Sequenced Mitochondrial Genomes of Bryophytes
... determined using electron microscopy and restriction endonuclease mapping. The mitochondrial genome of M. polymorpha was found to be a single circular molecule which consists of about 186609 base pairs (bp). Several genes including genes for three species of ribosomal RNA, transfer RNA and 30 open r ...
... determined using electron microscopy and restriction endonuclease mapping. The mitochondrial genome of M. polymorpha was found to be a single circular molecule which consists of about 186609 base pairs (bp). Several genes including genes for three species of ribosomal RNA, transfer RNA and 30 open r ...
ABSTRACT Title of Document: PROGRAMMED
... allows for translocation of the ribosome (Uemura et al, 2007). Eukaryotic initiation is much more complicated. eIF1, eIF1A, eIF3, and eIF5 combine to form the multifactor complex (Fekete et al, 2007). All of these factors are required to recruit the 43S pre-initiation complex, which includes the ini ...
... allows for translocation of the ribosome (Uemura et al, 2007). Eukaryotic initiation is much more complicated. eIF1, eIF1A, eIF3, and eIF5 combine to form the multifactor complex (Fekete et al, 2007). All of these factors are required to recruit the 43S pre-initiation complex, which includes the ini ...
Translational control of regA, a key gene controlling
... Kochert, 1982; Pommerville and Kochert, 1981; Schmitt, 2003). Three types of genes – gls, lag and regA – are responsible for cell differentiation in V. carteri. The regA gene is a master control gene that is expressed only in somatic cells, where it suppresses all reproductive activity. If the regA ...
... Kochert, 1982; Pommerville and Kochert, 1981; Schmitt, 2003). Three types of genes – gls, lag and regA – are responsible for cell differentiation in V. carteri. The regA gene is a master control gene that is expressed only in somatic cells, where it suppresses all reproductive activity. If the regA ...
Phylogenetic Relationships between the Western Aster Yellows
... conditions used for restriction enzymes, calf intestinal alkaline phosphatase (Boehringer Mannheim), exonuclease I11 (ExoIII) (Bethesda Research Laboratories), the Klenow enzyme (New England Biolabs), T4 DNA ligase (New England BioLabs), and Sequenase (Amersham) were those suggested by the manufactu ...
... conditions used for restriction enzymes, calf intestinal alkaline phosphatase (Boehringer Mannheim), exonuclease I11 (ExoIII) (Bethesda Research Laboratories), the Klenow enzyme (New England Biolabs), T4 DNA ligase (New England BioLabs), and Sequenase (Amersham) were those suggested by the manufactu ...
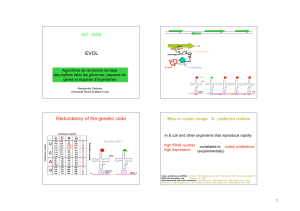
The Coevolution of Genes and Genetic Codes: Crick`s Frozen
... coevolution of codes and messages would have taken a different course. Crick suggests that at that stage the formation of the code proceeded by the introduction of new amino acids and by an increase in the precision of recognition of amino acids within classes. Such changes in the code were likely to ...
... coevolution of codes and messages would have taken a different course. Crick suggests that at that stage the formation of the code proceeded by the introduction of new amino acids and by an increase in the precision of recognition of amino acids within classes. Such changes in the code were likely to ...
Vaginal TM7 and the absorption of amino acids
... depleted resulting in an unbalanced microbiome. It has been shown to affect up to 36% of women, with half of these cases being asymptomatic, where the other half reported a “fishysmelling” discharge (Hay). Bacterial Vaginosis occurs when there is a depletion of Lactobacillus, and an overgrowth of an ...
... depleted resulting in an unbalanced microbiome. It has been shown to affect up to 36% of women, with half of these cases being asymptomatic, where the other half reported a “fishysmelling” discharge (Hay). Bacterial Vaginosis occurs when there is a depletion of Lactobacillus, and an overgrowth of an ...
Correlation of amino acid preference and
... Three types of analysis are conducted. At the genome level, crossvalidation is used to demonstrate the predictive capability of amino acid preference of viral genome type. Next at the sequence level, two types of resampling analysis are used to investigate the correlation of amino acid preference an ...
... Three types of analysis are conducted. At the genome level, crossvalidation is used to demonstrate the predictive capability of amino acid preference of viral genome type. Next at the sequence level, two types of resampling analysis are used to investigate the correlation of amino acid preference an ...
Localization and structural analysis of the ribosomal RNA operons of
... E. coli rRNA operons. The fact that insertion of a cartridge resulted in the loss of the wild type signal, leads to the conclusion that there are only three rRNA operons in R. sphaeroides. Thus rmA can be assigned to the 10 kb BamYU signal, rrnB to the 14 kb BamYU signal, and rmC to the 13 kb BamYU ...
... E. coli rRNA operons. The fact that insertion of a cartridge resulted in the loss of the wild type signal, leads to the conclusion that there are only three rRNA operons in R. sphaeroides. Thus rmA can be assigned to the 10 kb BamYU signal, rrnB to the 14 kb BamYU signal, and rmC to the 13 kb BamYU ...
mRNA Export - e
... (mRNPs). RBPs assemble on nascent and mature mRNAs. They serve as structural elements for mRNP packaging and modify the output of gene expression at all steps of the mRNA life cycle: ...
... (mRNPs). RBPs assemble on nascent and mature mRNAs. They serve as structural elements for mRNP packaging and modify the output of gene expression at all steps of the mRNA life cycle: ...
Mechanisms of translational regulation in bacteria
... the growing peptide chain is determined by triplets of nucleotides, so called codons. However, there are only 20 amino acids but 64 different triplets of nucleotides encoding them. Consequently, the genetic code is degenerate: Except for tryptophan and methionine, the amino acids are encoded by two, ...
... the growing peptide chain is determined by triplets of nucleotides, so called codons. However, there are only 20 amino acids but 64 different triplets of nucleotides encoding them. Consequently, the genetic code is degenerate: Except for tryptophan and methionine, the amino acids are encoded by two, ...
The mitochondrial genome of the soybean cyst nematode
... infest a number of indicator soybean cultivars. Genetic approaches for distinguishing H. glycines populations could provide a means of rapidly distinguishing these types. However, such genetic markers have not yet been developed. The mitochondrial genome of H. glycines may provide such genetic marke ...
... infest a number of indicator soybean cultivars. Genetic approaches for distinguishing H. glycines populations could provide a means of rapidly distinguishing these types. However, such genetic markers have not yet been developed. The mitochondrial genome of H. glycines may provide such genetic marke ...
Phylogenetic Network and Physicochemical Properties of
... higher than the estimates derived from phylogenetic analyses, suggesting that a significant fraction of mutations is removed by selection (Parsons et al. 1997). The effects of nonsynonymous mutations depend both on the position of the amino acid replacement in the protein sequence and on the physico ...
... higher than the estimates derived from phylogenetic analyses, suggesting that a significant fraction of mutations is removed by selection (Parsons et al. 1997). The effects of nonsynonymous mutations depend both on the position of the amino acid replacement in the protein sequence and on the physico ...
Module 5: Alternative Open Reading Frame
... over the first nucleotide of the highlighted in the start codon and a popup box will show up that has the nucleotide number indicated. Make a note of the number. Scroll down the page until you come to the highlighted stop codon in the same reading frame. Hover your cursor over the LAST nucleotide in ...
... over the first nucleotide of the highlighted in the start codon and a popup box will show up that has the nucleotide number indicated. Make a note of the number. Scroll down the page until you come to the highlighted stop codon in the same reading frame. Hover your cursor over the LAST nucleotide in ...
Ribosome profiling reveals post-transcriptional buffering of divergent
... C. Joel McManus,1 Gemma E. May, Pieter Spealman, and Alan Shteyman Carnegie Mellon University, Department of Biological Sciences, Pittsburgh, Pennsylvania 15213, USA Understanding the patterns and causes of phenotypic divergence is a central goal in evolutionary biology. Much work has shown that mRN ...
... C. Joel McManus,1 Gemma E. May, Pieter Spealman, and Alan Shteyman Carnegie Mellon University, Department of Biological Sciences, Pittsburgh, Pennsylvania 15213, USA Understanding the patterns and causes of phenotypic divergence is a central goal in evolutionary biology. Much work has shown that mRN ...
Improving Virus C type 4 Interferon using Bioinformatics Techniques
... the anticodon of tRNA carrying methionine. A large ribosomal subunit binds to the complex, and the reactions of protein synthesis itself can begin. The aminoacyl-tRNA to be called for next is determined by the next codon (the next three bases) on the mRNA as shown in figure [3]. Each amino acid is c ...
... the anticodon of tRNA carrying methionine. A large ribosomal subunit binds to the complex, and the reactions of protein synthesis itself can begin. The aminoacyl-tRNA to be called for next is determined by the next codon (the next three bases) on the mRNA as shown in figure [3]. Each amino acid is c ...
LESSON 4 Using Bioinformatics to Analyze Protein
... 16. Now it is time to practice translation. Show Slide #7, which contains a classic codon table, and remind students that amino acids are encoded by nucleotide triplets called codons. Walk students through the codon table, using the start codon ATG [AUG] as an example. Remind students that protein t ...
... 16. Now it is time to practice translation. Show Slide #7, which contains a classic codon table, and remind students that amino acids are encoded by nucleotide triplets called codons. Walk students through the codon table, using the start codon ATG [AUG] as an example. Remind students that protein t ...
Full-Text PDF
... involved, and mutations in these genes may lead to the various diseases mentioned. Almost all these genes are encoded by the nuclear genome, and only a few, coding for some components of the respiratory chain, tRNAs and rRNAs, are encoded by the mitochondrial genome. In humans, this amounts to a tot ...
... involved, and mutations in these genes may lead to the various diseases mentioned. Almost all these genes are encoded by the nuclear genome, and only a few, coding for some components of the respiratory chain, tRNAs and rRNAs, are encoded by the mitochondrial genome. In humans, this amounts to a tot ...
E.coli
... Can we use this signal to deduce some more biological information ? We determined the most important metabolic networks in a (translationally biased) organism Can we determine genes belonging to minimal gene sets ? ...
... Can we use this signal to deduce some more biological information ? We determined the most important metabolic networks in a (translationally biased) organism Can we determine genes belonging to minimal gene sets ? ...
Fig 1 - Centre for Biodiversity Genomics
... ranges fivefold (0.07–0.35). BIN Distances show great variation, ranging from 0.01 or less in 12 families to more than 0.25 in eight families. Patterns of amino acid substitution in COI are generally congruent with previously reported variation in nucleotide substitution rates in arachnids, but prov ...
... ranges fivefold (0.07–0.35). BIN Distances show great variation, ranging from 0.01 or less in 12 families to more than 0.25 in eight families. Patterns of amino acid substitution in COI are generally congruent with previously reported variation in nucleotide substitution rates in arachnids, but prov ...
1 Chapter 1 Chemistry On The Pyrimidine Ring
... The general mechanism for both of these enzymes first involves a sulfur transfer from a donor such as IscS for ThiI, and TusE for MnmA. This yields a protein persulfide intermediate. The enzymes then bind the tRNA and activates the uridine oxygen using ATP to form an adenylated intermediate. The su ...
... The general mechanism for both of these enzymes first involves a sulfur transfer from a donor such as IscS for ThiI, and TusE for MnmA. This yields a protein persulfide intermediate. The enzymes then bind the tRNA and activates the uridine oxygen using ATP to form an adenylated intermediate. The su ...
M2 RNA Pol Ⅰ genes
... B upstream binding factor (UBF) binds to both the UCE and the upstream part of the core element of the RNA Pol I promoter. C selectivity factor SLl stabilizes the UBF-DNA complex. D SL1 contains several subunits including the TATA-binding protein TBP. E in Acanthamoeba there is a single control elem ...
... B upstream binding factor (UBF) binds to both the UCE and the upstream part of the core element of the RNA Pol I promoter. C selectivity factor SLl stabilizes the UBF-DNA complex. D SL1 contains several subunits including the TATA-binding protein TBP. E in Acanthamoeba there is a single control elem ...
Constructing Sequences for Oxytocin and Vasopressin
... Constructing Sequences for Oxytocin and Vasopressin Introduction: The structure of oxytocin is very similar to that of the vasopressin (also known commonly as arginine vasopressin): Both are nonapeptides (peptides with nine amino acids) with a disulfide bridge and their amino acid sequence differs a ...
... Constructing Sequences for Oxytocin and Vasopressin Introduction: The structure of oxytocin is very similar to that of the vasopressin (also known commonly as arginine vasopressin): Both are nonapeptides (peptides with nine amino acids) with a disulfide bridge and their amino acid sequence differs a ...
Probable presence of an ubiquitous cryptic mitochondrial gene on
... Presentation of the hypothesis: A positionally conserved ORF has been found on the complementary strand of the cox1 genes of both eukaryotic mitochondria (protist, plant, fungal and animal) and alpha-proteobacteria. This putative gene has been named gau for gene antisense ubiquitous in mtDNAs. The l ...
... Presentation of the hypothesis: A positionally conserved ORF has been found on the complementary strand of the cox1 genes of both eukaryotic mitochondria (protist, plant, fungal and animal) and alpha-proteobacteria. This putative gene has been named gau for gene antisense ubiquitous in mtDNAs. The l ...
RNA Polymerases
... regulation of their transcription. Some promoters such as the U6 small nuclear RNA (U6 snRNA ) and small RNA genes from the Epstein-Barr virus use only regulatory sequences upstream from their transcription start sites. The coding region of the U6 snRNA has a characteristic A box. However, this sequ ...
... regulation of their transcription. Some promoters such as the U6 small nuclear RNA (U6 snRNA ) and small RNA genes from the Epstein-Barr virus use only regulatory sequences upstream from their transcription start sites. The coding region of the U6 snRNA has a characteristic A box. However, this sequ ...
DNA MUTATION, REPAIR, AND TRANSPOSITION
... Therefore, DNA molecule I is the least sensitive, while molecule III is the most sensitive. 24. Frameshift mutations are caused by insertions or deletions of bases (that are not multiples of 3). These will shift the reading frame for all codons downstream from the mutation. Single base-substitutions ...
... Therefore, DNA molecule I is the least sensitive, while molecule III is the most sensitive. 24. Frameshift mutations are caused by insertions or deletions of bases (that are not multiples of 3). These will shift the reading frame for all codons downstream from the mutation. Single base-substitutions ...
Transfer RNA

A transfer RNA (abbreviated tRNA and archaically referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length, that serves as the physical link between the mRNA and the amino acid sequence of proteins. It does this by carrying an amino acid to the protein synthetic machinery of a cell (ribosome) as directed by a three-nucleotide sequence (codon) in a messenger RNA (mRNA). As such, tRNAs are a necessary component of translation, the biological synthesis of new proteins according to the genetic code.The specific nucleotide sequence of an mRNA specifies which amino acids are incorporated into the protein product of the gene from which the mRNA is transcribed, and the role of tRNA is to specify which sequence from the genetic code corresponds to which amino acid. One end of the tRNA matches the genetic code in a three-nucleotide sequence called the anticodon. The anticodon forms three base pairs with a codon in mRNA during protein biosynthesis. The mRNA encodes a protein as a series of contiguous codons, each of which is recognized by a particular tRNA. On the other end of the tRNA is a covalent attachment to the amino acid that corresponds to the anticodon sequence. Each type of tRNA molecule can be attached to only one type of amino acid, so each organism has many types of tRNA (in fact, because the genetic code contains multiple codons that specify the same amino acid, there are several tRNA molecules bearing different anticodons which also carry the same amino acid).The covalent attachment to the tRNA 3’ end is catalyzed by enzymes called aminoacyl tRNA synthetases. During protein synthesis, tRNAs with attached amino acids are delivered to the ribosome by proteins called elongation factors (EF-Tu in bacteria, eEF-1 in eukaryotes), which aid in decoding the mRNA codon sequence. If the tRNA's anticodon matches the mRNA, another tRNA already bound to the ribosome transfers the growing polypeptide chain from its 3’ end to the amino acid attached to the 3’ end of the newly delivered tRNA, a reaction catalyzed by the ribosome.A large number of the individual nucleotides in a tRNA molecule may be chemically modified, often by methylation or deamidation. These unusual bases sometimes affect the tRNA's interaction with ribosomes and sometimes occur in the anticodon to alter base-pairing properties.