent large local deformations. In addition, direct visual comparison with results as displayed on the primary source generally reveal only modest bias or distortion relative to the other uncertainties in the parcellation or its representation on the cortical surface. Such comparisons are clearest when the input parcellation is available on explicit surface reconstructions (e.g., Figs 4, 5, 7, and 8). However, good correspondence is generally evident when comparisons are made in relation to section drawings or schematic brain drawings in the original published study in order to assess the fidelity of mapping to the atlas. It would be of interest to compare the LVD algorithm introduced here to other Landmark-SBR algorithms derived using different principles and approaches (e.g., Glaunes et al. 2004; Joshi et al. 2007; Pantazis et al. 2010) and also with EnergySBR methods (Fischl et al. 1999; Yeo et al. 2010) if adapted to Cerebral Cortex October 2012, V 22 N 10 2237 register macaque surfaces. We were unable to make direct comparisons using these other algorithms as our data sets. However, as noted above, the impact of the choice of algorithm depends greatly on the magnitude of deformations required and the density of landmarks or other features that constrain the registration. These issues are much more important when considering registration of human cortical surfaces, owing to the greater complexity and variability of human cortical convolutions (Van Essen et al. 2011) and for interspecies registration (Orban et al. 2004; Van Essen and Dierker 2007). Several other freely available macaque atlases besides the F99 atlas have been derived from MR scans and can be viewed as volumetric data sets in stereotaxic coordinates. Population average atlases include the 112RM-SL atlas for the rhesus macaque (McLaren et al. 2009), the Montreal Neurological Institute (MNI) rhesus plus cynomolgus macaque atlas (Frey et al. 2011), and the F6 cynomolgus atlas (Vincent et al. 2007), and the M. nemestrina atlas (Black et al. 2001). Single-subject atlases include the macaque D99 atlas (Saleem and Logothetis 2007). As with human stereotaxic atlases, methods for accurate interatlas registration are desirable, so that data generated or represented on one atlas can be easily transferred to any other. The F99 atlas considered here includes midthickness surfaces that have already been registered to D99, 112RM-SL, and F6 space. Registration of F99 to the MNI and D99 (112RM-SL) atlases will be straightforward to execute as well. Surface maps of cortical parcellations illustrated here can be readily converted to volumetric format in any of these alternate stereotaxic spaces using existing surface-to-volume capabilities in Caret. Reconciling Parcellations A fundamental challenge is to determine whether spatially overlapping parcels as defined in different studies are coextensive and therefore represent fundamentally the same underlying cortical domain, whether or not they were delineated by the same or different methods. Another challenge is to determine which parcellation most accurately reflects the underlying neurobiological organization (i.e., the ground truth). Atlas-based comparisons of parcellations as illustrated here do not resolve these issues, but they do provide useful insights. Area MT is an informative example, as it has been charted in many studies using multiple methods. The discrepancies among architectonically based delineations of MT likely reflect criterion differences between, for example, the cytoarchitectonic boundaries of PHT00 MT (Paxinos and Franklin 2000) and the more restricted myeloarchitectonic boundaries of LV00 MT (Lewis and Van Essen 2000) and FV91 MT (Felleman and Van Essen 1991). It is difficult to reconcile the discrepancies between maps of MT using retinotopic-fMRI (Kolster et al. 2009) versus any of the architectonic maps. Retinotopic fMRI involves complex analysis procedures applied to noisy experimental data; inaccurate estimation of areal boundaries using this modality might arise from a variety of factors, including incomplete mapping of the visual field and also inadequate registration of blood oxygen level--dependent EPI data sets to the anatomical MR volume. Achieving consistency across methods remains a highly desirable but an often elusive objective in the realm of cortical parcellation. We generated a composite pan-hemispheric parcellation in order to deal with limitations in available single-study 2238 Macaque Surface-Based Cortical Parcellations d Van Essen et al. parcellations—either in the amount of cortex covered or in the reliability of the parcellation in certain regions. For example, in OMPFC, the FOA00 parcellation has multiple advantages over other parcellations of this region. It is based on 5 independent architectonic criteria, it is a population average (n = 5) and thus less susceptible to the impact of individual variability, and connectivity data provide independent support for many of these areal assignments (Carmichael and Price 1996; Ferry et al. 2000). For other cortical regions, it is more difficult to assess which parcellation best reflects ground truth. Using a combination of the LV00 and PHT00, parcellations for non-OMPFC regions has advantages over either one alone but many alternative hybrids could plausibly be considered. One issue that arises is that the boundaries between separate parcellations rarely abut precisely. This leads to occasional gaps (unassigned cortex) and also modest overlaps that were adjudicated by trimming one or both areas. Patches, Gradients, Modules, and Parcellations Our estimate of ~140 cortical areas per hemisphere in the macaque provides one useful value for a fundamental feature of cortical organization. However, it is subject to the well-known tension between lumping and splitting of cortical areas. Some regions considered a single area in this composite parcellation arguably might be split into finer-grained subdivisions that warrant being considered separate areas. Alternatively, some fine-grained parcels arguably might be more appropriately lumped together in order to constitute neurobiologically welldefined areas. On what grounds and by what criteria, should any given transition in properties be considered an areal boundary versus heterogeneity within an area? There is no simple answer to such questions. An instructive example involves inferotemporal cortex, which is implicated in high-level processing of complex objects, including faces. IT neurons sharing similar functional properties tend to be clustered. Most notably, multiple face-selective patches have been identified by fMRI and neurophysiology (Tsao et al. 2006), and they are preferentially interconnected (Moeller et al. 2008). Should each face patch be considered a separate cortical area because it is distinct from neighboring cortex based on function and connectivity? A plausible alternative is to consider each face patch as a module within a larger area, whose other components might consist of 1) a additional modules that are comparably distinct but have different specializations (e.g., for other object categories) or 2) a heterogeneous region with fluctuations and gradients in functional properties that lack crisp modularity. For these and other reasons, the debate between cortical ‘‘lumpers’’ and ‘‘splitters’’ is unlikely to be quickly resolved. The concept of cortical areas remains a powerful organizing principle, but properties of modularity and internal gradients are valuable and complementary concepts as well. Progress on these issues will benefit from neuroimaging methods that enable systematic analysis across large cortical expanses and objective analysis of cortical properties and their spatial gradients (see Glasser and Van Essen 2011; Van Essen et al. 2011) Interspecies Comparisons A better understanding of macaque cortical parcellations will provide valuable comparative data that can aid in understanding human cortical parcellation. A companion study (Van Essen et al. 2011) also uses a surface-based analysis strategy to map published human cortical parcellations to an improved surface-based atlas. Progressively more accurate parcellations that are mapped to atlas surfaces in both species will facilitate evaluations of presumed homologies between species. This will also aid in quantitative assessments of differential cortical expansion during evolution (Orban et al. 2004; Van Essen and Dierker 2007). Interspecies comparisons will also be aided by analyses of features that tend to be evolutionarily conserved, such as myelin content (Glasser and Van Essen 2011) and cortical connectivity patterns. Funding National Institutes of Mental Health grant RO1 MH60974. Notes We thank S. Danker for assistance in manuscript preparation, E. Reid for assistance with the data analysis, and Drs J. L. Price, H. Kolster, G. Orban, H. Kennedy, N. Markov, and C. Galletti for providing data sets used to map cortical parcellations to the atlas. Conflict of Interest : None declared. References Bayliss GC, Rolls ET, Leonard CM. 1987. Functional subdivisions of the temporal lobe neocortex. J Neuroscience. 7:330--342. Black KJ, Koller JM, Snyder AZ, Perlmutter JS. 2001. Template images for nonhuman primate neuroimaging: 2. Macaque. Neuroimage. 14:744--748. Brodmann K. 1905. Beitraege zur histologischen Lokalisation der Grosshirnrinde: dritte Mitteilung: Die Rindenfelder der niederen Affen Journal fuer Psychologie und Neurologie. Neuroimage. 4:5/ 6:177--226. Carmichael ST, Price JL. 1994. Architectonic subdivision of the orbital and medial prefrontal cortex in the macaque monkey. J Comp Neurol. 346:366--402. Carmichael ST, Price JL. 1996. Connectional networks within the orbital and medial prefrontal cortex of macaque monkeys. J Comp Neurol. 371:179--207. Drury HA, Van Essen DC, Anderson CH, Lee CW, Coogan TA, Lewis JW. 1996. Computerized mappings of the cerebral cortex: a multiresolution flattening method and a surface-based coordinate system. J Cogn Neurosci. 8:1--28. Felleman DJ, Van Essen DC. 1991. Distributed hierarchical processing in the primate cerebral cortex. Cereb Cortex. 1:1--47. Ferry AT, Ongur D, An X, Price JL. 2000. Prefrontal cortical projections to the striatum in macaque monkeys: evidence for an organization related to prefrontal networks. J Comp Neurol. 425:447--470. Fischl B, Sereno MI, Tootell RB, Dale AM. 1999. High-resolution intersubject averaging and a coordinate system for the cortical surface. Hum Brain Mapp. 8:272--284. Frey S, Pandya DN, Chakravarty MM, Bailey L, Petrides M, Collins DL. 2011. An MRI based average macaque monkey stereotaxic atlas and space. MNI monkey space. Neuroimage. 55:1435--1442. Galletti C, Fattori P, Gamberini M, Kutz DF. 1999. The cortical visual area V6: brain location and visual topography. Eur J Neurosci. 11:3922--3936. Glasser M, Van Essen D. 2011. Mapping human cortical areas in vivo based on myelin content as revealed by T1 and T2-weighted MRI. J Neurosci. 31:11597--11616. Glaunes J, Vaillant M, Miller MI. 2004. Landmark matching via large deformation diffeomorphisms on the sphere. J Math Imaging Vis. 20:179--200. Joshi AA, Shattuck DW, Thompson PM, Leahy RM. 2007. Surfaceconstrained volumetric brain registration using harmonic mappings. IEEE Trans Med Imaging. 26:1657--1669. Joshi SC. 1997. Large deformation landmark based diffeomorphic for image matching [PhD thesis] [St. Louis (MO)]: Sever Institute, Washington University. Kaas JH. 2005. The future of mapping sensory cortex in primates: three of many remaining issues. Philos Trans R Soc Lond B Biol Sci. 360:653--664. Kolster H, Mandeville JB, Arsenault JT, Ekstrom LB, Wald LL, Vanduffel W. 2009. Visual field map clusters in macaque extrastriate visual cortex. J Neurosci. 29:7031--7039. Kolster H, Peeters R, Orban GA. 2010. The retinotopic organization of the human middle temporal area MT/V5 and its cortical neighbors. J Neurosci. 30:9801--9820. Lewis JW, Van Essen DC. 2000. Mapping of architectonic subdivisions in the macaque monkey, with emphasis on parieto-occipital cortex. J Comp Neurol. 428:79--111. Lyon DC, Kaas JH. 2002. Connectional evidence for dorsal and ventral V3, and other extrastriate areas in the prosimian primate, Galago garnetti. Brain Behav Evol. 59:114--129. Markov NT, Misery P, Falchier A, Lamy C, Vezolia J, Quilodran R, Giroud P, Gariel MA, Ercsey-Ravasz MM, Pilaz LJ, et al. 2011. Computational methods for reconstructing and unfolding the cerebral cortex. Cereb Cortex. 21:1254--1272. Maunsell JH, Van Essen DC. 1983. Functional properties of neurons in middle temporal visual area of the macaque monkey. I. Selectivity for stimulus direction, speed, and orientation. J Neurophysiol. 49:1127--1147. Maunsell JH, Van Essen DC. 1987. Topographic organization of the middle temporal visual area in the macaque monkey: representational biases and the relationship to callosal connections and myeloarchitectonic boundaries. J Comp Neurol. 266:535--555. McLaren DG, Kosmatka KJ, Oakes TR, Kroenke CD, Kohama SG, Matochik JA, Ingram DK, Johnson SC. 2009. A population-average MRI-based atlas collection of the rhesus macaque. Neuroimage. 45:52--59. Moeller S, Freiwald WA, Tsao DY. 2008. Patches with links: a unified system for processing faces in the macaque temporal lobe. Science. 320:1355--1359. Orban GA, Van Essen D, Vanduffel W. 2004. Comparative mapping of higher visual areas in monkeys and humans. Trends Cogn Sci. 8:315--324. Pantazis D, Joshi A, Jiang J, Shattuck DW, Bernstein LE, Damasio H, Leahy RM. 2010. Comparison of landmark-based and automatic methods for cortical surface registration. Neuroimage. 49:2479--2493. Paxinos G, Franklin KBJ. 2000. The mouse brain in stereotaxic coordinates. San Diego, CA: Academic Press. Pitzalis S, Galletti C, Huang RS, Patria F, Committeri G, Galati G, Fattori P, Sereno MI. 2006. Wide-field retinotopy defines human cortical visual area v6. J Neurosci. 26:7962--7973. Preuss TM, Goldman-Rakic PS. 1991. Architectonics of the parietal and temporal association cortex in the strepsirhine primate Galago compared to the anthropoid primate Macaca. J Comp Neurol. 310:475--506. Saad ZS, Reynolds RC, Argall RC, Japee S, Cox RW. 2004. SUMA: an interface for surface-based intra- and inter-subject analysis with AFNI. Proceedings of the 2004 International Symposium on Biomedical Imaging, Arlington, VA, IEEE. p. 1510--1513. Saleem KS, Logothetis NK. 2007. A combined MRI and histology atlas of the rhesus monkey brain. San Diego, CA: Academic Press. Seltzer B, Pandya DN. 1978. Afferent cortical connections and architectonics of the superior temporal sulcus and surrounding cortex in the rhesus monkey. Brain Res. 149:1--24. Seltzer B, Pandya DN. 1980. Converging visual and somatic sensory cortical input to the intraparietal sulcus of the rhesus monkey. Brain Res. 192:339--351. Seltzer B, Pandya DN. 1986. Posterior parietal projections to the intraparietal sulcus of the rhesus monkey. Exp Brain Res. 62:459--469. Tsao DY, Freiwald WA, Tootell RB, Livingstone MS. 2006. A cortical region consisting entirely of face-selective cells. Science. 311:670--674. Cerebral Cortex October 2012, V 22 N 10 2239 Ungerleider LG, Desimone R. 1986. Cortical connections of visual area MT in the macaque. J Comp Neurol. 248:190--222. Van Essen D, Newsome WT, Bixby JL. 1982. The middle temporal visual area in the macaque: myeloarchitecture, connections, functional properties and topographic organization. J Neurosci. 2:265--283. Van Essen D, Newsome WT, Maunsell JH. 1984. The visual field representation in striate cortex of the macaque monkey: asymmetries, anisotropies and individual variability. Vision Res. 24:420--448. Van Essen DC. 2002a. Surface-based atlases of cerebellar cortex in the human, macaque, and mouse. Ann N Y Acad Sci. 978:468--479. Van Essen DC. 2002b. Windows on the brain: the emerging role of atlases and databases in neuroscience. Curr Opini Neurobiol. 12:574--579. Van Essen DC. 2004. Surface-based approaches to spatial localization and registration in primate cerebral cortex. Neuroimage. 23(Suppl 1):S97--S107. Van Essen DC. 2005. A population-average, landmark- and surfacebased. PALS. Atlas of human cerebral cortex. Neuroimage. 28:635--662. Van Essen DC, Dierker DL. 2007. Surface-based and probabilistic atlases of primate cerebral cortex. Neuron. 56:209--225. Van Essen DC, Drury HA. 1997. Structural and functional analyses of human cerebral cortex using a surface-based atlas. J Neurosci. 17:7079--7102. Van Essen DC, Drury HA, Joshi S, Miller MI. 1998. Functional and structural mapping of human cerebral cortex: solutions are in the surfaces. Proc Natl Acad Sci U S A. 95:788--795. 2240 Macaque Surface-Based Cortical Parcellations d Van Essen et al. Van Essen DC, Glasser MF, Dierker D, Harwell J, Coalson T. 2011. Parcellations and hemispheric asymmetries of human cerebral cortex analyzed on surface-based atlases. Cerebral Cortex. (in press). Van Essen DC, Harwell J, Hanlon D, Dickson J. 2005. Surface-based atlases and a database of cortical structure and function. In: Koslow SH, Subramaniam S, editors. Databasing the brain: from data to knowledge. Neuroinformatics. Hoboken, NJ: John Wiley & Sons. p. 369--388. Van Essen DC, Lewis JW, Drury HA, Hadjikhani N, Tootell RB, Bakircioglu M, Miller MI. 2001. Mapping visual cortex in monkeys and humans using surface-based atlases. Vision Res. 41:1359--1378. Van Essen DC, Maunsell JH. 1980. Two-dimensional maps of the cerebral cortex. J Comp Neurol. 191:255--281. Van Essen DC, Maunsell JH, Bixby JL. 1981. The middle temporal visual area in the macaque: myeloarchitecture, connections, functional properties and topographic organization. J Comp Neurol. 199:293--326. Vincent JL, Patel GH, Fox MD, Snyder AZ, Baker JT, Van Essen DC, Zempel JM, Snyder LH, Corbetta M, Raichle ME. 2007. Intrinsic functional architecture in the anaesthetized monkey brain. Nature. 447:83--86. von Bonin G, Bailey P. 1947. The neocortex of Macaca Mulatta. Urbana (IL): University of Illinois. Yeo BT, Sabuncu MR, Vercauteren T, Ayache N, Fischl B, Golland P. 2010. Spherical Demons: fast diffeomorphic landmark-free surface registration. IEEE Trans Med Imaging. 29:650--668.