EE 400: Practice using NCBI, Blast and Clustal
... B. You will see a big box for “Input sequences.” Go to your Word document with all the sequences, and copy them all from the first > to the last amino acid. Paste this into the box. The format we have been using is called fastA, which is accepted by this program. Make sure you are not putting any bl ...
... B. You will see a big box for “Input sequences.” Go to your Word document with all the sequences, and copy them all from the first > to the last amino acid. Paste this into the box. The format we have been using is called fastA, which is accepted by this program. Make sure you are not putting any bl ...
Proteins - Madison Public Schools
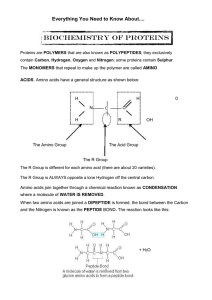
... The covalent bond that results from dehydration synthesis is known as a PEPTIDE BOND. When bonding together amino acids to build proteins, the carboxyl group from one amino acid has to match up with the amino group of the next amino acid. This is so that dehydration synthesis can take ...
... The covalent bond that results from dehydration synthesis is known as a PEPTIDE BOND. When bonding together amino acids to build proteins, the carboxyl group from one amino acid has to match up with the amino group of the next amino acid. This is so that dehydration synthesis can take ...
Dr Alanna Easton`s Travelling Scholarship Report, April 2014
... central hypothesis of my work states that differential neuronal structure, and the specific pattern of development in the medium spiny neurons of the striatum will predict impulsivity. In agreement with my own research interests and given the relatively short amount of time I had at Mount Sinai, I ...
... central hypothesis of my work states that differential neuronal structure, and the specific pattern of development in the medium spiny neurons of the striatum will predict impulsivity. In agreement with my own research interests and given the relatively short amount of time I had at Mount Sinai, I ...
Protein levels with and without Monensin for
... Service. Contents of this publication may be freely reproduced for educational purposes. All other rights reserved. Brand names appearing in this publication are for product identification purposes only. K-State Research and Extension is an equal opportunity provider and employer. ...
... Service. Contents of this publication may be freely reproduced for educational purposes. All other rights reserved. Brand names appearing in this publication are for product identification purposes only. K-State Research and Extension is an equal opportunity provider and employer. ...
Lecture 3: Contributions to protein stability
... energy of protein folding is the difference between two very large contributions: the large chain entropy loss upon folding and the large gain in hydrophobic interactions, it is not currently possible to predict overall protein stability even from high resolution crystal structures. • Much more ame ...
... energy of protein folding is the difference between two very large contributions: the large chain entropy loss upon folding and the large gain in hydrophobic interactions, it is not currently possible to predict overall protein stability even from high resolution crystal structures. • Much more ame ...
Media - Inside Cancer
... a phosphate group. The source of the phosphate group is ATP. Kinases serve as relay molecules in the cell signaling pathway. Once activated, they activate other proteins, hence transmitting the message through the cell. 2. Since all cell signaling pathways end with a cellular response, explain why i ...
... a phosphate group. The source of the phosphate group is ATP. Kinases serve as relay molecules in the cell signaling pathway. Once activated, they activate other proteins, hence transmitting the message through the cell. 2. Since all cell signaling pathways end with a cellular response, explain why i ...
Problem Set 1
... C. The blood of these worms contains a hemoglobin of 3780 kDa. Each molecule of this hemoglobin is comprised of 120 globin chains (each chain of 16.5 kDa and with one heme group) surrounding a core of 72 linker chains (each of 25 kDa). The P50 of the Pompeii worm Hb (Hbp) is 0.4 torr and the Hill co ...
... C. The blood of these worms contains a hemoglobin of 3780 kDa. Each molecule of this hemoglobin is comprised of 120 globin chains (each chain of 16.5 kDa and with one heme group) surrounding a core of 72 linker chains (each of 25 kDa). The P50 of the Pompeii worm Hb (Hbp) is 0.4 torr and the Hill co ...
Leatherbarrow talk
... • There are a few examples of successful drug leads that are targeted at Protein-Protein interfaces • However, there are currently NO marketed drugs that work this way… ...
... • There are a few examples of successful drug leads that are targeted at Protein-Protein interfaces • However, there are currently NO marketed drugs that work this way… ...
slides
... employ kinetic Monte Carlo simulation to model the time-dependent folding during transcription of riboswitch expression platforms. both that riboswitch transcriptional terminator sequences have been naturally selected for high folding efficiency, and that sequesterers can maintain their function eve ...
... employ kinetic Monte Carlo simulation to model the time-dependent folding during transcription of riboswitch expression platforms. both that riboswitch transcriptional terminator sequences have been naturally selected for high folding efficiency, and that sequesterers can maintain their function eve ...
Amino Acids 2
... 2. Gel-filtration chromatography separates a mixture of proteins on the basis of: A) size B) charge C) affinity for ligands in the column matrix D) density 3. What is the purpose of treating a protein with 2-mercaptoethanol? A) To hydrolyze the protein into its amino acids. B) To derivatize any free ...
... 2. Gel-filtration chromatography separates a mixture of proteins on the basis of: A) size B) charge C) affinity for ligands in the column matrix D) density 3. What is the purpose of treating a protein with 2-mercaptoethanol? A) To hydrolyze the protein into its amino acids. B) To derivatize any free ...
Protein Folding, Shape, and Function Activity Instructions
... proteins and other structures at the molecular level. In this activity you will explore the structure of proteins and the chemical interactions that drive each protein to fold into its specific structure. Each protein is made of a specific sequence of amino acids. There are 20 amino acids involved i ...
... proteins and other structures at the molecular level. In this activity you will explore the structure of proteins and the chemical interactions that drive each protein to fold into its specific structure. Each protein is made of a specific sequence of amino acids. There are 20 amino acids involved i ...
Comparative Proteomics Kit I: Protein Profiler Module
... • Traditional classification based upon traits: – Morphological – Behavioral ...
... • Traditional classification based upon traits: – Morphological – Behavioral ...
6. protein folding
... • Folding means arriving at the right combinations of angles for every amino acid residue in the sequence. • Proteins fold on a defined pathway (or a small number of alternative pathways); they don't randomly search all possible conformations until they arrive at the most stable (lowest free energy ...
... • Folding means arriving at the right combinations of angles for every amino acid residue in the sequence. • Proteins fold on a defined pathway (or a small number of alternative pathways); they don't randomly search all possible conformations until they arrive at the most stable (lowest free energy ...
No Slide Title
... (ie, random coil) • ~ uniform charge/mass ratio due to SDS • therefore endogenous charge and shape are not major factors • mobility is inverse of mass (charge) mobility (voltage) (mass) ...
... (ie, random coil) • ~ uniform charge/mass ratio due to SDS • therefore endogenous charge and shape are not major factors • mobility is inverse of mass (charge) mobility (voltage) (mass) ...
Resistance exercise volume affects myofibrillar protein synthesis
... Fed post-exercise. Fed-state MPS was transiently elevated above rest at 5 h for 1SET (2.3-fold) and returned to resting levels by 29 h post-exercise. However, the exercise induced increase in MPS following 3SET was superior in amplitude and duration as compared to 1SET at both 5 h (3.1-fold above re ...
... Fed post-exercise. Fed-state MPS was transiently elevated above rest at 5 h for 1SET (2.3-fold) and returned to resting levels by 29 h post-exercise. However, the exercise induced increase in MPS following 3SET was superior in amplitude and duration as compared to 1SET at both 5 h (3.1-fold above re ...
Water, Protein, and Nutrients
... Proteins are another type of ___________________________________ needed for life Meats such as ___________________________________contain large amounts of protein Like fats and carbs, proteins also ___________________________________to living things Proteins help control ____________________ ...
... Proteins are another type of ___________________________________ needed for life Meats such as ___________________________________contain large amounts of protein Like fats and carbs, proteins also ___________________________________to living things Proteins help control ____________________ ...
UTM EatWell
... What is Protein? Protein is one of the three major nutrients, along with carbohydrate and fat, that fuels our body. Dietary protein is digested into amino acids, which are the building blocks our body uses to build and maintain muscle, skin, hair, connective tissue and ...
... What is Protein? Protein is one of the three major nutrients, along with carbohydrate and fat, that fuels our body. Dietary protein is digested into amino acids, which are the building blocks our body uses to build and maintain muscle, skin, hair, connective tissue and ...
ProteinPrediction
... By definition, proteins that are more than 50% identical in amino acid sequence across their entire length are said to be members of a single family. Superfamilies are groups of protein families that are related by lower but still detectable levels of sequence similarity (and therefore have a common ...
... By definition, proteins that are more than 50% identical in amino acid sequence across their entire length are said to be members of a single family. Superfamilies are groups of protein families that are related by lower but still detectable levels of sequence similarity (and therefore have a common ...
Support vector machines for protein function prediction
... • A protein is classified as either belong (+) or not belong (-) to a functional family • By screening against all families, the function of this protein can be identified (example: SVMProt) ...
... • A protein is classified as either belong (+) or not belong (-) to a functional family • By screening against all families, the function of this protein can be identified (example: SVMProt) ...
AS Biology - Everything Protein
... The R Group is different for each amino acid (there are about 20 varieties). The R Group is ALWAYS opposite a lone Hydrogen off the central carbon. ...
... The R Group is different for each amino acid (there are about 20 varieties). The R Group is ALWAYS opposite a lone Hydrogen off the central carbon. ...
BiochemLecture07
... • RanGTP enhances binding between an exportin and its cargo but stimulates release of importin's cargo; RanGDT has the opposite effect, namely, it stimulates the release of exportin's cargo, but enhances the binding between an importin and its cargo. Therefore, the exportin and its cargo may move t ...
... • RanGTP enhances binding between an exportin and its cargo but stimulates release of importin's cargo; RanGDT has the opposite effect, namely, it stimulates the release of exportin's cargo, but enhances the binding between an importin and its cargo. Therefore, the exportin and its cargo may move t ...
Protein Modification, targeting and degradation Protein modification
... Ran helps move importins and exportins and their cargo in and out of the nucleus • RanGTP enhances binding between an exportin and its cargo but stimulates release of importin's cargo; RanGDT has the opposite effect, namely, it stimulates the release of exportin's cargo, but enhances the binding be ...
... Ran helps move importins and exportins and their cargo in and out of the nucleus • RanGTP enhances binding between an exportin and its cargo but stimulates release of importin's cargo; RanGDT has the opposite effect, namely, it stimulates the release of exportin's cargo, but enhances the binding be ...
Folds
... protein tertiary structures are divided into five main classes according to the secondary structure content of their domains: all-a domains, all-b domains (b-barrels, e.g. Greek key motif), a+b domains (irregular fashion of arrangement), a/b domains (b-a-b motifs) and “others” each class contains ma ...
... protein tertiary structures are divided into five main classes according to the secondary structure content of their domains: all-a domains, all-b domains (b-barrels, e.g. Greek key motif), a+b domains (irregular fashion of arrangement), a/b domains (b-a-b motifs) and “others” each class contains ma ...
Protein Folding
... • Collapsed, with native-like 2º structure (far UV CD) • Weak or transient side-chain interactions (near UV CD) • Binds hydrophobic dyes • Many proteins form molten globules at low pH • Model for early stages of protein folding (hydrophobic collapse) ...
... • Collapsed, with native-like 2º structure (far UV CD) • Weak or transient side-chain interactions (near UV CD) • Binds hydrophobic dyes • Many proteins form molten globules at low pH • Model for early stages of protein folding (hydrophobic collapse) ...
Bimolecular fluorescence complementation

Bimolecular fluorescence complementation (also known as BiFC) is a technology typically used to validate protein interactions. It is based on the association of fluorescent protein fragments that are attached to components of the same macromolecular complex. Proteins that are postulated to interact are fused to unfolded complementary fragments of a fluorescent reporter protein and expressed in live cells. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. This fluorescent signal can be detected and located within the cell using an inverted fluorescence microscope that allows imaging of fluorescence in cells. In addition, the intensity of the fluorescence emitted is proportional to the strength of the interaction, with stronger levels of fluorescence indicating close or direct interactions and lower fluorescence levels suggesting interaction within a complex. Therefore, through the visualisation and analysis of the intensity and distribution of fluorescence in these cells, one can identify both the location and interaction partners of proteins of interest.