* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download CDC Presentation
Survey
Document related concepts
Transcript
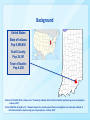
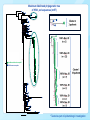
HIV-1 and HCV molecular epidemiology of a large community outbreak of HIV-1 infection linked to injection drug use of oxymorphone - Indiana, 2015 R.R. Galang, J. Gentry, P.J. Peters, S.J. Blosser, E.L. Chapman, C. Conrad, J.M. Duwve, L. Ganova-Raeva, W. Heneine, D. Hillman, H. Jia, L. Liu, W. Luo, J. Lovchik, S. Masciotra, S.M. Owen, A. Perez, P. Peyrani, P. Pontones, S. Ramachandran, J.C. Roseberry, M. Sandoval, A. Shankar, H. Thai, G. Xia, Y. Khudyakov, W.M. Switzer Romeo R. Galang MD MPH Epidemic Intelligence Service Officer International AIDS Society Conference Vancouver, Canada July 20, 2015 National Center for HIV/AIDS, Viral Hepatitis, STD, and TB Prevention Division of HIV/AIDS Prevention Disclosures Neither I nor Dr. Galang have actual or potential conflicts of interest in relation to this program/presentation. Background United States State of Indiana: Pop 6,596,850 Scott County: Pop 24,181 Town of Austin: Pop 4,200 Abstract A-729-0201-03716, J. Duwve et al. “Community outbreak of HIV infection linked to injection drug use of oxymorphone – Indiana, 2015“ Poster WELBPE25, M. Spiller et al. “ Network analysis of a contact network from an investigation of a community outbreak of HIV infection linked to injection drug use of oxymorphone – Indiana, 2015” Public Health Importance In an outbreak setting, determining the number, size, and recency of infection clusters is important to guide the most effective interventions Laboratory support in this outbreak was used to: • distinguish recent infections from prevalent infections • suggest a common source between related infection strains Traditionally these analyses are applied retrospectively once the entire outbreak cohort has been defined This investigation is one of few to use these techniques in realtime to inform decision making during the outbreak response Objective To infer the timing of HIV transmission relative to hepatitis C virus (HCV) transmission To infer the relatedness among HIV-positive isolates Specimen Collection Methods We analyzed serum and plasma samples from: • Residents of Scott County and surrounding counties • Positive antibody results for HIV and/or HCV • Collected from October 2014 through April 2015 Photo: EISO Romeo Galang reviews specimen collection with Disease Intervention Specialists Figure: Source Counties for specimen collection HIV-related Laboratory Methods Recency of HIV-1 infection was determined by avidity testing • Modified Bio-Rad HIV 1/2 plus O assay (BRAI) • “Recent” infections are those that occurred within prior 6 months Clustering was assessed by sequencing HIV-1 polymerase (pol) gene sequences by polymerase chain reaction (PCR) HIV-1 phylogenetic clusters were defined when: 1. HIV-1 pol sequences were highly genetically related (>97% nucleotide identity) 2. Statistical evidence supporting relatedness was high (Shimodaira-Hasegawa probabilities >0.99) HCV-related Laboratory Methods HCV NS5B and HVR1 gene sequences were amplified by PCR HCV phylogenetic clusters were defined when: • NS5B sequences belonged to the same genotype • HVR1 sequences were highly genetically related (>96.23% nucleotide identity) Results HIV-1 Recency Results (N=49) 100 90 Avidity Index (%) 80 70 60 50 40 Recent Prevalent 30 20 10 0 Dec-7 Dec-27 Jan-16 Feb-5 Feb-25 Mar-17 Apr-6 Date of Specimen Collection Note: Recency testing was performed using a Modified BioRad 1/2 Plus O Assay with Avidity Index cutoff set at 30% Maximum likelihood phylogenetic tree of HIV-1 pol sequences (n=57) 1 0.98 0.83 1 0.9 0.77 0.95 0.76 0.95 1 0.88 0.89 0.94 0.84 0.95 0.79 0.83 0.93 0.790.75 0.77 0.9 1 0.85 0.89 0.97 0.84 0.85 0.99 Indiana outbreak samples* References samples 1 1 1 0.97 0.93 0.92 0.98 0.96 1 0.93 0.83 0.86 0.92 0.86 0.91 1 0.87 0.93 0.97 0.9 0.780.84 0.85 0.98 0.83 0.77 0.89 0.79 0.7 0.78 1 1 0.97 1 0.02 * Tested as part of epidemiologic investigation Maximum likelihood phylogenetic tree of HCV NS5b sequences (n=119) Indiana HCV Isolates North America References Cluster 1 1a 1b 2b 3a Cluster 3 Example of non-clustering sequences among reference sequences Cluster 2 HCV Cluster distribution within HIV-1 pol phylogenetic tree (n=57) 0.98 1 0.83 1 0.9 0.77 0.95 0.76 0.95 1 0.88 0.89 0.94 0.84 0.95 0.79 0.83 0.93 0.790.75 0.77 0.9 1 Distribution of HCV Clusters within HIV Cluster 1 (n = 55) 0.85 0.89 0.97 0.84 HCV Neg-Unk 28% HCV Cluster 1 0.85 0.99 HCV Cluster 2 Cluster 1 55 patients HCV HCVNegative Neg-Unk 35% HCV Cluster 1 1 HCV no cluster 1 HCV Cluster 2 1 HCV No Cluster 0.97 0.93 0.92 0.98 0.96 0.02 0.83 0.86 0.92 0.86 0.91 1 0.87 0.93 0.97 0.9 0.780.84 0.85 0.98 0.83 0.77 0.89 0.79 0.7 0.78 1 0.97 1 1 1 0.93 15% 22% Limitations This analysis is limited to a subset of the outbreak cohort; however, specimen collection and analysis are ongoing Phylogenetic analysis only suggests a common source of infection and cannot determine direction of transmission Conclusions A single strain of HIV-1 was recently introduced into a population infected with multiple circulating HCV strains Laboratory analyses can be conducted in real-time to meet immediate decision making needs Public Health Response Tailored interventions were implemented in a geographically concentrated area rather than broadly across the region Retesting of contacts was performed given the large number of recent HIV infections and on-going risk behavior • Monitor pol sequences of new infections for spread of outbreak strain Future HIV infections are sequenced and compared with representative outbreak sequences Acknowledgements • • • • • • • • • Indiana State Department of Health (ISDH) Scott County Health Department Clark County Health Department Foundations Family Medicine Indiana University School of Medicine, Division of Infectious Diseases University of Louisville School of Medicine, Division of Infectious Diseases Division of HIV/AIDS Prevention (DHAP) Division of Viral Hepatitis (DVH) Epidemic Intelligence Service (EIS) Program Office For more information please contact Centers for Disease Control and Prevention 1600 Clifton Road NE, Atlanta, GA 30333 Telephone: 1-800-CDC-INFO (232-4636)/TTY: 1-888-232-6348 Visit: www.cdc.gov | Contact CDC at: 1-800-CDC-INFO or www.cdc.gov/info The findings and conclusions in this report are those of the authors and do not necessarily represent the official position of the Centers for Disease Control and Prevention. National Center for HIV/AIDS, Viral Hepatitis, STD, and TB Prevention Division Name in this space Thank You! For more information please contact Centers for Disease Control and Prevention 1600 Clifton Road NE, Atlanta, GA 30333 Telephone: 1-800-CDC-INFO (232-4636)/TTY: 1-888-232-6348 Visit: www.cdc.gov | Contact CDC at: 1-800-CDC-INFO or www.cdc.gov/info The findings and conclusions in this report are those of the authors and do not necessarily represent the official position of the Centers for Disease Control and Prevention. National Center for HIV/AIDS, Viral Hepatitis, STD, and TB Prevention Division Name in this space